

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146651.7 - phase: 0 /pseudo
(394 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
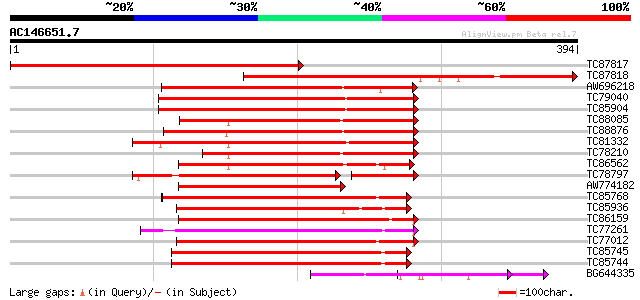
Score E
Sequences producing significant alignments: (bits) Value
TC87817 homologue to GP|20161224|dbj|BAB90151. katanin p60 subun... 399 e-111
TC87818 similar to GP|20197082|gb|AAC26698.2 putative katanin {A... 291 2e-79
AW696218 homologue to PIR|B96835|B96 CAD ATPase (AAA1) 35570-33... 230 8e-61
TC79040 homologue to PIR|F84674|F84674 probable AAA-type ATPase ... 221 5e-58
TC85904 similar to PIR|F84674|F84674 probable AAA-type ATPase [i... 219 1e-57
TC88085 similar to GP|20259341|gb|AAM13995.1 unknown protein {Ar... 170 1e-42
TC88876 similar to PIR|G96537|G96537 hypothetical protein F2J10.... 167 6e-42
TC81332 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22... 166 1e-41
TC78210 similar to GP|6056413|gb|AAF02877.1| Unknown protein {Ar... 159 2e-39
TC86562 similar to GP|21553404|gb|AAM62497.1 26S proteasome regu... 154 7e-38
TC78797 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22... 125 1e-36
AW774182 homologue to PIR|F84674|F84 probable AAA-type ATPase [i... 147 7e-36
TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle pr... 143 1e-34
TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidops... 128 3e-30
TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AA... 125 3e-29
TC77261 homologue to GP|8777330|dbj|BAA96920.1 26S proteasome AA... 122 2e-28
TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulator... 119 3e-27
TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome re... 116 2e-26
TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15... 115 4e-26
BG644335 similar to GP|18461184|db putative katanin {Oryza sativ... 110 9e-25
>TC87817 homologue to GP|20161224|dbj|BAB90151. katanin p60 subunit A 1-like
{Oryza sativa (japonica cultivar-group)}, partial (25%)
Length = 768
Score = 399 bits (1024), Expect = e-111
Identities = 195/204 (95%), Positives = 197/204 (95%)
Frame = +2
Query: 1 MAADDEPMPTRWSFEEFKKYYDVRLGRKKLVENGENAVSNGNSSGIASNGNSHGKVTSDR 60
MAADDEPMPTRWSFEEFKKYYDVRLGRKKLVENGENAVSNGNSSGIASNGNSHGKVTSDR
Sbjct: 134 MAADDEPMPTRWSFEEFKKYYDVRLGRKKLVENGENAVSNGNSSGIASNGNSHGKVTSDR 313
Query: 61 AIYDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVK 120
AIYDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVK
Sbjct: 314 AIYDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVK 493
Query: 121 WESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECN 180
WESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECN
Sbjct: 494 WESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECN 673
Query: 181 TTFFNISASSIVSKWRGDSEKLVK 204
TTFFNI + KWR DS+KLVK
Sbjct: 674 TTFFNIFSIVYCHKWRSDSKKLVK 745
Score = 29.6 bits (65), Expect = 2.1
Identities = 19/50 (38%), Positives = 24/50 (48%)
Frame = +3
Query: 162 GPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELAR 211
GP G + L + + F SASSIV+ + KVLFELAR
Sbjct: 618 GPQGLERQCLQRQLQLSATLHFLIFSASSIVTNGEVIQKN**KVLFELAR 767
>TC87818 similar to GP|20197082|gb|AAC26698.2 putative katanin {Arabidopsis
