

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
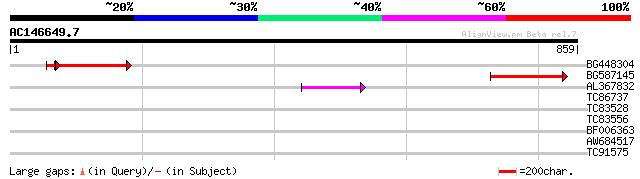
Query= AC146649.7 - phase: 0 /pseudo
(859 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG448304 160 9e-41
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 115 5e-26
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 49 1e-05
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 36 0.055
TC83528 35 0.094
TC83556 33 0.61
BF006363 similar to GP|23496060|gb hypothetical protein {Plasmod... 32 0.80
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 32 1.4
TC91575 similar to GP|20073183|gb|AAH27195.1 Similar to xylosylp... 29 6.7
>BG448304
Length = 637
Score = 160 bits (404), Expect(2) = 9e-41
Identities = 75/117 (64%), Positives = 88/117 (75%)
Frame = +2
Query: 68 KDAGVKPPKAHSQEKRRVDKTKYCRFYKCHGHVTDHCIHLKDAIKMLIQ*GRLN*FTKNS 127
K G PPK +QE + DKTKYCRF+KCH H+TD CIHLKD I++LIQ GRLN TKN
Sbjct: 44 KKLGSNPPKTPTQENKGTDKTKYCRFHKCHRHLTDDCIHLKDTIEILIQRGRLNQLTKNP 223
Query: 128 ETERQAVKLKTDGKEKNPAVAMSVEQLDTFLEHMDTTPYACSWEHFPDANIIVGGTS 184
E E+Q VKL TD K K+ VAMSVEQLD F+EH+D TPY+C+WE FP AN+I GG S
Sbjct: 224 EPEKQTVKLITDKKNKDIVVAMSVEQLDEFVEHVDITPYSCTWEPFPTANVITGGIS 394
Score = 26.2 bits (56), Expect(2) = 9e-41
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = +1
Query: 56 EKILVEISVAELKDAGVKPPK 76
+KIL I++ +LK+ G+KP K
Sbjct: 7 KKILA*IAITDLKEVGIKPAK 69
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 115 bits (289), Expect = 5e-26
Identities = 54/117 (46%), Positives = 80/117 (68%)
Frame = +3
Query: 729 GMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVLSRPDEGE 788
G + +L RF+++S LPF+ LL +F W +CE+A + LK L+ PPVLS+P+ G+
Sbjct: 408 GRIAALNRFISRSTDKCLPFYKLLCGNKRFVWDEKCEEAFEQLKQYLTTPPVLSKPEAGD 587
Query: 789 TLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALVTAAR 845
TL LY+++SS AVS L E + QKP ++TSK + PETRY +EK+A A++T+AR
Sbjct: 588 TLSLYIAISSTAVSSVLIREDRGEQKPIFYTSKRMTDPETRYPTLEKMAFAVITSAR 758
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 48.5 bits (114), Expect = 1e-05
Identities = 25/96 (26%), Positives = 46/96 (47%)
Frame = -1
Query: 443 KLGKGIPEHARAHLIACLWLNTDLFARSITDMPDIDPSVACHQLTVSPSASVVAQRRKKQ 502
K+G + E + + L D+FA S DMP +DP + H++ P V + ++
Sbjct: 303 KVGAALEEGVKRKIFQLLREYLDIFACSYEDMPGLDPKIVEHRIPTKPECPPVR*KLRRT 124
Query: 503 FPEKAEGTEKFVKDLAEEKFIAEAKYTTWLSNILLV 538
P+ A + V+ + F+ +Y W++NI+ V
Sbjct: 123 HPDMALKIKSEVQKQIDAGFLMTVEYPEWVANIVPV 16
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 36.2 bits (82), Expect = 0.055
Identities = 31/110 (28%), Positives = 42/110 (38%), Gaps = 1/110 (0%)
Frame = +1
Query: 737 FVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVLSRPDEGETLYLYLSV 796
F+ ++ P L +K+ F W E E A LK + PVL D +
Sbjct: 1198 FIPGYSEITEPLTRLTRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPEAVTTVETDC 1377
