

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
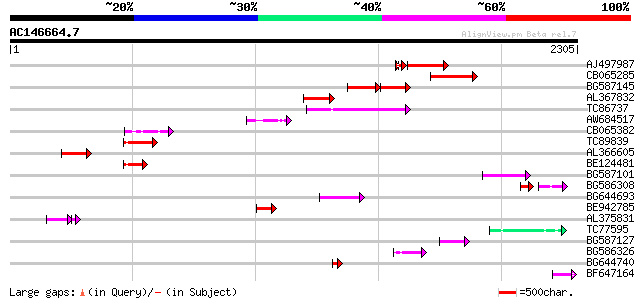
Query= AC146664.7 - phase: 0
(2305 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 334 3e-91
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 291 2e-78
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 147 9e-56
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 214 4e-55
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 189 8e-48
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 149 9e-36
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 146 8e-35
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 142 1e-33
AL366605 134 5e-31
BE124481 115 1e-25
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 113 7e-25
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 62 1e-18
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 91 7e-18
BE942785 82 2e-15
AL375831 67 6e-15
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 71 6e-12
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 70 9e-12
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 54 5e-07
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 50 1e-05
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 50 1e-05
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 334 bits (856), Expect = 3e-91
Identities = 151/167 (90%), Positives = 163/167 (97%)
Frame = -2
Query: 1616 KTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRT 1675
KTCCALAWAAKRLRHYMINHTTWL+SKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYR+
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1676 QKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPESEWGLIF 1735
QKA+KGSILA+HLAHQP+EDY+PIKFDFPD+E+MYLK KDCDEP+FGEGPDP+S WGLIF
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 1736 DGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEACIMGI 1782
DGAVNVYG+GIGAVL+TPKG HIPFTARLRFDCTNNI EYEACIMGI
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
Score = 47.4 bits (111), Expect(2) = 1e-05
Identities = 21/31 (67%), Positives = 24/31 (76%)
Frame = +2
Query: 1582 CVLGQQDETGKKEHVIYYLSKKFTDCESRYS 1612
CVLGQQDETG+KEH IYYLS+K + R S
Sbjct: 38 CVLGQQDETGRKEHAIYYLSRKLSSTVWRIS 130
Score = 21.9 bits (45), Expect(2) = 1e-05
Identities = 12/13 (92%), Positives = 12/13 (92%)
Frame = +1
Query: 1569 LIMYLTVLEDSMG 1581
LIMYLTVLE SMG
Sbjct: 1 LIMYLTVLE-SMG 36
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 291 bits (746), Expect = 2e-78
Identities = 137/196 (69%), Positives = 161/196 (81%), Gaps = 8/196 (4%)
Frame = -2
Query: 1712 KAKDCDEPVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNN 1771
K+KDC+EP+ GEGPDP S+WGL+FDGAVN YG GIGAV+++P+G +IPFTAR+ F+CTNN
Sbjct: 591 KSKDCEEPLIGEGPDPNSKWGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNN 412
Query: 1772 IVEYEACIMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFF 1831
+ EYEACI GIEEAID+RIK + IYGDSALVINQIKGEWET H LIPYRDYARRLLT+F
Sbjct: 411 MAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYF 232
Query: 1832 NKVELHHVPRDENQMADALATLSSMINVNGHNTVPVINVQFLDRPAYVF--------VAE 1883
KVELHH+PRDENQMADALATLSSM VN N VP+I VQ L+RP++VF E