thaliana}, partial (62%)
Length = 984
Score = 291 bits (746), Expect = 2e-79
Identities = 170/242 (70%), Positives = 179/242 (73%), Gaps = 10/242 (4%)
Frame = +2
Query: 163 PPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDE 222
P G + MLAKAVATECNTTFFNISASS+ SKWRGDSEKLVKVLFELARHHAPATIFLDE
Sbjct: 5 PQGLERPMLAKAVATECNTTFFNISASSMFSKWRGDSEKLVKVLFELARHHAPATIFLDE 184
Query: 223 IDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLE 282
IDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLE
Sbjct: 185 IDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLE 364
Query: 283 KR--NQKQEEQCLRNSF--LCSLMRNRCPMIY-----WWTGQKDTQARIFAYFVKKQQCS 333
KR E + R F L L + PM Y G + R+ Q
Sbjct: 365 KRILVPLPEPEARRAMFEELLPLQPDEEPMPYDLLVDRTEGYSGSDIRLLCKETAMQPLR 544
Query: 334 H*DA**HNLNKNQMWCP-KKLPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQ 392
L + P ++LPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQ
Sbjct: 545 RLMT---QLEQEPDVVPEEELPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQ 715
Query: 393 IL 394
IL
Sbjct: 716 IL 721
>AW696218 homologue to PIR|B96835|B96 CAD ATPase (AAA1) 35570-33019
[imported] - Arabidopsis thaliana, partial (34%)
Length = 556
Score = 230 bits (586), Expect = 8e-61
Identities = 107/185 (57%), Positives = 148/185 (79%), Gaps = 7/185 (3%)
Frame = +3
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPG 165
E L RD++ SP V+W+ + GL AKRLL+EAVV+P+ P+YF G+ PWKG+L+FGPPG
Sbjct: 3 EMLERDVLETSPGVRWDDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPG 182
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
TGKT+LAKAVATEC TTFFN+S++++ SKWRG+SE++V+ LF+LAR +AP+TIF+DEID+
Sbjct: 183 TGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDS 362
Query: 226 IISQRGEGRSEHEASRRLKTELLIQMDGLAR-------TDELVFVLAATNLPWELDAAML 278
+ + RG EHE+SRR+K+ELL+Q+DG++ + +LV VLAATN PW++D A+
Sbjct: 363 LCNARG-ASGEHESSRRVKSELLVQVDGVSNSATNEDGSRKLVMVLAATNFPWDIDEALR 539
Query: 279 RRLEK 283
RRLEK
Sbjct: 540 RRLEK 554
>TC79040 homologue to PIR|F84674|F84674 probable AAA-type ATPase [imported]
- Arabidopsis thaliana, partial (74%)
Length = 1193
Score = 221 bits (562), Expect = 5e-58
Identities = 107/181 (59%), Positives = 140/181 (77%)
Frame = +3
Query: 104 LAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGP 163
L L IIR PNVKW + GLE+AK+ L+EAV++P+K+P++FTG PW+ LL+GP
Sbjct: 6 LRAGLISAIIREKPNVKWNDVAGLESAKQALQEAVILPVKFPQFFTGKRRPWRAFLLYGP 185
Query: 164 PGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEI 223
PGTGK+ LAKAVATE ++TFF+IS+S +VSKW G+SEKLV LF++AR AP+ IF+DEI
Sbjct: 186 PGTGKSYLAKAVATEADSTFFSISSSDLVSKWMGESEKLVSNLFQMARESAPSIIFVDEI 365
Query: 224 DAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEK 283
D++ QRGEG +E EASRR+KTELL+QM G+ D+ V VLAATN P+ LD A+ RR +K
Sbjct: 366 DSLCGQRGEG-NESEASRRIKTELLVQMQGVGNNDQKVLVLAATNTPYALDQAIRRRFDK 542
Query: 284 R 284
R
Sbjct: 543 R 545
>TC85904 similar to PIR|F84674|F84674 probable AAA-type ATPase [imported] -
Arabidopsis thaliana, complete
Length = 1793
Score = 219 bits (559), Expect = 1e-57
Identities = 105/181 (58%), Positives = 140/181 (77%)
Frame = +2
