

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146632.13 - phase: 0
(126 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
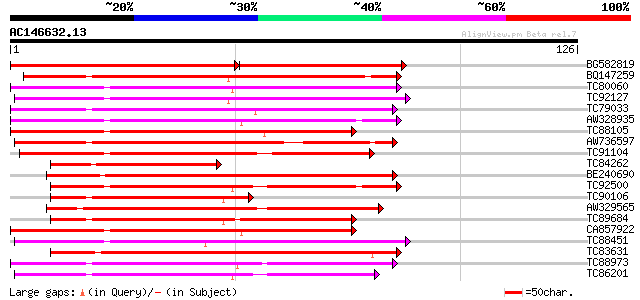
BG582819 similar to GP|17979045|gb unknown protein {Arabidopsis ... 82 8e-22
BQ147259 similar to PIR|C96654|C96 hypothetical protein F16P17.1... 79 4e-16
TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.... 64 1e-11
TC92127 weakly similar to GP|16040952|dbj|BAB69683. receptor kin... 64 2e-11
TC79033 similar to GP|21554229|gb|AAM63304.1 somatic embryogenes... 62 6e-11
AW328935 similar to PIR|T01711|T01 probable serine/threonine-spe... 62 8e-11
TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein ki... 60 3e-10
AW736597 similar to GP|12054894|e receptor protein kinase {Pinus... 59 4e-10
TC91104 similar to GP|9758799|dbj|BAB09252.1 serine/threonine pr... 58 8e-10
TC84262 similar to GP|15528705|dbj|BAB64771. putative receptor p... 57 1e-09
BE240690 similar to PIR|T49986|T49 lectin-like protein kinase-li... 57 2e-09
TC92500 similar to GP|14194119|gb|AAK56254.1 At1g01540/F22L4_6 {... 57 2e-09
TC90106 similar to GP|3641252|gb|AAC36318.1| leucine-rich recept... 57 2e-09
AW329565 homologue to GP|19347728|gb putative receptor protein k... 56 3e-09
TC89684 similar to PIR|T10659|T10659 probable serine/threonine-s... 55 5e-09
CA857922 similar to PIR|T47793|T47 receptor-like protein kinase ... 55 7e-09
TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein ki... 55 9e-09
TC83631 similar to PIR|T10659|T10659 probable serine/threonine-s... 55 9e-09
TC88973 similar to PIR|T50817|T50817 protein serine/threonine ki... 54 1e-08
TC86201 similar to PIR|T49003|T49003 protein kinase-like protein... 54 1e-08
>BG582819 similar to GP|17979045|gb unknown protein {Arabidopsis thaliana},
partial (19%)
Length = 753
Score = 82.4 bits (202), Expect(2) = 8e-22
Identities = 35/51 (68%), Positives = 45/51 (87%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGED 51
+S++ QSALGY+APE AC+ +++ EKCDVYGFGV++LE VTGKRPVEY ED
Sbjct: 244 LSSKIQSALGYMAPEFACKTVKITEKCDVYGFGVLVLETVTGKRPVEYMED 396
Score = 36.2 bits (82), Expect(2) = 8e-22
Identities = 15/37 (40%), Positives = 24/37 (64%)
Frame = +2
Query: 52 NVLILNDHVRVLLEHGNALECVDPSLMSEYPEDEVLP 88
+V++L D VR L+ G EC+D L ++P +EV+P
Sbjct: 398 DVVVLCDMVRGALDEGRVEECIDERLQGKFPVEEVIP 508
>BQ147259 similar to PIR|C96654|C96 hypothetical protein F16P17.10 [imported]
- Arabidopsis thaliana, partial (16%)
Length = 662
Score = 79.3 bits (194), Expect = 4e-16
Identities = 43/85 (50%), Positives = 62/85 (72%), Gaps = 1/85 (1%)
Frame = +2
Query: 4 RFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE-YGEDNVLILNDHVRV 62
+F +A+GYVAPELA Q R +EKCDVY FGV++LE+VTG++PVE V++L ++VR
Sbjct: 86 KFHNAVGYVAPELA-QSFRQSEKCDVYSFGVILLELVTGRKPVESVTAHEVVVLCEYVRS 262
Query: 63 LLEHGNALECVDPSLMSEYPEDEVL 87
LLE G+A C D +L + E+E++
Sbjct: 263 LLETGSASNCFDRNLQG-FVENELI 334
>TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.24
[imported] - Arabidopsis thaliana, partial (32%)
Length = 1547
Score = 64.3 bits (155), Expect = 1e-11
Identities = 37/88 (42%), Positives = 52/88 (59%), Gaps = 1/88 (1%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
+S R +GY+APE A + + +K DVY FGV+ LEIV+G Y ++ + L D