Sbjct: 231 TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGNVIDQTGE 52
Query: 1884 AIDDDKPWYHDIQVFL 1899
+ D KPWY+DI+ FL
Sbjct: 51 NVVDYKPWYYDIKQFL 4
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 147 bits (372), Expect(2) = 9e-56
Identities = 69/136 (50%), Positives = 100/136 (72%)
Frame = +2
Query: 1373 MAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVK 1432
M P+D EKT+FIT G +CY VMPFGL NAG+TYQR + ++F D + +EVY+DDM+VK
Sbjct: 2 MHPDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVK 181
Query: 1433 SGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIR 1492
S +H+ +L + F+ L +Y ++LNP KCTFGV SG+ LG+IV+Q+GIEV+P ++ AI
Sbjct: 182 SLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAIL 361
Query: 1493 EMPVPKTEKQVRGFLG 1508
++P PK ++V+ G
Sbjct: 362 DLPSPKNSREVQRLTG 409
Score = 90.1 bits (222), Expect(2) = 9e-56
Identities = 45/120 (37%), Positives = 74/120 (61%)
Frame = +3
Query: 1508 GRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDDCQKAFDQIKEYLLEPPILVPPVDGR 1567
GR+ ++RFIS T C P +KLL ++ W++ C++AF+Q+K+YL PP+L P G
Sbjct: 408 GRIAALNRFISRSTDKCLPFYKLLCGNKRFVWDEKCEEAFEQLKQYLTTPPVLSKPEAGD 587
Query: 1568 PLIMYLTVLEDSMGCVLGQQDETGKKEHVIYYLSKKFTDCESRYSVLEKTCCALAWAAKR 1627
L +Y+ + ++ VL ++D +K I+Y SK+ TD E+RY LEK A+ +A++
Sbjct: 588 TLSLYIAISSTAVSSVLIREDRGEQKP--IFYTSKRMTDPETRYPTLEKMAFAVITSARK 761
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 214 bits (544), Expect = 4e-55
Identities = 96/128 (75%), Positives = 118/128 (92%)
Frame = -1
Query: 1194 QERKTIQPHQEEIEIINLGTEEDKKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDMP 1253
QERK IQPHQEEIE+INLGTEE+K+EIK+GA+L+ +K+++ +L+REY+DIFA SY+DMP
Sbjct: 384 QERKAIQPHQEEIELINLGTEENKREIKVGAALEEGVKRKIFQLLREYLDIFACSYEDMP 205
Query: 1254 GLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANI 1313
GLDP +VEHR+P KPECPPV+ KLRR+HPDMALKIK EV+KQIDAGFL+T EYP+W+ANI
Sbjct: 204 GLDPKIVEHRIPTKPECPPVR*KLRRTHPDMALKIKSEVQKQIDAGFLMTVEYPEWVANI 25
Query: 1314 VPVPKKDG 1321
VPVPKKDG
Sbjct: 24 VPVPKKDG 1
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 189 bits (481), Expect = 8e-48
Identities = 129/428 (30%), Positives = 220/428 (51%), Gaps = 7/428 (1%)
Frame = +1
Query: 1207 EIINLGTEEDKKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDMPGLDPDVVEHRLPL 1266
+++++ E+ +K + A +D K ++ E+ D+F +++H +PL
Sbjct: 256 QVMSVSLEDIEKALAPKAPVDPR-KHVPAGVLEEFPDLFNPEKAYQVPASRGLLDHAIPL 432
Query: 1267 KPEC----PPVKQ-KLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANIVPVPKKDG 1321
P+ PP+ L L +K+ + +D GF+ S A ++ V K G
Sbjct: 433 IPDKDGNDPPLPWGPLYGMSRQELLVLKKTLEDLLDKGFIKASGSAAG-APVLFVRKPGG 609
Query: 1322 KIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREKT 1381
IR CVDYR LN + KD +PLP I + A + F+ +D + ++++R+ ED+EKT
Sbjct: 610 GIRFCVDYRALNAITKKDRYPLPLISETLRRVAGARWFTKLDVVAAFHKMRIKDEDQEKT 789
Query: 1382 SFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIV-KSGTEEEHV 1440
+F T +G F ++V PFGL A AT+QR + K H+ + + Y+DD+++ +G++++H
Sbjct: 790 AFRTRYGLFEWIVCPFGLTGAPATFQRYINKTLHEFLDDFVTAYIDDVLIYTTGSKKDHE 969
Query: 1441 EYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQ-KGIEVDPDKVRAIREMPVPKT 1499
+ ++ +RL L L+P KC F V + K +GFI++ KG+ DP K+ AIR+ P +