Query: 104 LAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGP 163
L L+ I+R PNVKW + GLE+AK+ L+EAV++P+K+P++FTG PW+ LL+GP
Sbjct: 524 LRAGLNSAIVREKPNVKWNDVAGLESAKQSLQEAVILPVKFPQFFTGKRRPWRAFLLYGP 703
Query: 164 PGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEI 223
PGTGK+ LAKAVATE ++TFF++S+S +VSKW G+SEKLV LFE+AR AP+ IF+DEI
Sbjct: 704 PGTGKSYLAKAVATEADSTFFSVSSSDLVSKWMGESEKLVSNLFEMARESAPSIIFVDEI 883
Query: 224 DAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEK 283
D++ RGEG +E EASRR+KTELL+QM G+ D+ V VLAATN P+ LD A+ RR +K
Sbjct: 884 DSLCGTRGEG-NESEASRRIKTELLVQMQGVGHNDQKVLVLAATNTPYALDQAIRRRFDK 1060
Query: 284 R 284
R
Sbjct: 1061R 1063
>TC88085 similar to GP|20259341|gb|AAM13995.1 unknown protein {Arabidopsis
thaliana}, partial (29%)
Length = 1261
Score = 170 bits (430), Expect = 1e-42
Identities = 92/169 (54%), Positives = 116/169 (68%), Gaps = 3/169 (1%)
Frame = +1
Query: 119 VKWESIKGLENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFGPPGTGKTMLAKAVA 176
V + I LEN K LKE V++P++ P+ F L P KGILLFGPPGTGKTMLAKAVA
Sbjct: 754 VSFNDIGALENVKDTLKELVMLPLQRPELFCKGQLTKPCKGILLFGPPGTGKTMLAKAVA 933
Query: 177 TECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSE 236
TE F NIS SSI SKW G+ EK VK +F LA AP+ IF+DE+D+++ +R E E
Sbjct: 934 TEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKIAPSVIFVDEVDSMLGRR-ENPGE 1110
Query: 237 HEASRRLKTELLIQMDGLARTD-ELVFVLAATNLPWELDAAMLRRLEKR 284
HEA R++K E ++ DGL D E V VLAATN P++LD A++RRL +R
Sbjct: 1111HEAMRKMKNEFMVNWDGLRTKDRERVLVLAATNRPFDLDEAVIRRLPRR 1257
>TC88876 similar to PIR|G96537|G96537 hypothetical protein F2J10.1 [imported]
- Arabidopsis thaliana, partial (72%)
Length = 1572
Score = 167 bits (423), Expect = 6e-42
Identities = 84/180 (46%), Positives = 125/180 (68%), Gaps = 3/180 (1%)
Frame = +2
Query: 108 LSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILLFGPPG 165
+S + G VK++ I LE+ K+ L+E V++P++ P+ F+ LL P KGILLFGPPG
Sbjct: 878 VSAVVAPGEIGVKFDDIGALEDVKKALQELVILPMRRPELFSHGNLLRPCKGILLFGPPG 1057
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
TGKT+LAKA+ATE F +++ S++ SKW GD+EKL K LF A AP IF+DE+D+
Sbjct: 1058 TGKTLLAKALATEAGANFISVTGSTLTSKWFGDAEKLTKALFSFASKLAPVIIFVDEVDS 1237
Query: 226 IISQRGEGRSEHEASRRLKTELLIQMDGL-ARTDELVFVLAATNLPWELDAAMLRRLEKR 284
++ RG G EHEA+RR++ E + DGL ++ ++ + +L ATN P++LD A++RRL +R
Sbjct: 1238 LLGARG-GAHEHEATRRMRNEFMAAWDGLRSKENQRILILGATNRPFDLDDAVIRRLPRR 1414
>TC81332 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22
{Arabidopsis thaliana}, partial (39%)
Length = 1514
Score = 166 bits (420), Expect = 1e-41
Identities = 92/209 (44%), Positives = 135/209 (64%), Gaps = 10/209 (4%)
Frame = +1
Query: 86 EKPKKSLLPPFESAEMRT------LAESLSRDIIRGSP-NVKWESIKGLENAKRLLKEAV 138
EKPK+S A ++ E + +++I + V + I L++ K L+EAV
Sbjct: 709 EKPKESKKDGDIKASAKSDSPDNAFEECIRQELIPANEIKVTFSDIGALDDVKESLQEAV 888
Query: 139 VMPIKYPKYFTG--LLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWR 196
++P++ P F G +L P KG+LLFGPPGTGKTMLAKA+A E +F N+S S+I S W+
Sbjct: 889 MLPLRRPDLFKGDGVLKPCKGVLLFGPPGTGKTMLAKAIANEAGASFINVSPSTITSMWQ 1068
Query: 197 GDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDG-LA 255
