

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
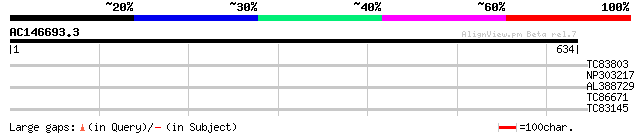
Query= AC146693.3 - phase: 0 /pseudo
(634 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC83803 weakly similar to GP|14209566|dbj|BAB56062. putative rec... 32 0.97
NP303217 NP303217|AJ297721.1|CAC20726.1 putative repetitive prol... 32 0.97
AL388729 PIR|S42643|S42 ubiquitin / ribosomal protein S27a - pot... 30 2.2
TC86671 similar to PIR|T06782|T06782 extensin - soybean, partial... 29 4.8
TC83145 weakly similar to GP|16604509|gb|AAL24260.1 AT5g58730/mz... 28 8.2
>TC83803 weakly similar to GP|14209566|dbj|BAB56062. putative
receptor-protein kinase {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 754
Score = 31.6 bits (70), Expect = 0.97
Identities = 19/45 (42%), Positives = 25/45 (55%), Gaps = 2/45 (4%)
Frame = +2
Query: 312 SPRTEKPNRISERLSPNSLPRLILS--PHTPKCLRPKSPKWPNKW 354
S RT K + I+++L ++ LIL P TP CLRP P P W
Sbjct: 113 STRTNKKHEINDKL*NHNHHPLILFHLPLTPSCLRPN*PF*PQLW 247
>NP303217 NP303217|AJ297721.1|CAC20726.1 putative repetitive proline-rich
protein
Length = 525
Score = 31.6 bits (70), Expect = 0.97
Identities = 26/73 (35%), Positives = 33/73 (44%), Gaps = 4/73 (5%)
Frame = +1
Query: 311 KSPRTEKPNRISERL--SPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVN- 367
K P T KP + + SPN P L L+P PK P+K PP K P+N
Sbjct: 175 KPPPTYKPPIKKQPINKSPNKKPLL---------LKPPLPKPPHKEPPSRKR----PINT 315
Query: 368 -PKQTPKLMSTPF 379
P + P L+ PF
Sbjct: 316 PPDKKPPLLKPPF 354
>AL388729 PIR|S42643|S42 ubiquitin / ribosomal protein S27a - potato
(fragment), partial (33%)
Length = 425
Score = 30.4 bits (67), Expect = 2.2
Identities = 33/114 (28%), Positives = 48/114 (41%), Gaps = 1/114 (0%)
Frame = +1
Query: 317 KPNRISERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFP-VNPKQTPKLM 375
KP +++ S + RLIL P T W + WP LL + P VNP +P
Sbjct: 100 KPRSKTKKESHRTKQRLILLPGTTS--------WED-WPALLA*RTTIPEVNPTLSP--- 243
Query: 376 STPFLMGVVSWKGQLLKPKELRERVLGFLVRKMRNRLIRTRPLTH*GLLSSTWR 429
++ V W G+ K + +ER +K +PL H*G + WR
Sbjct: 244 ---LVLPVFRWVGRRKKDGK-KERT*QLKAQKKE-----FKPLRH*GRVEKLWR 378
>TC86671 similar to PIR|T06782|T06782 extensin - soybean, partial (41%)
Length = 991
Score = 29.3 bits (64), Expect = 4.8
Identities = 26/78 (33%), Positives = 34/78 (43%), Gaps = 8/78 (10%)
Frame = +3
Query: 309 K*KSPRTEKPNRISERLSPNSLPRLILSPHTP----KCLRPKSPKWPNKW----PPLLKH 360
K KSP P + + SP + SP P K L P PK P K+ PP+ K+
Sbjct: 225 KYKSPPPPLPKKPYKYPSPPPPIYMYKSPPPPIYKYKSLPPPPPKKPYKYPSPPPPVYKY 404
Query: 361 *ESFPVNPKQTPKLMSTP 378
+PK+T K S P
Sbjct: 405 KSPPSPSPKKTYKYPSPP 458
>TC83145 weakly similar to GP|16604509|gb|AAL24260.1 AT5g58730/mzn1_180
{Arabidopsis thaliana}, partial (33%)
Length = 904
Score = 28.5 bits (62), Expect = 8.2
Identities = 18/48 (37%), Positives = 23/48 (47%), Gaps = 2/48 (4%)
Frame = +1
Query: 327 PNSLPRLILSPHTPKCLRPKSPKWPN--KWPPLLKH*ESFPVNPKQTP 372
P S+P L LS + K + K P W PLLK S P + + TP
Sbjct: 295 PFSIPSLSLSISSLKWAQISPTKQPTFLSWSPLLKQPSSMPTSDQITP 438
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.364 0.164 0.631
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,161,546
Number of Sequences: 36976
Number of extensions: 364381
Number of successful extensions: 5010
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1517
Number of HSP's successfully gapped in prelim test: 207
Number of HSP's that attempted gapping in prelim test: 3437
Number of HSP's gapped (non-prelim): 1877
length of query: 634
length of database: 9,014,727
effective HSP length: 102
effective length of query: 532
effective length of database: 5,243,175
effective search space: 2789369100
effective search space used: 2789369100
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146693.3