Sbjct: 970 AQVRRVLRRLADAGLSLDPKKCEFSVTTVKYVGFILTAGKGVSCDPLKLAAIRDWLPPGS 1149
Query: 1500 EKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDDCQKAFDQIKEYLLEPPI 1559
K R FLG NY FI + P+ +L RKD +W + + AF ++K E P+
Sbjct: 1150 VKGARSFLGFCNYYKDFIPGYSEITEPLTRLTRKDFPFRWGAEQEAAFTKLKRLFAEEPV 1329
Query: 1560 LVPPVDGRPLIMYLTVLEDSMGCVLGQQDETGKKEHVIYYLSKKFTDCESRYSVLEKTCC 1619
L + ++G VL Q+D TG H + + S++ + E Y + +K
Sbjct: 1330 LRMFDPEAVTTVETDCSGFALGGVLTQEDGTG-AAHPVAFHSQRLSPAEYNYPIHDKELL 1506
Query: 1620 ALAWAAKR 1627
A+ WA R
Sbjct: 1507 AV-WACLR 1527
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana chromosome
II BAC F26H6; putative retroelement pol polyprotein,
partial (1%)
Length = 488
Score = 149 bits (377), Expect = 9e-36
Identities = 85/186 (45%), Positives = 111/186 (58%), Gaps = 2/186 (1%)
Frame = +1
Query: 963 VTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLKFVSRGKLITVSGESAFLISNLSAF 1022
+TFQVM+I AS+SCLLGRPWIHDAGAVTSTLHQKLKFV G
Sbjct: 1 ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKFVKNG------------------- 123
Query: 1023 SVIGGSGSDGPSFQGFSAE-ESVGKIETCMASLKDARRVIQEGKTEGWGQLVELPENKRK 1081
++G +FQG S E K+ MASLKDA++ +QEG+ WG+L++L ENKRK
Sbjct: 124 ------SAEGTAFQGLSMEGAEPKKVGAAMASLKDAQKAVQEGQAADWGKLIQLCENKRK 285
Query: 1082 EGIGFLNSKPGMFDPTRGSFHSAGFIHDSPETNAILDDAPGGV-TPVFVTPGGACCNWIA 1140
EG+ F + + G+FHSAGF+ N + ++ V P+FV PGG +W A
Sbjct: 286 EGLRFSPTS----GVSTGTFHSAGFV------NTLAEEVARFVPRPLFVIPGGIAKDWDA 435
Query: 1141 VDIPSV 1146
VD+PS+
Sbjct: 436 VDVPSI 453
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 146 bits (369), Expect = 8e-35
Identities = 92/203 (45%), Positives = 118/203 (57%), Gaps = 4/203 (1%)
Frame = -2
Query: 468 QPYNPQQLSQPPYYPQQP-YQQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALLPSLLAQ 526
+P P L PP P +QQR QQ RP P +PM Y LP+LL +
Sbjct: 569 RPMIP*LLC*PPRNNHPPVHQQRQ*QQARPTFPL-----------IPMLYAE*LPTLLLR 423
Query: 527 N---LVQTIPPPRIPDSLPRWYRPDLHCIYHQGAPGHDVERCFALKKEVQKLINSKELTF 583
+ Q PPP D LP +R DL C +HQGA GHDVE C+ALK V+KLIN +LTF
Sbjct: 422 GHCTIRQGKPPP---DPLPPRFRSDLKCDFHQGALGHDVEGCYALKHIVKKLINQGKLTF 252
Query: 584 TDPDAVAQNNPLPTHGPAVNMIQDDQEEARILSVGDIKTPLVPIHVKMCKATLFNHNHEA 643
+ +NPLP H AVNMI + EEA L V ++ TPLVP+H+K+C+A+LF+H+H
Sbjct: 251 ENNVPHVLDNPLPNHA-AVNMI-EVYEEAPGLDVRNVTTPLVPLHIKLCQASLFDHDHAN 78
Query: 644 CDICLMDPRGCIQVQNDMQGLLN 666
C +P GC VQND+Q L+N
Sbjct: 77 CQE*FYNPLGCCVVQNDIQSLMN 9
Score = 32.7 bits (73), Expect = 1.7
Identities = 15/43 (34%), Positives = 27/43 (61%)
Frame = -1
Query: 6 QENAQLREELVNLKGEMERMALMMETMMAEREQATISNSTPVV 48
+ENAQLR EL +++ E+ + M ++A +EQ++ + T V
Sbjct: 624 EENAQLRTELASMREELAKANDTMTALLAAQEQSSTGSPTTTV 496
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 142 bits (359), Expect = 1e-33
Identities = 73/148 (49%), Positives = 95/148 (63%), Gaps = 9/148 (6%)
Frame = +3
Query: 463 PPLYQQ-PYNPQQLSQPPYYPQQPYQQRPQQQ------PRPQVPHNQQYQRQQFDPLPMT 515
PPL QQ P +P Q P PQ+P QQ P+Q P+ +P NQ + Q DP+P+
Sbjct: 228 PPLVQQRPQSPYQ----PMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVK 395
Query: 516 YGALLPSLLAQNLVQTIPPPRIPDSLPRWYRPDLHCIYHQGAPGHDVERCFALKKEVQKL 575
