

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146705.12 - phase: 0 /pseudo
(811 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
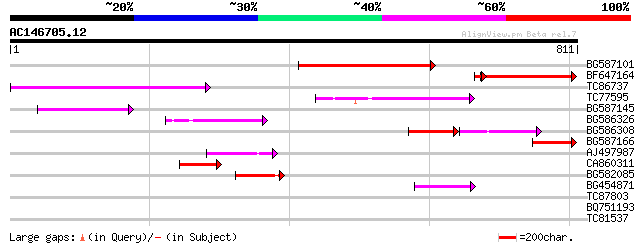
Score E
Sequences producing significant alignments: (bits) Value
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 309 2e-84
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 180 3e-51
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 167 2e-41
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 105 5e-23
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 93 4e-19
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 87 3e-17
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 59 4e-16
BG587166 similar to GP|8778801|gb| T32E20.9 {Arabidopsis thalian... 67 3e-11
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 58 1e-08
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 53 5e-07
BG582085 51 2e-06
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 50 5e-06
TC87803 homologue to GP|1871187|gb|AAB63547.1| unknown protein {... 30 4.8
BQ751193 similar to GP|21622336|em probable multidrug resistance... 30 4.8
TC81537 similar to GP|7630037|emb|CAB88331.1 putative protein {A... 29 8.2
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 309 bits (792), Expect = 2e-84
Identities = 140/197 (71%), Positives = 165/197 (83%)
Frame = +2
Query: 413 DLKHYYWDEPYLFRRGSDGIFRRCIPENEVSSILTHCHSSSYGGHASTQKTSFKILHSGF 472
D++ Y WDEP+L+++ +D I+ RC+ E E+ IL HCH S+Y GH + KT KI +GF
Sbjct: 8 DVRRYLWDEPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGF 187
Query: 473 WWPSLFKDVHLFISKCDKCQRTGSITKRNEMPLNNILEVEIFDVWGIDFMGPFPSSFGNQ 532
WWP++FKD H FISKCD CQR G+I+ RNEMP N ILEVE+FDVWGIDFMGPFPSS+ N+
Sbjct: 188 WWPTMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPSSYNNK 367
Query: 533 YILVAVDYVSKWVEAIASPTNDAQVVIKMFKKVIFPRFGVPRVVISDGGSHFISRHFEKL 592
YILVAVDYVSKWVEAIASPTNDA VV+KMFK VIFPRFGVPRVVISDGGSHFI++ FEKL
Sbjct: 368 YILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKL 547
Query: 593 LQKLGVRHKIATPYHPQ 609
L+K GVRHK+AT YHPQ
Sbjct: 548 LKKNGVRHKVATAYHPQ 598
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 180 bits (457), Expect(2) = 3e-51
Identities = 88/137 (64%), Positives = 106/137 (77%)
Frame = +1
Query: 674 LEHKAYWAIRNLNLDPNLAGDKRKLQLNELEELRMDAYENARIYKERTKTWHDKKIIKRH 733
L K I+ LN+D AG KR L+LNELEELR+DAYENA+IYKERTK WHD++II+R
Sbjct: 28 LSIKPIGPIKALNMDYKAAGGKRVLELNELEELRLDAYENAKIYKERTKKWHDRRIIRRE 207
Query: 734 FKSGDLVLLFNSRLKLFPGKLRSRWSGPFQVRTVYPYGAIEIFSEETGSFTVNGQRLKIY 793
F+ G+LVLLFNSRLKLFPGKLRS WSGPFQV+ V P GA+E++SE TG FTVNGQRLK Y
Sbjct: 208 FREGELVLLFNSRLKLFPGKLRSHWSGPFQVKNVMPSGAVEVWSESTGPFTVNGQRLKHY 387
Query: 794 NTGEVNEVVADFTLSDP 810
+GE + + L+ P
Sbjct: 388 TSGENVDKGVRYALTKP 438
Score = 40.8 bits (94), Expect(2) = 3e-51
Identities = 16/16 (100%), Positives = 16/16 (100%)
Frame = +3
Query: 666 KSCHLPVELEHKAYWA 681
KSCHLPVELEHKAYWA
Sbjct: 3 KSCHLPVELEHKAYWA 50
