

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146631.5 - phase: 0 /pseudo
(359 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
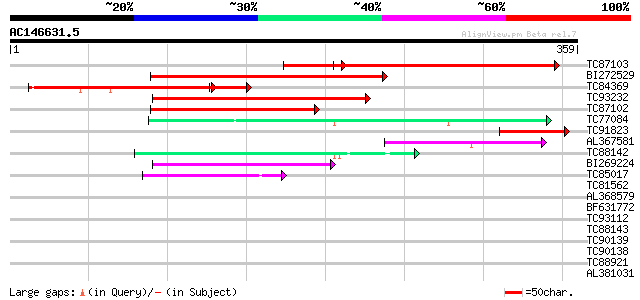
Score E
Sequences producing significant alignments: (bits) Value
TC87103 similar to GP|15292735|gb|AAK92736.1 unknown protein {Ar... 150 1e-43
BI272529 weakly similar to GP|6478925|gb| hypothetical protein {... 165 3e-41
TC84369 similar to GP|19310729|gb|AAL85095.1 unknown protein {Ar... 137 5e-40
TC93232 weakly similar to GP|21739227|emb|CAD40982. OSJNBa0072F1... 140 1e-33
TC87102 similar to PIR|T02589|T02589 hypothetical protein T16B24... 138 3e-33
TC77084 similar to GP|15529155|gb|AAK97672.1 AT3g30390/T6J22_16 ... 68 5e-12
TC91823 weakly similar to GP|19310729|gb|AAL85095.1 unknown prot... 60 1e-09
AL367581 similar to GP|15529155|gb AT3g30390/T6J22_16 {Arabidops... 47 9e-06
TC88142 weakly similar to GP|14587360|dbj|BAB61261. contains EST... 47 1e-05
BI269224 weakly similar to GP|11862972|db hypothetical protein {... 45 4e-05
TC85017 weakly similar to GP|9758059|dbj|BAB08638.1 amino acid t... 44 1e-04
TC81562 weakly similar to GP|6671946|gb|AAF23206.1| putative ami... 40 0.002
AL368579 similar to GP|14194151|gb AT5g40780/K1B16_3 {Arabidopsi... 39 0.004
BF631772 similar to PIR|F86385|F86 hypothetical protein AAG12739... 38 0.005
TC93112 weakly similar to GP|6671946|gb|AAF23206.1| putative ami... 38 0.007
TC88143 weakly similar to GP|20453045|gb|AAM19768.1 AT3g56200/F1... 36 0.020
TC90139 similar to GP|15290010|dbj|BAB63704. contains ESTs C9952... 35 0.034
TC90138 similar to GP|15290010|dbj|BAB63704. contains ESTs C9952... 33 0.22
TC88921 similar to SP|P92966|RS41_ARATH Arginine/serine-rich spl... 30 1.1
AL381031 29 2.5
>TC87103 similar to GP|15292735|gb|AAK92736.1 unknown protein {Arabidopsis
thaliana}, partial (51%)
Length = 1621
Score = 150 bits (380), Expect(2) = 1e-43
Identities = 68/144 (47%), Positives = 100/144 (69%), Gaps = 1/144 (0%)
Frame = +3
Query: 206 LHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTT-VGFHHT 264
L+ + LF V+ AL +LPTVWL+DL V+SY+S GG+ A++L+++ + +G VGF +
Sbjct: 507 LNXNPQTLFAVVAALAVLPTVWLRDLSVLSYISAGGVIASVLVVLCLLWIGIEDVGFQRS 686
Query: 265 GRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVM 324
G +N +P AIG+YG+C++GH+VFPNIY SMA Q+ L+ CF +C L+Y AVM
Sbjct: 687 GTTLNLGTLPVAIGLYGYCYSGHAVFPNIYTSMAKPNQFPAVLVACFGVCTLLYAGGAVM 866
Query: 325 GFLSFGDDTLSQITLNMPAGAFAS 348
G+ + G+DTLSQ TLN+P A+
Sbjct: 867 GYKNVGEDTLSQFTLNLPQDLVAT 938
Score = 43.5 bits (101), Expect(2) = 1e-43
Identities = 17/40 (42%), Positives = 26/40 (64%)
Frame = +2