G SEK V+ LF LA AP IF+DE+D+++ QR R EH + RR+K E + + DG L+
Sbjct: 1069GQSEKNVRALFSLAAKVAPTIIFIDEVDSMLGQRSSTR-EHSSMRRVKNEFMSRWDGLLS 1245
Query: 256 RTDELVFVLAATNLPWELDAAMLRRLEKR 284
+ DE + VLAATN+P++LD A++RR ++R
Sbjct: 1246KPDEKITVLAATNMPFDLDEAIIRRFQRR 1332
>TC78210 similar to GP|6056413|gb|AAF02877.1| Unknown protein {Arabidopsis
thaliana}, partial (23%)
Length = 1254
Score = 159 bits (402), Expect = 2e-39
Identities = 84/153 (54%), Positives = 108/153 (69%), Gaps = 3/153 (1%)
Frame = +3
Query: 135 KEAVVMPIKYPKYFTG--LLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIV 192
KE V++P+K P+ F L P KGILLFGPPGTGKTMLAKAVATE F NIS SSI
Sbjct: 3 KELVMLPLKRPELFCKGQLTKPCKGILLFGPPGTGKTMLAKAVATEAGANFINISMSSIT 182
Query: 193 SKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMD 252
SKW G+ EK VK +F LA AP+ IF+DE+D+++ +R E EHEA R++K E ++ D
Sbjct: 183 SKWFGEGEKYVKAVFSLASKIAPSVIFVDEVDSMLGRR-ENPGEHEAMRKMKNEFMVNWD 359
Query: 253 GL-ARTDELVFVLAATNLPWELDAAMLRRLEKR 284
GL + E + VLAATN P++LD A++RRL +R
Sbjct: 360 GLRTKEKERILVLAATNRPFDLDEAVIRRLPRR 458
>TC86562 similar to GP|21553404|gb|AAM62497.1 26S proteasome regulatory
particle chain RPT6-like protein {Arabidopsis thaliana},
partial (77%)
Length = 1564
Score = 154 bits (388), Expect = 7e-38
Identities = 85/168 (50%), Positives = 115/168 (67%), Gaps = 4/168 (2%)
Frame = +1
Query: 118 NVKWESIKGLENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFGPPGTGKTMLAKAV 175
+V+++SI GLE K+ L E V++P++ P F+ LL P KG+LL+GPPGTGKTMLAKA+
Sbjct: 394 DVEFDSIGGLETIKQTLFELVILPLQRPDLFSHGKLLGPQKGVLLYGPPGTGKTMLAKAI 573
Query: 176 ATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRS 235
A E F N+ S+++SKW GD++KLV +F LA P+ IF+DE+D+ + QR S
Sbjct: 574 AKESGAVFINVRISNLMSKWFGDAQKLVAAVFSLAHKLQPSIIFIDEVDSFLGQRRS--S 747
Query: 236 EHEASRRLKTELLIQMDGLARTDE--LVFVLAATNLPWELDAAMLRRL 281
+HEA +KTE + DG A TD+ V VLAATN P ELD A+LRRL
Sbjct: 748 DHEAVLNMKTEFMALWDGFA-TDQSARVMVLAATNRPSELDEAILRRL 888
>TC78797 similar to GP|15810167|gb|AAL06985.1 At1g64110/F22C12_22 {Arabidopsis
thaliana}, partial (74%)
Length = 2270
Score = 125 bits (315), Expect(2) = 1e-36
Identities = 71/150 (47%), Positives = 94/150 (62%), Gaps = 5/150 (3%)
Frame = +1
Query: 86 EKP----KKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMP 141
EKP K S +PP E R E + + I +V + I LE K L+E V++P
Sbjct: 1606 EKPVAASKASEVPPDNEFEKRIRPEVIPANEI----DVTFSDIGALEETKESLQELVMLP 1773
Query: 142 IKYPKYFTG-LLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSE 200
++ P FTG LL P +GILLFGPPGTGKTMLAKA+A E +F N+S S+I SKW G+ E
Sbjct: 1774 LRRPDLFTGGLLKPCRGILLFGPPGTGKTMLAKAIAKEAGASFINVSMSTITSKWFGEDE 1953
Query: 201 KLVKVLFELARHHAPATIFLDEIDAIISQR 230
K V+ LF LA +P IF+DE+D+++ QR
Sbjct: 1954 KNVRALFTLASKVSPTIIFVDEVDSMLGQR 2043
Score = 45.1 bits (105), Expect(2) = 1e-36
Identities = 22/48 (45%), Positives = 33/48 (67%), Gaps = 1/48 (2%)
Frame = +3
Query: 238 EASRRLKTELLIQMDGLA-RTDELVFVLAATNLPWELDAAMLRRLEKR 284
+A R++K E + DGL + E + VLAATN P++LD A++RR E+R
Sbjct: 2064 KAMRKIKNEFMSHWDGLTTKQGERILVLAATNKPFDLDEAIIRRFERR 2207
>AW774182 homologue to PIR|F84674|F84 probable AAA-type ATPase [imported] -