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 167 bits (422), Expect = 2e-41
Identities = 103/290 (35%), Positives = 150/290 (51%), Gaps = 3/290 (1%)
Frame = +1
Query: 1 NKATRKDHFPLPFIDQMLERLAKHSHFCYLDGYSGFFQIPIHPNDQEKTTFTCPFGTFAY 60
N T+KD +PLP I + L R+A F LD + F ++ I DQEKT F +G F +
Sbjct: 643 NAITKKDRYPLPLISETLRRVAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEW 822
Query: 61 RRMPFGLCNAPATFQRCMMSIFSDFVEKIMEVFMDDFSVHGS-NFDDCLTNLEKVLERCE 119
PFGL APATFQR + +F++ + ++DD ++ + + D + +VL R
Sbjct: 823 IVCPFGLTGAPATFQRYINKTLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLA 1002
Query: 120 QVNLVLNWEKCHFMVREGIVLGH-LVFDRGIEVDRAKIEIIKKMLPPTSVKEIRSFLGHA 178
L L+ +KC F V +G L +G+ D K+ I+ LPP SVK RSFLG
Sbjct: 1003DAGLSLDPKKCEFSVTTVKYVGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFC 1182
Query: 179 GFYRRFIKDFSSITKPLTSLLLKDADFTFDDSCLQAFCRLKEALITAPIIQPPDWNLPFE 238
+Y+ FI +S IT+PLT L KD F + AF +LK P+++ D
Sbjct: 1183NYYKDFIPGYSEITEPLTRLTRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPEAVTT 1362
Query: 239 IMCDASDYAVGAVLGQRNDK-KMHAIYYASKTLDGAQVNYATTEKELLAV 287
+ D S +A+G VL Q + H + + S+ L A+ NY +KELLAV
Sbjct: 1363VETDCSGFALGGVLTQEDGTGAAHPVAFHSQRLSPAEYNYPIHDKELLAV 1512
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial
(14%)
Length = 1708
Score = 105 bits (263), Expect = 5e-23
Identities = 69/236 (29%), Positives = 116/236 (48%), Gaps = 8/236 (3%)
Frame = +2
Query: 438 PENEV-SSILTHCHSSSYGGHASTQKTSFKILHSGFWWPSLFKDVHLFISKCDKCQ---- 492
P NE+ + ++ H S+ GH T +I+ F+WP + V F+ CD C
Sbjct: 155 PLNELRTKLVQESHDSTAAGHPGRNGT-LEIVSRKFFWPGQSQTVRRFVRNCDVCGGIHI 331
Query: 493 -RTGSITKRNEMPLNNILEVEIFDVWGIDFMGPFPSSFG--NQYILVAVDYVSKWVEAIA 549
R +P+ N L ++ +DF+ P + G +QY+ V VD +SK V
Sbjct: 332 WRQAKRGFLKPLPVPNRLHSDL----SMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEE 499
Query: 550 SPTNDAQVVIKMFKKVIFPRFGVPRVVISDGGSHFISRHFEKLLQKLGVRHKIATPYHPQ 609
T +A+ + F + G+P+ ++SD GS+++ R + + + GV ++T YHPQ
Sbjct: 500 MDTMEAEACAQRFLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQ 679
Query: 610 TSGQVEVSNRQIKAILEKTVSTSRTDWSNKLDDALWAYRTAYKTPIGMTPFKLVYG 665
T G E N++I+A+L V S+ +W + L A R + + IG TPF + +G
Sbjct: 680 TDGGTERWNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHG 847
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 93.2 bits (230), Expect = 4e-19
Identities = 50/136 (36%), Positives = 79/136 (57%)
Frame = +2
Query: 41 IHPNDQEKTTFTCPFGTFAYRRMPFGLCNAPATFQRCMMSIFSDFVEKIMEVFMDDFSVH 100
+HP+D EKT F GT+ Y+ MPFGL NA +T+QR + +F+D + MEV++DD V
Sbjct: 2 MHPDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVK 181
Query: 101 GSNFDDCLTNLEKVLERCEQVNLVLNWEKCHFMVREGIVLGHLVFDRGIEVDRAKIEIIK 160
D L +L++ + ++ + LN KC F V G LG++V +GIEV+ +I I
Sbjct: 182 SLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAIL 361
Query: 161 KMLPPTSVKEIRSFLG 176
+ P + +E++ G
Sbjct: 362 DLPSPKNSREVQRLTG 409
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 86.7 bits (213), Expect = 3e-17