Query: 174 YTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHLDSMHL 213
Y + C+E+I LEGDNL LFP L++GG+ +S ++
Sbjct: 410 YVVMQGCCIEYIILEGDNLASLFPNAYLNLGGIEXESTNI 529
>BI272529 weakly similar to GP|6478925|gb| hypothetical protein {Arabidopsis
thaliana}, partial (23%)
Length = 625
Score = 165 bits (417), Expect = 3e-41
Identities = 73/150 (48%), Positives = 108/150 (71%)
Frame = +2
Query: 90 CSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREG 149
CS+ ++V NG N++ G+GL++ PY VK+ G++ L L+L+FA +CCYT L+ C +S G
Sbjct: 131 CSFARSVINGTNLLCGIGLMTMPYAVKEGGFLSLALLLLFAVICCYTGILLMRCLQSYPG 310
Query: 150 LTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHLD 209
L +YPDIG+AAFG GR+ ++IIL EL+ VEF+TL DNL+ LFP TS+ + G L
Sbjct: 311 LDTYPDIGQAAFGYAGRLGIAIILSLELFGASVEFLTLVSDNLSALFPNTSMIVAGTELS 490
Query: 210 SMHLFGVLTALVILPTVWLKDLRVISYLSV 239
+ +F + AL++LPT WLK+L ++SY+SV
Sbjct: 491 THQVFTIAAALLVLPTGWLKNLSLLSYISV 580
>TC84369 similar to GP|19310729|gb|AAL85095.1 unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 614
Score = 137 bits (346), Expect(2) = 5e-40
Identities = 78/125 (62%), Positives = 92/125 (73%), Gaps = 6/125 (4%)
Frame = +3
Query: 13 YNRETTDSYTIATAPNFVSILRGPSSIYSSF--SNRSKSDLDI-DGKTPCLSG--LDGTT 67
YN E D TIA APN SILRGPS +YSSF S SKS L++ DGKT LSG +
Sbjct: 174 YN-EAIDPLTIAAAPNLGSILRGPSVLYSSFVGSFSSKSYLELHDGKTSFLSGNQIQEGF 350
Query: 68 QSTWWEKRSTQ-NLVGEMPLGYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLM 126
+TWWEK S Q + E+P+GYGC++TQT+FNGIN+MAGVGLLSTP TVKQAGW+ LV+M
Sbjct: 351 PTTWWEKASIQMQIPEELPIGYGCTFTQTIFNGINIMAGVGLLSTPDTVKQAGWVSLVVM 530
Query: 127 LIFAS 131
LIF S
Sbjct: 531 LIFCS 545
Score = 44.7 bits (104), Expect(2) = 5e-40
Identities = 19/27 (70%), Positives = 21/27 (77%)
Frame = +1
Query: 127 LIFASVCCYTATLMRHCFESREGLTSY 153
L FA VC YTA LM HCF+SREG+ SY
Sbjct: 532 LFFAVVCXYTAXLMXHCFQSREGIISY 612
>TC93232 weakly similar to GP|21739227|emb|CAD40982. OSJNBa0072F16.6 {Oryza
sativa}, partial (56%)
Length = 695
Score = 140 bits (352), Expect = 1e-33
Identities = 62/138 (44%), Positives = 88/138 (62%)
Frame = +3
Query: 91 SYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREGL 150
S+ T NG+N ++GVG+LS PY + GW+ L L+ A+ Y+ LM+ C E +
Sbjct: 165 SFFHTCVNGLNAISGVGILSVPYALASGGWLSLALLFCIAAAAFYSGILMKRCMEKNSNI 344
Query: 151 TSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHLDS 210
+YPDIGE AFG+ GR+ VSI +YTELY + F+ LEGDNL+ LFP + GL + +
Sbjct: 345 KTYPDIGELAFGKIGRLIVSISMYTELYLVSIGFLILEGDNLSNLFPIEEFQVFGLSIGA 524
Query: 211 MHLFGVLTALVILPTVWL 228
F +L A++ILPT+WL
Sbjct: 525 KKFFVILVAVIILPTIWL 578
>TC87102 similar to PIR|T02589|T02589 hypothetical protein T16B24.23 -
