

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146588.5 - phase: 0 /pseudo
(637 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
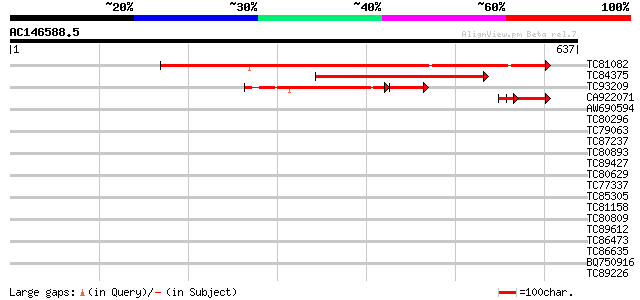
Score E
Sequences producing significant alignments: (bits) Value
TC81082 similar to GP|17385678|dbj|BAB78631. putative co-repress... 624 e-179
TC84375 similar to GP|17385678|dbj|BAB78631. putative co-repress... 387 e-108
TC93209 similar to PIR|T51447|T51447 transcription regulator-lik... 176 4e-60
CA922071 similar to GP|8247350|emb| pSbaNS5 protein {Sorghum bic... 77 2e-14
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 39 0.005
TC80296 similar to PIR|H86265|H86265 protein F3F19.18 [imported]... 35 0.088
TC79063 similar to GP|15215854|gb|AAK91471.1 At2g43170/F14B2.11 ... 35 0.088
TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-inf... 35 0.088
TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {... 35 0.12
TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed pr... 35 0.12
TC80629 35 0.12
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 34 0.15
TC85305 aquaporin 34 0.20
TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidops... 33 0.34
TC80809 similar to GP|22136852|gb|AAM91770.1 unknown protein {Ar... 33 0.34
TC89612 similar to GP|6934300|gb|AAF31706.1| unknown {Euphorbia ... 33 0.44
TC86473 similar to PIR|E84828|E84828 probable WD-40 repeat prote... 33 0.44
TC86635 weakly similar to GP|9758600|dbj|BAB09233.1 gene_id:MDF2... 32 0.57
BQ750916 weakly similar to SP|P03186|TEGU Large tegument protein... 32 0.75
TC89226 weakly similar to GP|9759519|dbj|BAB10985.1 gb|AAF16763.... 32 0.75
>TC81082 similar to GP|17385678|dbj|BAB78631. putative co-repressor protein
{Oryza sativa (japonica cultivar-group)}, partial (20%)
Length = 1461
Score = 624 bits (1608), Expect = e-179
Identities = 324/444 (72%), Positives = 365/444 (81%), Gaps = 6/444 (1%)
Frame = +3
Query: 170 GLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNF 229
GL+ PSSEG DS R STNGA+ T+ R +ES FKSEREEGELSPNGDFEEDNF
Sbjct: 30 GLEGPSSEGGDSARLYASTNGAVAGGTEVCRYQEESNPQFKSEREEGELSPNGDFEEDNF 209
Query: 230 AVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGEN----DDESDGSPHRSSDDSEN 285
AVY +AG EAVHKG D + YQNRR EQ+CG A GEN DDE + SP RSS+ SEN
Sbjct: 210 AVYGDAGSEAVHKGNDGGLKRQYQNRRGEQVCGEARGENYADADDEGEESPQRSSEGSEN 389
Query: 286 ASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSER 345
AS N DVSG+ESADGEECS+EEH+ DG+HD+K ESEGEAEGMADA+DVEGDG SLP+SER
Sbjct: 390 ASGNVDVSGSESADGEECSQEEHD-DGEHDDKAESEGEAEGMADAHDVEGDGTSLPFSER 566
Query: 346 FLLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKW 405
FLL KPL K+V PV H +++N ++FYGNDSFYVL RLHQTLYERI +AK+NSSS E+KW
Sbjct: 567 FLLNVKPLAKHVPPVLHVEDKNSRVFYGNDSFYVLIRLHQTLYERIHAAKVNSSSTERKW 746
Query: 406 RASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQ 465
+ASN+TSSTD+Y RFM +LYSLLDGSSDNSKFEDDCRAIIGTQSY+LFTLDKLIYKLVKQ
Sbjct: 747 KASNNTSSTDQYDRFMIALYSLLDGSSDNSKFEDDCRAIIGTQSYLLFTLDKLIYKLVKQ 926
Query: 466 LQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECS--PTRL 523
LQAVAS DEMDNKLLQLYAYE+SRK G F D+VYHENARVLLHDENIYRIE S P L
Sbjct: 927 LQAVAS--DEMDNKLLQLYAYEKSRKPGKFVDIVYHENARVLLHDENIYRIEYSPKPKTL 1100
Query: 524 SIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELS 583
SIQLMD G++K EVTAV +DPNFS YLHNDFLS+ D +K GIFLKRNK A SDE S
Sbjct: 1101SIQLMDCGNEKHEVTAVLVDPNFSAYLHNDFLSIASD--KKLGIFLKRNKRGYACSDEFS 1274
Query: 584 SQVMDGIQVINGLECKIACGSSKV 607
S+ M+G+Q+INGLECKIA SSKV
Sbjct: 1275SEAMEGLQIINGLECKIASNSSKV 1346
>TC84375 similar to GP|17385678|dbj|BAB78631. putative co-repressor protein
{Oryza sativa (japonica cultivar-group)}, partial (10%)
Length = 587
Score = 387 bits (995), Expect = e-108
Identities = 193/195 (98%), Positives = 194/195 (98%)
Frame = +3