Identities = 52/145 (35%), Positives = 81/145 (55%)
Frame = +2
Query: 224 TAPIIQPPDWNLPFEIMCDASDYAVGAVLGQRNDKKMHAIYYASKTLDGAQVNYATTEKE 283
+API+ P+ + + + DAS +G VL Q I YAS+ L + NY T + E
Sbjct: 11 SAPILVLPEL-ITYVVYTDASITGLGCVLTQHEK----VIAYASRQLRKHEGNYPTHDLE 175
Query: 284 LLAVVYAIDKFRQYLVGSKIIVYTDHSAIKYLLNKKDAKPRLIRWILLLQEFDLEIKDKK 343
+ AVV+A+ +R YL G+K+ ++TDH ++KY+ + + R RW+ + ++DL+I
Sbjct: 176 MAAVVFALKIWRSYLYGAKVQIHTDHKSLKYIFTQPELNLRQRRWMEFVADYDLDITYYP 355
Query: 344 GVENVVADHLSRLRETNKDELPLDD 368
G N+VAD LSR R E DD
Sbjct: 356 GKANLVADALSRRRVDVSAEREADD 430
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 686
Score = 58.5 bits (140), Expect(2) = 4e-16
Identities = 26/71 (36%), Positives = 45/71 (62%)
Frame = -2
Query: 571 GVPRVVISDGGSHFISRHFEKLLQKLGVRHKIATPYHPQTSGQVEVSNRQIKAILEKTVS 630
G+P +++D GSHFIS F + ++ +R A+P +PQ++GQ E SN+ I L+K +
Sbjct: 682 GLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKIIIDGLKKRLD 503
Query: 631 TSRTDWSNKLD 641
+ W+++LD
Sbjct: 502 LKKGCWADELD 470
Score = 44.7 bits (104), Expect(2) = 4e-16
Identities = 31/119 (26%), Positives = 57/119 (47%), Gaps = 2/119 (1%)
Frame = -3
Query: 644 LWAYRTAYKTPIGMTPFKLVYGKSCHLPVELEHKAYWAIRNLNLDPNLAGDKRKLQLNEL 703
LW++RT + TPF + + P E+ + R + + L D+ L +
Sbjct: 462 LWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSR-MPQNIELNNDRLFNALETI 286
Query: 704 EELRMDAYENARIYKERTKTWHDKKIIKRHFKSGDLVL--LFNSRLKLFPGKLRSRWSG 760
EE R A + Y+ + +++++K + + K GD+VL +F + +L GKL + W G
Sbjct: 285 EERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNWEG 109
>BG587166 similar to GP|8778801|gb| T32E20.9 {Arabidopsis thaliana}, partial
(2%)
Length = 756
Score = 67.0 bits (162), Expect = 3e-11
Identities = 34/62 (54%), Positives = 42/62 (66%)
Frame = +1
Query: 749 LFPGKLRSRWSGPFQVRTVYPYGAIEIFSEETGSFTVNGQRLKIYNTGEVNEVVADFTLS 808
LFPGKL+SR SGPF+V+ V PYGAI ++S + FTVNGQR+K+Y E LS
Sbjct: 1 LFPGKLKSR*SGPFKVKEVRPYGAIVLWSTDGRDFTVNGQRVKLYMATAPEEDGVSTPLS 180
Query: 809 DP 810
DP
Sbjct: 181 DP 186
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 58.2 bits (139), Expect = 1e-08
Identities = 29/102 (28%), Positives = 55/102 (53%), Gaps = 1/102 (0%)
Frame = -2
Query: 282 KELLAVVYAIDKFRQYLVGSKIIVYTDHSAIKYLLNKKDAKPRLIRWILLLQEFDLEIKD 341
K A+ +A + R Y++ + + IKY+ K R+ RW +LL E+D+E +
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 342 KKGVE-NVVADHLSRLRETNKDELPLDDSFPDDQLFLLAQTD 382
+K ++ +++ADHL+ + +D P+ FPD+++ L D
Sbjct: 455 QKAIKGSILADHLA--HQPLEDYRPIKFDFPDEEIMYLKMKD 336
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 52.8 bits (125), Expect = 5e-07
Identities = 27/61 (44%), Positives = 38/61 (62%), Gaps = 1/61 (1%)
Frame = +1
Query: 243 ASDYAVGAVLGQRNDKKM-HAIYYASKTLDGAQVNYATTEKELLAVVYAIDKFRQYLVGS 301
A D+ +GA L Q++++ H I YAS+ L A+ NY E+E LA ++AI FR YL G
Sbjct: 1 APDWGIGAGLSQKDEENHEHPIAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYLHGP 180
Query: 302 K 302
K
Sbjct: 181 K 183
>BG582085
Length = 824
Score = 51.2 bits (121), Expect = 2e-06