Arabidopsis thaliana, partial (27%)
Length = 487
Score = 147 bits (371), Expect = 7e-36
Identities = 66/116 (56%), Positives = 92/116 (78%)
Frame = -1
Query: 118 NVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVAT 177
NVKW + GL++AK+ L+ AV++P+KY ++ TG PW+ LL+GPPGTGK+ LAKAVAT
Sbjct: 364 NVKWNDVAGLKSAKQSLQ*AVILPVKYRQFLTGKRRPWRAFLLYGPPGTGKSYLAKAVAT 185
Query: 178 ECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEG 233
+ ++TFF++S+S +VSKW G+SEKLV LFE+AR AP+ IF+DEID++ RGEG
Sbjct: 184 KADSTFFSVSSSDLVSKWMGESEKLVSNLFEMARESAPSIIFVDEIDSLCGTRGEG 17
>TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle protein 48
homolog (Valosin containing protein homolog) (VCP).
[Soybean], partial (95%)
Length = 2785
Score = 143 bits (360), Expect = 1e-34
Identities = 74/175 (42%), Positives = 111/175 (63%), Gaps = 2/175 (1%)
Frame = +2
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S R+ + PN W+ I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 1490 SALRETVVEVPNCSWDDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 1669
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
GKT+LAKA+A EC F +I +++ W G+SE V+ +F+ AR AP +F DE+D+
Sbjct: 1670 CGKTLLAKAIANECQANFISIKGPELLTMWFGESEANVREIFDKARGSAPCVLFFDELDS 1849
Query: 226 IISQRGEGRSE-HEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
I +QRG + A+ R+ +LL +MDG++ + VF++ ATN P +D A+LR
Sbjct: 1850 IATQRGSSVGDAGGAADRVLNQLLTEMDGMS-AKKTVFIIGATNRPDIIDPALLR 2011
Score = 118 bits (295), Expect = 5e-27
Identities = 67/175 (38%), Positives = 107/175 (60%), Gaps = 1/175 (0%)
Frame = +2
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 164
E + R+ V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 668 EPIKREDENRLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 847
Query: 165 GTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEID 224
G+GKT++A+AVA E FF I+ I+SK G+SE ++ FE A +AP+ IF+DEID
Sbjct: 848 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 1027
Query: 225 AIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+I +R ++ E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R
Sbjct: 1028SIAPKR--EKTHGEVERRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRR 1183
>TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidopsis
thaliana}, partial (98%)
Length = 1633
Score = 128 bits (322), Expect = 3e-30
Identities = 73/167 (43%), Positives = 109/167 (64%), Gaps = 4/167 (2%)
Frame = +1
Query: 117 PNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAV 175
P+ ++ I GL+ + +KE + +PIK+P+ F L ++ KG+LL+GPPGTGKT+LA+AV
Sbjct: 607 PDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLLARAV 786
Query: 176 ATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQR---GE 232
A + TF +S S +V K+ G+ ++V+ LF +AR HAP+ IF+DEID+I S R G
Sbjct: 787 AHHTDCTFIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSARMESGS 966
Query: 233 GRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
G + E R + ELL Q+DG +++ + VL ATN LD A+LR
Sbjct: 967 GNGDSEVQRTM-LELLNQLDGFEASNK-IKVLMATNRIDILDQALLR 1101
>TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AAA-ATPase