Arabidopsis thaliana (fragment), partial (44%)
Length = 964
Score = 138 bits (348), Expect = 3e-33
Identities = 62/107 (57%), Positives = 78/107 (71%)
Frame = +2
Query: 90 CSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREG 149
CS+ Q V NGINV+ GVG+LSTPY K+ GW+GL ++ IF + YT L+R C +S G
Sbjct: 608 CSFGQAVLNGINVLCGVGILSTPYAAKEGGWLGLSILFIFGILSFYTGLLLRSCLDSEPG 787
Query: 150 LTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLF 196
L +YPDIG+AAFG GRI +SI+LY ELY C+E+I LEGDNL LF
Sbjct: 788 LETYPDIGQAAFGTAGRIAISIVLYVELYGCCIEYINLEGDNLAFLF 928
>TC77084 similar to GP|15529155|gb|AAK97672.1 AT3g30390/T6J22_16
{Arabidopsis thaliana}, partial (88%)
Length = 2353
Score = 68.2 bits (165), Expect = 5e-12
Identities = 62/274 (22%), Positives = 110/274 (39%), Gaps = 19/274 (6%)
Frame = +1
Query: 89 GCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWM-GLVLMLIFASVCCYTATLMRHCFESR 147
G S++ VFN + G G+++ P T+KQ G + GL +L+ A + + L+ F
Sbjct: 469 GASFSGAVFNLATTIIGAGIMALPATLKQLGLIPGLCAILLMAFLTEKSIELLIR-FTRA 645
Query: 148 EGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIG--- 204
SY + +FG+YG+ I + + ++ + GD L+G G
Sbjct: 646 GKAVSYAGLMGDSFGKYGKAMAQICVIVNNIGVLIVYMIIIGDVLSGTSSSGEHHYGILE 825
Query: 205 ---GLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTTVGF 261
G+H + F VL V + T R+ S ++ + ++ V +VG ++
Sbjct: 826 GWFGVHWWTGRTFVVLLTTVAIFTPLASFKRIDSLRFTSALSVALAVVFLVIAVGISIVK 1005
Query: 262 HHTGRVVNWSGIPFA------------IGVYGFCFAGHSVFPNIYQSMADKKQYTKALIT 309
+G + P + V+ + H +I + D Q + T
Sbjct: 1006IISGGITMPRLFPAVTDATSIVNLFTVVPVFVTAYICHYNVHSIDNELEDNSQMQGVVRT 1185
Query: 310 CFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPA 343
LC +Y ++ GFL FG+ TL + N A
Sbjct: 1186ALGLCSSVYLMISFFGFLLFGEGTLDDVLANFDA 1287
>TC91823 weakly similar to GP|19310729|gb|AAL85095.1 unknown protein
{Arabidopsis thaliana}, partial (20%)
Length = 926
Score = 60.1 bits (144), Expect = 1e-09
Identities = 28/44 (63%), Positives = 34/44 (76%)
Frame = +3
Query: 311 FVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWT 354
FVL + +YGSV GFL FG+ T SQITL++P AFASKV+LWT
Sbjct: 48 FVLPVFLYGSVGAAGFLMFGERTSSQITLDLPRDAFASKVSLWT 179
>AL367581 similar to GP|15529155|gb AT3g30390/T6J22_16 {Arabidopsis
thaliana}, partial (33%)
Length = 525
Score = 47.4 bits (111), Expect = 9e-06
Identities = 33/114 (28%), Positives = 49/114 (42%), Gaps = 11/114 (9%)
Frame = -2
Query: 238 SVGGIAATILIIISVFSVGTTVGFHHTGRVVNWSGIPFAIG-VYGFCFAGHSVFP----- 291
+VG + ++II V S T+ G HH+G + W G+ + G + F +VF
Sbjct: 458 NVGVLIVYMIIIGDVVSGTTSSGIHHSGILEGWFGVHWWTGRTFVLTFTTLAVFAPLVSF 279