Y LLP LL +NL+QT+P PR+P+SLP WYRPDL+C++HQGAPGHD E+C+ LK+EVQKL
Sbjct: 396 YADLLPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHDTEQCYPLKEEVQKL 575
Query: 576 INSKELTFTDPD--AVAQNNPLPTHGPA 601
I + +F D D + Q L H A
Sbjct: 576 IENNVWSFDDQDIKVLLQQQHLAPHSGA 659
>AL366605
Length = 422
Score = 134 bits (336), Expect = 5e-31
Identities = 65/124 (52%), Positives = 88/124 (70%), Gaps = 2/124 (1%)
Frame = -3
Query: 211 DLCLVPDVNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGP 270
DLCLV V++P KFK+P F++Y G TCPQNH+ Y+R+M YK+N L+IHCFQDSL
Sbjct: 372 DLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMED 193
Query: 271 AHTWFMGL--KGVTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQ 328
A W+ L + TF++LA AF Y +NT L P+R+ L+SL+QK +ESF+EYAQR+
Sbjct: 192 AAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRG 13
Query: 329 KAAQ 332
AA+
Sbjct: 12 AAAR 1
>BE124481
Length = 538
Score = 115 bits (289), Expect = 1e-25
Identities = 56/106 (52%), Positives = 72/106 (67%), Gaps = 7/106 (6%)
Frame = +3
Query: 463 PPLYQQ-PYNPQQLSQPPYYPQQPYQQRPQQQ------PRPQVPHNQQYQRQQFDPLPMT 515
PPL QQ P +P Q P PQ+P QQ P+Q P+ +P NQ + Q DP+P+
Sbjct: 231 PPLVQQRPQSPYQ----PMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVK 398
Query: 516 YGALLPSLLAQNLVQTIPPPRIPDSLPRWYRPDLHCIYHQGAPGHD 561
Y LLP LL +NL+QT+P PR+P+SLP WYRPDL+C++HQGAPGHD
Sbjct: 399 YADLLPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHD 536
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 113 bits (283), Expect = 7e-25
Identities = 60/196 (30%), Positives = 100/196 (50%)
Frame = +2
Query: 1922 RFFLNEDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHAMAKKILRAGYYWI 1981
R+ +E LYK+ D + +RCV + E ++ H ++ H KI +AG++W
Sbjct: 17 RYLWDEPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGFWWP 196
Query: 1982 TMHADCYNHAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMIGRIEPKASNGHRF 2041
TM D ++ +C CQ + N I F +WGID +G +N ++
Sbjct: 197 TMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPSSYNN--KY 370
Query: 2042 ILVAIDYFTKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDNGTNLNNNMMKELC 2101
ILVA+DY +KWVEA + VVV+ K+ + R+GVP +I+D G++ N + ++L
Sbjct: 371 ILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLL 550
Query: 2102 DDFKIQHHNSSPYRPQ 2117
++H ++ Y PQ
Sbjct: 551 KKNGVRHKVATAYHPQ 598
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 61.6 bits (148), Expect(2) = 1e-18
Identities = 42/122 (34%), Positives = 68/122 (55%), Gaps = 4/122 (3%)
Frame = -3
Query: 2151 LYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIPSL---RVLMEAELSEAEWCQSRYDQL 2207
L+ +RT+ R +T +TPFS+ + +EA+ P EV + SL R+ EL+ ++ L
Sbjct: 462 LWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNIELNN----DRLFNAL 295
Query: 2208 NLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEGDLVL-KCIKSFQPDPRGKWTPNY 2266
IEE+R AL Q YQ +++ ++K V + K GD+VL K ++ + GK N+
Sbjct: 294 ETIEERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNW 115
Query: 2267 EG 2268
EG
Sbjct: 114 EG 109
Score = 52.0 bits (123), Expect(2) = 1e-18
Identities = 22/52 (42%), Positives = 36/52 (68%)
Frame = -2
Query: 2078 YGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI 2129