subunit RPT4a-like protein {Arabidopsis thaliana},
complete
Length = 1640
Score = 125 bits (314), Expect = 3e-29
Identities = 68/171 (39%), Positives = 111/171 (64%), Gaps = 4/171 (2%)
Frame = +2
Query: 118 NVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAVA 176
N+ + ++ GL + R L+E++ +P+ P+ F + + P KG+LL+GPPGTGKT+LA+A+A
Sbjct: 473 NISYSAVGGLSDQIRELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLARAIA 652
Query: 177 TECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQR-GEGRS 235
+ + F + +S+I+ K+ G+S +L++ +F AR H P IF+DEIDAI +R EG S
Sbjct: 653 SNIDANFLKVVSSAIIDKYIGESARLIREMFGYARDHQPCIIFMDEIDAIGGRRFSEGTS 832
Query: 236 EHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR--RLEKR 284
+R ELL Q+DG + ++ ++ ATN P LD A+LR RL+++
Sbjct: 833 ADREIQRTLMELLNQLDGFDQLGKVKMIM-ATNRPDVLDPALLRPGRLDRK 982
>TC77261 homologue to GP|8777330|dbj|BAA96920.1 26S proteasome AAA-ATPase
subunit RPT3 {Arabidopsis thaliana}, partial (94%)
Length = 1575
Score = 122 bits (306), Expect = 2e-28
Identities = 74/197 (37%), Positives = 119/197 (59%), Gaps = 4/197 (2%)
Frame = +2
Query: 92 LLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL 151
+LPP + + L++S P+V + I G + K+ ++EAV +P+ + + + +
Sbjct: 506 VLPPEADSSISLLSQS-------EKPDVTYNDIGGCDIQKQEIREAVELPLTHHELYKQI 664
Query: 152 -LSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELA 210
+ P +G+LL+GPPGTGKTMLAKAVA F + S V K+ G+ ++V+ +F LA
Sbjct: 665 GIDPPRGVLLYGPPGTGKTMLAKAVANHTTAAFIRVVGSEFVQKYLGEGPRMVRDVFRLA 844
Query: 211 RHHAPATIFLDEIDAIISQRGEGRSEHEAS-RRLKTELLIQMDGLARTDELVFVLAATNL 269
+ +APA IF+DE+DAI + R + ++ + +R+ ELL QMDG +T V V+ ATN
Sbjct: 845 KENAPAIIFIDEVDAIATARFDAQTGADREVQRILMELLNQMDGFDQTVN-VKVIMATNR 1021
Query: 270 PWELDAAMLR--RLEKR 284
LD A+LR RL+++
Sbjct: 1022ADTLDPALLRPGRLDRK 1072
>TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulatory subunit
6A homolog (TAT-binding protein homolog 1) (TBP-1)
(Mg(2+), complete
Length = 1793
Score = 119 bits (297), Expect = 3e-27
Identities = 70/172 (40%), Positives = 108/172 (62%), Gaps = 4/172 (2%)
Frame = +1
Query: 117 PNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAV 175
P + I GLE + L EA+V+P+ + + F L + P KG+LL+GPPGTGKT++A+A
Sbjct: 625 PTEDYNDIGGLEKQIQELVEAIVLPMTHKERFQKLGIRPPKGVLLYGPPGTGKTLMARAC 804
Query: 176 ATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGR- 234
A + N TF ++ +V + GD KLV+ F+LA+ +P IF+DEIDAI ++R +
Sbjct: 805 AAQTNATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDAIGTKRFDSEV 984
Query: 235 SEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR--RLEKR 284
S +R ELL Q+DG + +D+ + V+AATN LD A++R RL+++
Sbjct: 985 SGDREVQRTMLELLNQLDGFS-SDDRIKVIAATNRADILDPALMRSGRLDRK 1137
>TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome regulatory
particle triple-A ATPase subunit2b {Oryza sativa
(japonica cultivar-group), partial (98%)
Length = 1696
Score = 116 bits (290), Expect = 2e-26
Identities = 69/170 (40%), Positives = 105/170 (61%), Gaps = 3/170 (1%)
Frame = +3
Query: 113 IRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTML 171
+ +P + I GL+ + +KEAV +P+ +P+ + + + P KG++L+G PGTGKT+L