Sbjct: 667 ISTRIAGTIGYMAPEYAMRGY-LTDKADVYSFGVVALEIVSGMSNTNYRPKEEFVYLLDW 843
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E GN LE VDP+L S+Y +E +
Sbjct: 844 AYVLQEQGNLLELVDPTLGSKYSSEEAM 927
>TC92127 weakly similar to GP|16040952|dbj|BAB69683. receptor kinase 5
{Brassica rapa}, partial (18%)
Length = 507
Score = 63.9 bits (154), Expect = 2e-11
Identities = 38/89 (42%), Positives = 51/89 (56%), Gaps = 1/89 (1%)
Frame = +3
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE-YGEDNVLILNDHV 60
+NR GY++PE A + + + K DVY FGV++LEI+ G++ Y ED L L H
Sbjct: 162 TNRIVGTYGYMSPEYAMEGV-FSTKSDVYSFGVLLLEIINGEKNNSFYCEDRPLNLVGHA 338
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLPC 89
L + G LE VDP L + EDEVL C
Sbjct: 339 WELWKEGVVLELVDPLLNESFSEDEVLRC 425
>TC79033 similar to GP|21554229|gb|AAM63304.1 somatic embryogenesis
receptor-like kinase putative {Arabidopsis thaliana},
partial (77%)
Length = 1762
Score = 62.0 bits (149), Expect = 6e-11
Identities = 33/87 (37%), Positives = 51/87 (57%), Gaps = 1/87 (1%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNV-LILNDH 59
++ R + LGY+APE A + + NE CDVY FG+++LE+ +GK+P+E +V +ND
Sbjct: 937 VTTRVKGTLGYLAPEYA-MLGKANESCDVYSFGILLLELASGKKPLEKLSSSVKRAINDW 1113
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEV 86
L E DP L +Y E+E+
Sbjct: 1114 ALPLACEKKFSELADPRLNGDYVEEEL 1194
>AW328935 similar to PIR|T01711|T01 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) A_IG002N01.22 - Arabidopsis
thaliana, partial (30%)
Length = 572
Score = 61.6 bits (148), Expect = 8e-11
Identities = 33/88 (37%), Positives = 53/88 (59%), Gaps = 1/88 (1%)
Frame = +3
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGE-DNVLILNDH 59
++ R GYVAPE AC + + EK DVY FG++I+E++TG+ PV+YG + L +
Sbjct: 60 VTTRVMGTFGYVAPEYACTGM-LTEKSDVYSFGILIMELITGRSPVDYGRPQGEVNLIEW 236
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
++ ++ + A + VDP L E P + L
Sbjct: 237 LKTMVGNRKAEDVVDPKL-PELPSSKAL 317
>TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein kinase
{Arabidopsis thaliana}, partial (72%)
Length = 2379
Score = 59.7 bits (143), Expect = 3e-10
Identities = 32/78 (41%), Positives = 50/78 (64%), Gaps = 1/78 (1%)
Frame = +2
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLI-LNDH 59
++ R GYVAPE A L +NEK DVY FGV++LE +TG+ PV+Y + L D
Sbjct: 1199 ITTRVMGTFGYVAPEYANSGL-LNEKSDVYSFGVLLLEAITGRDPVDYNRSAAEVNLVDW 1375
Query: 60 VRVLLEHGNALECVDPSL 77
+++++ + +A E VDP++
Sbjct: 1376 LKMMVGNRHAEEVVDPNI 1429
>AW736597 similar to GP|12054894|e receptor protein kinase {Pinus
sylvestris}, partial (15%)
Length = 567
Score = 59.3 bits (142), Expect = 4e-10
Identities = 33/85 (38%), Positives = 53/85 (61%)
Frame = +1
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVR 61
SN + GY+APE ++++ EK DVY +GV++LE++TGK+P++ + L + D VR
Sbjct: 253 SNTVAGSYGYIAPEYG-YMMKITEKSDVYSYGVVLLEVLTGKQPIDPTIPDGLHVVDWVR 429
Query: 62 VLLEHGNALECVDPSLMSEYPEDEV 86
LE +DP+L+S PE E+
Sbjct: 430 ----QKRGLEVLDPTLLSR-PESEI 489
>TC91104 similar to GP|9758799|dbj|BAB09252.1 serine/threonine protein
kinase-like {Arabidopsis thaliana}, partial (29%)
Length = 996
Score = 58.2 bits (139), Expect = 8e-10
Identities = 28/79 (35%), Positives = 52/79 (65%)
Frame = +2
Query: 3 NRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRV 62