+G+P I+TDNG++ +N +E C+ ++I+ + +SP PQ NG EA+NK I
Sbjct: 685 HGLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKII 530
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 90.5 bits (223), Expect = 7e-18
Identities = 57/184 (30%), Positives = 95/184 (50%)
Frame = +2
Query: 1260 VEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANIVPVPKK 1319
++ + L P P+ R +P +K +++ ++ GF+ S YP + ++ + KK
Sbjct: 35 IDFGIDLLPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKK 211
Query: 1320 DGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDRE 1379
DG +RM +DY LN + K +PLP ID L DN K F +D G +Q R+ ED
Sbjct: 212 DGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQHRVIGEDVP 391
Query: 1380 KTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEH 1439
KT+F +G + +VM FG N + M ++F D + + V+ +D+++ S E EH
Sbjct: 392 KTAFRIRYGHYEILVMSFG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDILIYSKNENEH 571
Query: 1440 VEYL 1443
+L
Sbjct: 572 ENHL 583
>BE942785
Length = 460
Score = 82.4 bits (202), Expect = 2e-15
Identities = 40/81 (49%), Positives = 57/81 (69%), Gaps = 1/81 (1%)
Frame = -2
Query: 1003 GKLITVSGESAFLISNLSAFSVIGGSGSDGPSFQGFSAEESVGKIE-TCMASLKDARRVI 1061
G L+T+ GE A+LIS LS+FS I ++G +FQG + E + K + T MASLKDA+R +
Sbjct: 375 GXLVTIHGEEAYLISQLSSFSCIEAGSAEGTAFQGLTVEGTEPKRDGTAMASLKDAQRAV 196
Query: 1062 QEGKTEGWGQLVELPENKRKE 1082
QE + GWG+L++L ENK K+
Sbjct: 195 QESQAAGWGRLIQLRENKHKD 133
>AL375831
Length = 467
Score = 67.4 bits (163), Expect(2) = 6e-15
Identities = 40/108 (37%), Positives = 55/108 (50%), Gaps = 1/108 (0%)
Frame = +2
Query: 148 IPQATMTQAGPTVHVGPQHEE-QIYHSDSIMGDDKAIDWEERFGALEKKMSNMRGKETVV 206
+PQ T P VH PQ H + EE L ++ RG V
Sbjct: 44 LPQTTAAVTEPLVHTLPQGININTQHRSIPVTKTMEEMMEELAKELRHEIQANRGNADSV 223
Query: 207 QSIYDLCLVPDVNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKN 254
++ DLCLV V++P KFK+P F++Y G TCPQNH+ Y+R+M YK+
Sbjct: 224 KT-QDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 364
Score = 33.5 bits (75), Expect(2) = 6e-15
Identities = 16/36 (44%), Positives = 21/36 (57%), Gaps = 2/36 (5%)
Frame = +3
Query: 253 KNNVPLLIHCFQDSLTGPAHTWFMGL--KGVTTFEQ 286
K N L+IHCFQDSL A W+ L + TF++
Sbjct: 360 KTNDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDE 467
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 70.9 bits (172), Expect = 6e-12
Identities = 72/320 (22%), Positives = 129/320 (39%), Gaps = 8/320 (2%)
Frame = +2
Query: 1950 KLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPS 2009
KL+ E H+ + H G +I+ ++W + C C IHI
Sbjct: 176 KLVQESHDSTAAGHP-GRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVC----GGIHIWRQ 340
Query: 2010 MLNVISSPWPF-----SMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQ 2064
P P S +D I + P G +++ V +D +K V + +
Sbjct: 341 AKRGFLKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAE 520
Query: 2065 VVVRFIKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEA 2124
+ + +G+P I++D G+N +E C + S+ Y PQ +G E
Sbjct: 521 ACAQRFLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTER 700
Query: 2125 ANKNIKKIIQKMVVTYKD-WHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEI 2183