Sbjct: 624 VEKAPLESYADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLL 803
Query: 172 AKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRG 231
AKAVA + TF + S ++ K+ GD KLV+ LF +A +PA +F+DEIDA+ ++R
Sbjct: 804 AKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPAIVFIDEIDAVGTKRY 983
Query: 232 EGRSEHEAS-RRLKTELLIQMDGL-ARTDELVFVLAATNLPWELDAAMLR 279
+ S E +R ELL Q+DG +R D V V+ ATN LD A+LR
Sbjct: 984 DAHSGGEREIQRTMLELLNQLDGFDSRGD--VKVILATNRIESLDPALLR 1127
>TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15.70 -
Arabidopsis thaliana, complete
Length = 1629
Score = 115 bits (287), Expect = 4e-26
Identities = 68/170 (40%), Positives = 105/170 (61%), Gaps = 3/170 (1%)
Frame = +2
Query: 113 IRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTML 171
+ +P + I GL+ + +KEAV +P+ +P+ + + + P KG++L+G PGTGKT+L
Sbjct: 626 VEKAPLESYADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLL 805
Query: 172 AKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRG 231
AKAVA + TF + S ++ K+ GD KLV+ LF +A +P+ +F+DEIDA+ ++R
Sbjct: 806 AKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAVGTKRY 985
Query: 232 EGRSEHEAS-RRLKTELLIQMDGL-ARTDELVFVLAATNLPWELDAAMLR 279
+ S E +R ELL Q+DG +R D V V+ ATN LD A+LR
Sbjct: 986 DAHSGGEREIQRTMLELLNQLDGFDSRGD--VKVILATNRIESLDPALLR 1129
>BG644335 similar to GP|18461184|db putative katanin {Oryza sativa (japonica
cultivar-group)}, partial (41%)
Length = 566
Score = 110 bits (275), Expect = 9e-25
Identities = 68/148 (45%), Positives = 82/148 (54%), Gaps = 8/148 (5%)
Frame = -1
Query: 210 ARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNL 269
ARHHAP+TIFLDEIDAIISQRGE RSEHE+SRRLKTE L DG +D + + N
Sbjct: 566 ARHHAPSTIFLDEIDAIISQRGEARSEHESSRRLKTE-LAHTDGRFESDR*TCLCSGGNK 390
Query: 270 P----WELDAAMLRRLEK----RNQKQEEQCLRNSFLCSLMRNRCPMIYWWTGQKDTQAR 321
W + + R R QKQ CLRN + L R MIYW QK
Sbjct: 389 SFPGNWMQQCSDVLRSGSLCPFRRQKQGAGCLRNYYHPFLNRRHFHMIYW*KRQKAFPVL 210
Query: 322 IFAYFVKKQQCSH*DA**HNLNKNQMWC 349
IF Y+ ++ C+H D * NL + + WC
Sbjct: 209 IFGYYARRLPCNHYDV*WQNLIRGKNWC 126
Score = 50.1 bits (118), Expect = 2e-06
Identities = 41/115 (35%), Positives = 55/115 (47%), Gaps = 10/115 (8%)
Frame = -3
Query: 270 PWELDAAMLRRLEKRN----QKQEEQCLRNSFLCSLMRNRCPMIYWWTGQKD-----TQA 320
PWELDAAMLRRLEKR + E +C L + + P+ Y +K +
Sbjct: 384 PWELDAAMLRRLEKRILVPLPEAEARCGMFEELLPSLPEQAPLPYDLLVEKTKGFSGSDI 205
Query: 321 RIFAYFVKKQQCSH*DA**HNLNKNQMWCPK-KLPKVGPVVPEDVEAALRNTRPS 374
R+ Q A L+K + P+ +LP VGP+ D+E AL NTRPS
Sbjct: 204 RLLCKEAAMQPLRCLMA---ELDKREELVPEDELPNVGPITVTDIEMALMNTRPS 49
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,770,258
Number of Sequences: 36976
Number of extensions: 156051
Number of successful extensions: 914
Number of sequences better than 10.0: 99
Number of HSP's better than 10.0 without gapping: 873
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 881
length of query: 394
length of database: 9,014,727
effective HSP length: 98
effective length of query: 296
effective length of database: 5,391,079
effective search space: 1595759384
effective search space used: 1595759384
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146651.7