Identities = 27/72 (37%), Positives = 46/72 (63%), Gaps = 2/72 (2%)
Frame = -3
Query: 323 PRLIRWILLLQEFDLEIKDKK--GVENVVADHLSRLRETNKDELPLDDSFPDDQLFLLAQ 380
P +R ILL QEFDLE + ++ G+ ++A R ++ ++E+ + + FP++QLF ++Q
Sbjct: 771 P*RLRCILLCQEFDLESRQERETGMW*LIAFPCLRNKQVTQNEISIKERFPNEQLFAISQ 592
Query: 381 TDAPWYADFVNF 392
PW+AD NF
Sbjct: 591 --RPWFADMTNF 562
Score = 29.6 bits (65), Expect = 4.8
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = -1
Query: 436 CIPENEVSSILTHCHSSSYGGHAS 459
C+ E + IL H HSSSY GH S
Sbjct: 530 CVGEKVIKDIL*HHHSSSYMGHHS 459
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 49.7 bits (117), Expect = 5e-06
Identities = 27/88 (30%), Positives = 43/88 (48%)
Frame = +2
Query: 579 DGGSHFISRHFEKLLQKLGVRHKIATPYHPQTSGQVEVSNRQIKAILEKTVSTSRTDWSN 638
+G S S +++L + G +++ YHP + GQ E N+ + L + T WS
Sbjct: 11 EG*SSLYSNFWKQLFKLHGTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSK 190
Query: 639 KLDDALWAYRTAYKTPIGMTPFKLVYGK 666
A + Y T+Y MTPFK +YG+
Sbjct: 191 AFPWAEYWYNTSYNISAAMTPFKALYGR 274
>TC87803 homologue to GP|1871187|gb|AAB63547.1| unknown protein {Arabidopsis
thaliana}, partial (0%)
Length = 840
Score = 29.6 bits (65), Expect = 4.8
Identities = 15/35 (42%), Positives = 18/35 (50%)
Frame = +2
Query: 464 SFKILHSGFWWPSLFKDVHLFISKCDKCQRTGSIT 498
S K S W SLF V ISKC C +T +I+
Sbjct: 110 SLKKFDSSIWCRSLFISVESRISKCSPCYKTMNIS 214
>BQ751193 similar to GP|21622336|em probable multidrug resistance protein 2
{Neurospora crassa}, partial (33%)
Length = 797
Score = 29.6 bits (65), Expect = 4.8
Identities = 19/67 (28%), Positives = 30/67 (44%), Gaps = 10/67 (14%)
Frame = -1
Query: 677 KAYWAIRNLNLDPN---------LAGDKRKLQLNELEELRMDAYENARIYKERTKTWHDK 727
K+ WA+R LNL+PN L + N L + R++ + + H++
Sbjct: 659 KSDWALRRLNLEPNRL*NTGLPVLQFKEAACSANALHQFRVEGRKAEKTGSRERSVHHER 480
Query: 728 -KIIKRH 733
KI KRH
Sbjct: 479 CKISKRH 459
>TC81537 similar to GP|7630037|emb|CAB88331.1 putative protein {Arabidopsis
thaliana}, partial (34%)
Length = 751
Score = 28.9 bits (63), Expect = 8.2
Identities = 17/73 (23%), Positives = 34/73 (46%)
Frame = +1
Query: 697 KLQLNELEELRMDAYENARIYKERTKTWHDKKIIKRHFKSGDLVLLFNSRLKLFPGKLRS 756
+ LNE EE+ + + E+ ++ + KKI + + + + P +L +
Sbjct: 337 RTSLNEGEEVPLSSPESIEAFRMLNPRYRKKKIEEMGVTEEEYLAKQFEIKEDVPEELET 516
Query: 757 RWSGPFQVRTVYP 769
+W+GP V+ V P
Sbjct: 517 KWAGPLVVKLVPP 555
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.140 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,815,737
Number of Sequences: 36976
Number of extensions: 400604
Number of successful extensions: 1808
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 1795
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1803
length of query: 811
length of database: 9,014,727
effective HSP length: 104
effective length of query: 707
effective length of database: 5,169,223
effective search space: 3654640661
effective search space used: 3654640661
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146705.12