N+ I+ +++ V +D W ++LP R +S GATPF + +G V P+
Sbjct: 701 WNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYH-VDPIPTVE 877
Query: 2184 PSLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPRE-FK 2242
+ V+ E E + + D I+ + +AA Q R + + +K P + ++
Sbjct: 878 DTGGVVSEGEAAAQLLVKRMKDVTGFIQAEIVAA-------QQRSEASANKRRCPADRYQ 1036
Query: 2243 EGDLVLKCIKSFQ-PDPRGK 2261
GD V + +++ P P K
Sbjct: 1037 VGDKVWLNVSNYKSPRPSKK 1096
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 70.1 bits (170), Expect = 9e-12
Identities = 43/121 (35%), Positives = 60/121 (49%)
Frame = +3
Query: 1747 GAVLITPKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRIKKIVIYGDSALVINQI 1806
G L +P + + RL F +NN YEA I G+ A L+I+ I Y DS LV +Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1807 KGEWETRHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADALATLSSMINVNGHNTVP 1866
GE+E R + Y ++L + L +PR EN ADALA L+S + +P
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALASSSDPELKRVIP 362
Query: 1867 V 1867
V
Sbjct: 363 V 365
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 54.3 bits (129), Expect = 5e-07
Identities = 41/140 (29%), Positives = 67/140 (47%), Gaps = 3/140 (2%)
Frame = +2
Query: 1558 PILVPPVDGRPLIMYLTVLEDS---MGCVLGQQDETGKKEHVIYYLSKKFTDCESRYSVL 1614
PILV P LI Y+ + S +GCVL Q E VI Y S++ E Y
Sbjct: 17 PILVLP----ELITYVVYTDASITGLGCVLTQH------EKVIAYASRQLRKHEGNYPTH 166
Query: 1615 EKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYR 1674
+ A+ +A K R Y+ + + +KYIF +P L R RW +++YD++
Sbjct: 167 DLEMAAVVFALKIWRSYLYGAKVQIHTDHKSLKYIFTQPELNLRQRRWMEFVADYDLDI- 343
Query: 1675 TQKAVKGSILAEHLAHQPIE 1694
T K +++A+ L+ + ++
Sbjct: 344 TYYPGKANLVADALSRRRVD 403
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 49.7 bits (117), Expect = 1e-05
Identities = 22/41 (53%), Positives = 28/41 (67%)
Frame = -1
Query: 1311 ANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVD 1351
A ++ V KKDG RMC+DYR NK + K+ +PLP ID L D
Sbjct: 238 AALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFD 116
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 49.7 bits (117), Expect = 1e-05
Identities = 33/99 (33%), Positives = 53/99 (53%), Gaps = 1/99 (1%)
Frame = +1
Query: 2206 QLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEGDLVLKCIKSFQPDPRGKWTPN 2265
+LN +EE R+ A + ++Y+ R K+ D+ + REF+EG+LVL + P GK +
Sbjct: 103 ELNELEELRLDAYENAKIYKERTKKWHDRRIIRREFREGELVLLFNSRLKLFP-GKLRSH 279
Query: 2266 YEGPYVVKRAFSGGAL-ILTNMDGEELPRPVNSDAVKKY 2303
+ GP+ VK GA+ + + G P VN +K Y
Sbjct: 280 WSGPFQVKNVMPSGAVEVWSESTG---PFTVNGQRLKHY 387
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 71,196,195
Number of Sequences: 36976
Number of extensions: 1087927
Number of successful extensions: 8352
Number of sequences better than 10.0: 254
Number of HSP's better than 10.0 without gapping: 6109
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7322
length of query: 2305
length of database: 9,014,727
effective HSP length: 112
effective length of query: 2193
effective length of database: 4,873,415
effective search space: 10687399095
effective search space used: 10687399095
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146664.7