Query: 292 -----NIYQSMADKKQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLN 340
NI + D + LC +Y + GFL FGDDTL + N
Sbjct: 278 KQIVHNIDNELEDSSWMHGVVRASLTLCSSVYLLTSFFGFLLFGDDTLPDVLAN 117
>TC88142 weakly similar to GP|14587360|dbj|BAB61261. contains ESTs
AU163711(E4375) C25327(S5952) D40001(S1713)~similar to
Oryza sativa chromosome 6, partial (23%)
Length = 788
Score = 47.0 bits (110), Expect = 1e-05
Identities = 44/189 (23%), Positives = 76/189 (39%), Gaps = 9/189 (4%)
Frame = +1
Query: 80 LVGEMPLGYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATL 139
L G P S VFN + G G++S P +K G + +++ +V +
Sbjct: 100 LTGSNPAAVPGSIPGAVFNVATSIIGSGIMSIPAILKVLGVLPSFALILIVAVLAEVSVD 279
Query: 140 MRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGT 199
F T+Y + + AFG G + + + + V F+ + GD L+G G
Sbjct: 280 FLMRFTHAGVTTTYSGVMKEAFGSVGALSAQVCVIITNFGGLVLFLIIIGDVLSGKESGG 459
Query: 200 SLDIG------GLH---LDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIII 250
+ +G G+H + LF + LV+LP V K + + Y S GI+ + +
Sbjct: 460 EVHLGLLQQWFGIHWWNSREVALF-ITLVLVMLPLVLYKRVESLKYSS--GISTLLAVAF 630
Query: 251 SVFSVGTTV 259
G +
Sbjct: 631 VTICSGLAI 657
>BI269224 weakly similar to GP|11862972|db hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (26%)
Length = 537
Score = 45.1 bits (105), Expect = 4e-05
Identities = 31/116 (26%), Positives = 53/116 (44%)
Frame = +2
Query: 91 SYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREGL 150
S++ +VFN + G G+++ P +K G + +IF ++ +T+ + F
Sbjct: 143 SFSGSVFNLSTTIIGAGIMALPAAMKVLGLTIGIASIIFLALLSHTSLDILMRFSRVAKA 322
Query: 151 TSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGL 206
SY D+ AFG GR+ I + + V +I + GD L+G S G L
Sbjct: 323 QSYGDVMGYAFGSLGRLLFQISVLFNNFGILVVYIIIIGDVLSGTTSSGSHHFGVL 490
>TC85017 weakly similar to GP|9758059|dbj|BAB08638.1 amino acid
transporter-like protein {Arabidopsis thaliana}, partial
(11%)
Length = 481
Score = 43.9 bits (102), Expect = 1e-04
Identities = 28/92 (30%), Positives = 47/92 (50%), Gaps = 1/92 (1%)
Frame = +1
Query: 85 PLGYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWM-GLVLMLIFASVCCYTATLMRHC 143
P+ + + VFN M G G++S P T+K G + GL+++++ A + T M C
Sbjct: 196 PIPQNGTVSGAVFNISTTMVGAGIMSIPATIKVLGIIPGLLVIVLVAVITDLTVEFMLRC 375
Query: 144 FESREGLTSYPDIGEAAFGRYGRIFVSIILYT 175
S + +T +GE +FG G + V I + T
Sbjct: 376 TSSGKAVTYAGMVGE-SFGSVGSLAVKICVIT 468
>TC81562 weakly similar to GP|6671946|gb|AAF23206.1| putative amino acid
transporter protein {Arabidopsis thaliana}, partial
(36%)
Length = 673
Score = 39.7 bits (91), Expect = 0.002
Identities = 35/144 (24%), Positives = 59/144 (40%), Gaps = 9/144 (6%)
Frame = +1