Query: 797 SSEAVSGTLNIETQEG-QKPTYFTSKALLGPETRYQKIEKVALALVTAAR 845
S A+ G L E G P F S+ L E Y +K LA+ R
Sbjct: 1378 SGFALGGVLTQEDGTGAAHPVAFHSQRLSPAEYNYPIHDKELLAVWACLR 1527
>TC83528
Length = 555
Score = 35.4 bits (80), Expect = 0.094
Identities = 19/50 (38%), Positives = 26/50 (52%), Gaps = 3/50 (6%)
Frame = +3
Query: 304 GDGNRTKTIKIPFLVIEV---TFLYNCIIGMTGLVQLGAVC*TAHLKLKY 350
G G TKTIK+ + V+E +YN ++G L L AV A +KY
Sbjct: 390 GKGENTKTIKVRYFVVESPPSVSIYNVVLGWPSLKDLKAVLSVAEFTIKY 539
>TC83556
Length = 842
Score = 32.7 bits (73), Expect = 0.61
Identities = 25/86 (29%), Positives = 44/86 (51%), Gaps = 4/86 (4%)
Frame = +1
Query: 125 KNSETERQAVKLKTD-GKEKNPAVAMSVEQ---LDTFLEHMDTTPYACSWEHFPDANIIV 180
K +++ + A K K++ K+K V +S+ + + FL M+ + AC I+
Sbjct: 427 KATDSNKAAAKEKSEESKDKGLQVPISITKQ*DFNIFLAIMEMSNNACI--------ILD 582
Query: 181 GGTSTLSLGSMKRKFEELLSVNHLVH 206
GG ++G KRKFEE++ + VH
Sbjct: 583 GGLCHKNMGFNKRKFEEMMCIT*WVH 660
>BF006363 similar to GP|23496060|gb hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 583
Score = 32.3 bits (72), Expect = 0.80
Identities = 14/36 (38%), Positives = 23/36 (63%), Gaps = 3/36 (8%)
Frame = -1
Query: 80 QEKRRVDKTKYCRFYKCHGHVTDHCIH---LKDAIK 112
+E D +KYCRF++C G+ T+ C +K++IK
Sbjct: 526 KETSWTDISKYCRFHRCTGYDTNDCFQ*EAIKESIK 419
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 488
Score = 31.6 bits (70), Expect = 1.4
Identities = 14/37 (37%), Positives = 22/37 (58%)
Frame = +1
Query: 314 IPFLVIEVTFLYNCIIGMTGLVQLGAVC*TAHLKLKY 350
I F V+++ Y+C++G + GAV T H KLK+
Sbjct: 1 ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKF 111
>TC91575 similar to GP|20073183|gb|AAH27195.1 Similar to xylosylprotein
beta1 4-galactosyltransferase polypeptide 7
(galactosyltransferase I), partial (4%)
Length = 722
Score = 29.3 bits (64), Expect = 6.7
Identities = 13/40 (32%), Positives = 20/40 (49%)
Frame = -3
Query: 398 YSGVQPDRSRSTLHKVIVERAEEGKQGPTQRGNTSPFGDD 437
YSG+ P+ +KVI +E+G GPT+ + D
Sbjct: 414 YSGIDPNLKEINPNKVIPTLSEQGLAGPTEASQVGSYSLD 295
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.335 0.146 0.469
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,736,323
Number of Sequences: 36976
Number of extensions: 379646
Number of successful extensions: 2679
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1482
Number of HSP's successfully gapped in prelim test: 104
Number of HSP's that attempted gapping in prelim test: 1178
Number of HSP's gapped (non-prelim): 1623
length of query: 859
length of database: 9,014,727
effective HSP length: 104
effective length of query: 755
effective length of database: 5,169,223
effective search space: 3902763365
effective search space used: 3902763365
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146649.7