++ + GY+APE + V+EK DV+ FGV++LE+V+G+R ++Y + ++++ +
Sbjct: 89 SKLKGTFGYLAPEYLLHGI-VDEKTDVFAFGVLLLELVSGRRALDYSQQSLVL---WAKP 256
Query: 63 LLEHGNALECVDPSLMSEY 81
LL+ + +E VDPSL E+
Sbjct: 257 LLKKNDIMELVDPSLAGEF 313
>TC84262 similar to GP|15528705|dbj|BAB64771. putative receptor protein
kinase {Oryza sativa (japonica cultivar-group)},
partial (8%)
Length = 769
Score = 57.4 bits (137), Expect = 1e-09
Identities = 25/38 (65%), Positives = 33/38 (86%)
Frame = +1
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE 47
GY+APE AC +L++ EK DVY FGV++LEI+TGKRPV+
Sbjct: 31 GYIAPEYAC-MLKITEKSDVYSFGVVLLEIITGKRPVD 141
>BE240690 similar to PIR|T49986|T49 lectin-like protein kinase-like -
Arabidopsis thaliana, partial (25%)
Length = 634
Score = 57.0 bits (136), Expect = 2e-09
Identities = 28/78 (35%), Positives = 48/78 (60%)
Frame = +2
Query: 9 LGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLEHGN 68
+GY+APE A R +++ DVY FG++ LEI G++P+ ++N + + + V L G
Sbjct: 389 MGYMAPECATTG-RASKETDVYSFGIVALEIACGRKPIINAQENEINIVEWVWGLYGRGR 565
Query: 69 ALECVDPSLMSEYPEDEV 86
+E VDP L +Y E+++
Sbjct: 566 IVEAVDPRLDGDYEEEQI 619
>TC92500 similar to GP|14194119|gb|AAK56254.1 At1g01540/F22L4_6 {Arabidopsis
thaliana}, partial (34%)
Length = 569
Score = 56.6 bits (135), Expect = 2e-09
Identities = 33/82 (40%), Positives = 52/82 (63%), Gaps = 4/82 (4%)
Frame = +2
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY----GEDNVLILNDHVRVLLE 65
GYVAPE AC + + E+ DVY FG++I+E++TG+ PV+Y GE N++ + ++ ++
Sbjct: 182 GYVAPEYACTGM-LTERSDVYSFGILIMELITGRSPVDYSRPHGEVNLV---EWLKNMVG 349
Query: 66 HGNALECVDPSLMSEYPEDEVL 87
A E VDP + SE P + L
Sbjct: 350 SRRAEEVVDPKI-SEKPSSKAL 412
>TC90106 similar to GP|3641252|gb|AAC36318.1| leucine-rich receptor-like
protein kinase {Malus x domestica}, partial (25%)
Length = 1241
Score = 56.6 bits (135), Expect = 2e-09
Identities = 28/47 (59%), Positives = 38/47 (80%), Gaps = 2/47 (4%)
Frame = +1
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV--EYGEDNVL 54
GY+APE A L+VNEK D+Y FGV+ILE+VTG+RPV E+GE +++
Sbjct: 364 GYIAPEYA-YTLKVNEKSDIYSFGVVILELVTGRRPVDPEFGEKDLV 501
>AW329565 homologue to GP|19347728|gb putative receptor protein kinase
{Arabidopsis thaliana}, partial (32%)
Length = 545
Score = 56.2 bits (134), Expect = 3e-09
Identities = 29/75 (38%), Positives = 47/75 (62%)
Frame = +1
Query: 9 LGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLEHGN 68
LGYV+PE A + ++ DVY FG+++LE++TGKRPV + +D ++ V+ L+ G
Sbjct: 67 LGYVSPE-AILTSEITKESDVYSFGIVLLELLTGKRPVMFTQDEDIV--KWVKKQLQRGQ 237
Query: 69 ALECVDPSLMSEYPE 83
E ++P L+ PE
Sbjct: 238 ITELLEPGLLELDPE 282
>TC89684 similar to PIR|T10659|T10659 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) T5F17.100 - Arabidopsis
thaliana, partial (20%)
Length = 828
Score = 55.5 bits (132), Expect = 5e-09
Identities = 33/70 (47%), Positives = 49/70 (69%), Gaps = 2/70 (2%)
Frame = +3
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV--EYGEDNVLILNDHVRVLLEHG 67
GY+APE L+V+EK DVY +GV++LE+VTGKRP+ E+GE +V I+ R + E+
Sbjct: 288 GYIAPEYG-YALKVDEKIDVYSYGVVLLELVTGKRPLDSEFGE-SVDIVEWIRRKIRENK 461
Query: 68 NALECVDPSL 77
+ E +DPS+
Sbjct: 462 SLEEALDPSV 491
>CA857922 similar to PIR|T47793|T47 receptor-like protein kinase -
Arabidopsis thaliana, partial (47%)