Query: 152 SYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFP--GTSLDIGGLHLD 209
SY D+G +G GR I+ + + ++ G NL +F G SL
Sbjct: 247 SYGDLGYRCYGILGRYLTEFIIVVAQCAGAIAYLVFIGQNLDSVFQKYGPSL-------- 402
Query: 210 SMHLFGVLTALVILPTVWLKDLRVIS----YLSVGGIAATILIIISVFSVGTTVGFHHTG 265
+ ++F ++ V + W+ L ++ + V + A +++ V GF
Sbjct: 403 ASYIFMLVP--VEIGMSWIGSLSALAPFSIFADVCNVLAMGIVVKEDVQVALEDGFSFRE 576
Query: 266 RVV---NWSGIPFAIGVYGFCFAG 286
R N G+PFA G+ FCF G
Sbjct: 577 RTAITSNIGGLPFAAGMAVFCFEG 648
>AL368579 similar to GP|14194151|gb AT5g40780/K1B16_3 {Arabidopsis thaliana},
partial (29%)
Length = 504
Score = 38.5 bits (88), Expect = 0.004
Identities = 31/135 (22%), Positives = 57/135 (41%), Gaps = 6/135 (4%)
Frame = +1
Query: 43 FSNRSKSDLDIDGKTPCLSGLDGTTQSTWWEKRSTQNLVGE-MPLGYG--CSYTQTVFNG 99
F S S + + + P + +D EK Q + + +P+ + + F+
Sbjct: 31 FHTNSLSQVTMGTQAPTATDID--------EKLKRQKAIDDWLPISSSRNAKWWYSAFHN 186
Query: 100 INVMAGVGLLSTPYTVKQAGWMGLVLMLIFA-SVCCYTATLMRHCFESREG--LTSYPDI 156
+ M G G+LS PY + Q GW V +L+ + + YT M E G Y ++
Sbjct: 187 VTAMVGAGVLSLPYAMAQLGWGPEVTVLVLSWFITLYTLWQMVEMHEMVPGKRFDRYHEL 366
Query: 157 GEAAFGRYGRIFVSI 171
+ AFG +++ +
Sbjct: 367 SQYAFGEKLGLYIVV 411
>BF631772 similar to PIR|F86385|F86 hypothetical protein AAG12739.1
[imported] - Arabidopsis thaliana, partial (29%)
Length = 523
Score = 38.1 bits (87), Expect = 0.005
Identities = 23/73 (31%), Positives = 34/73 (46%), Gaps = 5/73 (6%)
Frame = +2
Query: 95 TVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREGLTS-- 152
+ F+ + M G G+LS PY + GW +LML+ + C T M + E +
Sbjct: 128 STFHTVTAMIGAGVLSLPYAMAYLGWGPGILMLLLS--WCLTLNTMWQMIQLHECVPGTR 301
Query: 153 ---YPDIGEAAFG 162
Y D+G AFG
Sbjct: 302 FDRYIDLGRHAFG 340
>TC93112 weakly similar to GP|6671946|gb|AAF23206.1| putative amino acid
transporter protein {Arabidopsis thaliana}, partial
(39%)
Length = 680
Score = 37.7 bits (86), Expect = 0.007
Identities = 17/47 (36%), Positives = 28/47 (59%)
Frame = +3
Query: 296 SMADKKQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMP 342
SM DK ++ K L F L+Y + G+++FG++T +TLN+P
Sbjct: 39 SMKDKAKFPKLLAQTFSGITLVYILFGLCGYMAFGEETRDIVTLNLP 179
>TC88143 weakly similar to GP|20453045|gb|AAM19768.1 AT3g56200/F18O21_160
{Arabidopsis thaliana}, partial (51%)
Length = 892
Score = 36.2 bits (82), Expect = 0.020
Identities = 20/86 (23%), Positives = 40/86 (46%)
Frame = +1
Query: 270 WSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVMGFLSF 329
++ +P + + F F H I +A A+ + C ++Y ++ + G+L F
Sbjct: 325 FTAVPVVVTAFTFHFNVHP----IGFELASPSHMKTAVRLAIMFCAVVYFTIGLFGYLLF 492
Query: 330 GDDTLSQITLNMPAGAFASKVALWTT 355
GD T S I +N A ++ +L+ +
Sbjct: 493 GDSTQSDILINFDQNADSAVGSLFNS 570