Length = 810
Score = 55.1 bits (131), Expect = 7e-09
Identities = 32/78 (41%), Positives = 47/78 (60%), Gaps = 1/78 (1%)
Frame = +2
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGE-DNVLILNDH 59
++ R GYVAPE A L +NEK D+Y FGV++LE VTG+ PV+Y N + L +
Sbjct: 323 ITTRVMGTFGYVAPEYANSGL-LNEKSDIYSFGVLLLEAVTGRDPVDYARPSNEVNLVEW 499
Query: 60 VRVLLEHGNALECVDPSL 77
+++++ A E VD L
Sbjct: 500 LKMMVGARRAEEVVDSRL 553
>TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein kinase
{Ipomoea trifida}, partial (28%)
Length = 1276
Score = 54.7 bits (130), Expect = 9e-09
Identities = 35/89 (39%), Positives = 47/89 (52%), Gaps = 1/89 (1%)
Frame = +3
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTG-KRPVEYGEDNVLILNDHV 60
+NR GY++PE A + + + K DVY FGV++LEIV G K Y D L L H
Sbjct: 705 TNRIVGTYGYMSPEYAMEGV-CSTKSDVYSFGVLLLEIVCGIKNNSFYDVDRPLNLIGHA 881
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLPC 89
L G L+ +DP+L + DEV C
Sbjct: 882 WELWNDGEYLKLMDPTLNDTFVPDEVKRC 968
>TC83631 similar to PIR|T10659|T10659 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) T5F17.100 - Arabidopsis
thaliana, partial (24%)
Length = 975
Score = 54.7 bits (130), Expect = 9e-09
Identities = 27/79 (34%), Positives = 52/79 (65%), Gaps = 1/79 (1%)
Frame = +1
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLEHGNA 69
GY+APE L+V+EK D+Y FG+++LE++TGKRP++ + + +R ++ +
Sbjct: 412 GYIAPEYGYS-LKVDEKIDIYSFGIVLLELITGKRPIDPDFGESVDIVGWIRRKIDKNSP 588
Query: 70 LECVDPSLMS-EYPEDEVL 87
E +DPS+ + ++ ++E+L
Sbjct: 589 EEALDPSVGNCKHVQEEML 645
>TC88973 similar to PIR|T50817|T50817 protein serine/threonine kinase-like
protein - Arabidopsis thaliana, partial (42%)
Length = 839
Score = 54.3 bits (129), Expect = 1e-08
Identities = 31/90 (34%), Positives = 54/90 (59%), Gaps = 4/90 (4%)
Frame = +3
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYG----EDNVLIL 56
++ + + +G++APE + +EK DV+ +G+M+LE+VTG+R +++ ED+VL+L
Sbjct: 564 VTTQIRGTMGHIAPEYL-STGKPSEKTDVFSYGIMLLELVTGQRAIDFSRLEDEDDVLLL 740
Query: 57 NDHVRVLLEHGNALECVDPSLMSEYPEDEV 86
DHV+ L VD +L Y +EV
Sbjct: 741 -DHVKKLQRDKRLDAIVDSNLNKNYNIEEV 827
>TC86201 similar to PIR|T49003|T49003 protein kinase-like protein -
Arabidopsis thaliana, partial (86%)
Length = 2127
Score = 54.3 bits (129), Expect = 1e-08
Identities = 31/85 (36%), Positives = 47/85 (54%), Gaps = 4/85 (4%)
Frame = +3
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY----GEDNVLILN 57
S R GY APE A ++ +K DVY FGV++LE++TG++PV++ G+ +++
Sbjct: 912 STRVLGTFGYHAPEYA-MTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLV--- 1079
Query: 58 DHVRVLLEHGNALECVDPSLMSEYP 82
L +CVDP L EYP
Sbjct: 1080TWATPRLSEDKVKQCVDPKLKGEYP 1154
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,515,125
Number of Sequences: 36976
Number of extensions: 25965
Number of successful extensions: 456
Number of sequences better than 10.0: 311
Number of HSP's better than 10.0 without gapping: 421
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 422
length of query: 126
length of database: 9,014,727
effective HSP length: 85
effective length of query: 41
effective length of database: 5,871,767
effective search space: 240742447
effective search space used: 240742447
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146632.13