>TC90139 similar to GP|15290010|dbj|BAB63704. contains ESTs C99526(E20351)
C73742(E20351)~unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (26%)
Length = 680
Score = 35.4 bits (80), Expect = 0.034
Identities = 35/120 (29%), Positives = 49/120 (40%), Gaps = 6/120 (5%)
Frame = +2
Query: 108 LLSTPYTVKQAGW-MGLVLMLIFASVCCYTATLMRHCFESREGLTS----YPDIGEAAFG 162
LLS PY GW G+ ++I A V Y+ L+ E + L + + D+ G
Sbjct: 254 LLSLPYAFTLLGWTAGIFFLVIGAMVTFYSYNLLSRVLEHQAQLGNRQLRFRDMARDILG 433
Query: 163 -RYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALV 221
R+GR FV I + Y V L G + + H+DS H LT LV
Sbjct: 434 PRWGRYFVGPIQFAVCYGAVVACTLLGGQCMKVILNPCDTH---THIDS-HFLHALTILV 601
>TC90138 similar to GP|15290010|dbj|BAB63704. contains ESTs C99526(E20351)
C73742(E20351)~unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (30%)
Length = 573
Score = 32.7 bits (73), Expect = 0.22
Identities = 27/95 (28%), Positives = 40/95 (41%), Gaps = 6/95 (6%)
Frame = +3
Query: 108 LLSTPYTVKQAGW-MGLVLMLIFASVCCYTATLMRHCFESREGLTS----YPDIGEAAFG 162
LLS PY GW G+ ++I A V Y+ L+ E + L + + D+ G
Sbjct: 237 LLSLPYAFTLLGWTAGIFFLVIGAMVTFYSYNLLSRVLEHQAQLGNRQLRFRDMARDILG 416
Query: 163 -RYGRIFVSIILYTELYSYCVEFITLEGDNLTGLF 196
R+GR FV I Y V L G + ++
Sbjct: 417 PRWGRYFVGPIQVAVCYGAVVACTLLGGQCMKAVY 521
>TC88921 similar to SP|P92966|RS41_ARATH Arginine/serine-rich splicing
factor RSP41. [Mouse-ear cress] {Arabidopsis thaliana},
partial (33%)
Length = 1419
Score = 30.4 bits (67), Expect = 1.1
Identities = 21/54 (38%), Positives = 28/54 (50%), Gaps = 8/54 (14%)
Frame = -1
Query: 232 RVISYLSVGGIAAT-ILIIISVFS---VGTTVGFHH----TGRVVNWSGIPFAI 277
R+ S+ G +A T IL I F VGT GF H T ++ W+G+PF I
Sbjct: 969 RLKSFFCRG*LALTAILFIHGTFRTTIVGTVPGFRHSIWATSIIIVWAGVPFCI 808
>AL381031
Length = 336
Score = 29.3 bits (64), Expect = 2.5
Identities = 10/31 (32%), Positives = 18/31 (57%)
Frame = +1
Query: 250 ISVFSVGTTVGFHHTGRVVNWSGIPFAIGVY 280
+ V+ VG +HH G ++N PF++ V+
Sbjct: 52 VKVYGVGKF*NYHHLGEILNIGAFPFSVLVF 144
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.140 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,240,674
Number of Sequences: 36976
Number of extensions: 174766
Number of successful extensions: 1234
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 1217
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1228
length of query: 359
length of database: 9,014,727
effective HSP length: 97
effective length of query: 262
effective length of database: 5,428,055
effective search space: 1422150410
effective search space used: 1422150410
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146631.5