Query: 344 ERFLLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEK 403
E FLLTAKPLVKYVSPVFHGKEENVQIFYGND+FYVLFRLHQTLYERIRSAKINSSSAEK
Sbjct: 3 EGFLLTAKPLVKYVSPVFHGKEENVQIFYGNDAFYVLFRLHQTLYERIRSAKINSSSAEK 182
Query: 404 KWRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLV 463
KWRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLV
Sbjct: 183 KWRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLV 362
Query: 464 KQLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECSPTRL 523
KQLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECSPTRL
Sbjct: 363 KQLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECSPTRL 542
Query: 524 SIQLMDYGHDKPEVT 538
SIQLMDYGHDKPEVT
Sbjct: 543 SIQLMDYGHDKPEVT 587
>TC93209 similar to PIR|T51447|T51447 transcription regulator-like protein -
Arabidopsis thaliana, partial (10%)
Length = 721
Score = 176 bits (445), Expect(2) = 4e-60
Identities = 103/170 (60%), Positives = 125/170 (72%), Gaps = 7/170 (4%)
Frame = +3
Query: 264 AGGENDDESDGSPHRSSDDSENASENG-DVSGTESADGEECSREEHEEDG----DHDNKV 318
AGG+ND ++D +DSEN SE G DVSG+ESA G+ECSRE+HEE+ D D K
Sbjct: 102 AGGDNDADAD------DEDSENVSEAGEDVSGSESA-GDECSREDHEEEDMEHDDVDGKA 260
Query: 319 ESEGEAEGMADANDVEG-DGASLPYSERFLLTAKPLVKYVSPV-FHGKEENVQIFYGNDS 376
ESEGEAEGM DA+ G DG+SLP SERFL T KPL K+VS V F ++ ++FYGND
Sbjct: 261 ESEGEAEGMCDADAQTGVDGSSLPLSERFLSTVKPLTKHVSAVSFVEDVKDSRVFYGNDD 440
Query: 377 FYVLFRLHQTLYERIRSAKINSSSAEKKWRASNDTSSTDRYARFMNSLYS 426
F+VLFRLHQ LYERI SAK NS+SAE KW+ + D SSTD YARFM++LY+
Sbjct: 441 FFVLFRLHQILYERILSAKENSTSAEIKWK-TKDASSTDLYARFMDALYN 587
Score = 74.7 bits (182), Expect(2) = 4e-60
Identities = 34/44 (77%), Positives = 41/44 (92%)
Frame = +1
Query: 427 LLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVA 470
LLDGS +N+KFE++CRAI+G QSYVLFTLDKLIYKL++QLQ VA
Sbjct: 589 LLDGSPENAKFENECRAILGNQSYVLFTLDKLIYKLIRQLQTVA 720
>CA922071 similar to GP|8247350|emb| pSbaNS5 protein {Sorghum bicolor},
partial (14%)
Length = 808
Score = 77.4 bits (189), Expect = 2e-14
Identities = 39/49 (79%), Positives = 40/49 (81%)
Frame = -1
Query: 559 PDSKEKSGIFLKRNKLKCARSDELSSQVMDGIQVINGLECKIACGSSKV 607
P +E K NKLKCARSDELSSQVMDGIQVINGLECKIACGSSKV
Sbjct: 778 PTVRESQEFS*KGNKLKCARSDELSSQVMDGIQVINGLECKIACGSSKV 632
Score = 43.5 bits (101), Expect = 2e-04
Identities = 20/22 (90%), Positives = 20/22 (90%)
Frame = -3
Query: 550 LHNDFLSVVPDSKEKSGIFLKR 571
LHNDFLSV PDSK KSGIFLKR
Sbjct: 806 LHNDFLSVSPDSKRKSGIFLKR 741
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 39.3 bits (90), Expect = 0.005
Identities = 47/202 (23%), Positives = 77/202 (37%), Gaps = 16/202 (7%)
Frame = +3
Query: 268 NDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGM 327
N++E +G D+ + E+ E + EE EE EED D + E E E E
Sbjct: 54 NEEEDEGEVEEEEDEHDEEEED------EHDEEEEEEEEEEEEDEHDDEEEEEEEEEEEE 215
Query: 328 ADANDVEGDGASLPYSERFLLTAKPLVKYV----------------SPVFHGKEENVQIF 371
+ +D EG+ + +R PL+ ++ S + E +Q+F
Sbjct: 216 EEDDDEEGEEDEI---DRISTLPDPLLLHILSFLPTRTSVATMSLASRKWRNLWEQLQVF 386
Query: 372 YGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGS 431
DS V H Y++ KK+ D+ R +R + + L G
Sbjct: 387 NFKDSDPVFHTPHDCEYKKF-----------KKFAYYVDSVLALRRSRNIRK-FRLSCGY 530
Query: 432 SDNSKFEDDCRAIIGTQSYVLF 453
+D +KF+ RA G L+
Sbjct: 531 ADENKFKVWIRAATGPHLQQLY 596
>TC80296 similar to PIR|H86265|H86265 protein F3F19.18 [imported] -
Arabidopsis thaliana, partial (34%)
Length = 1734
Score = 35.0 bits (79), Expect = 0.088
Identities = 24/75 (32%), Positives = 33/75 (44%)
Frame = +3
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEG 322
V+ +DD GS + SDD E +G +D E ++E GD + VE EG
Sbjct: 828 VSLNSDDDNQLGSDNTGSDDDEAEDHDG------VSDDENDRSSDYETSGDDADNVEDEG 989
Query: 323 EAEGMADANDVEGDG 337
+ D D E DG
Sbjct: 990 D-----DLEDSEEDG 1019
>TC79063 similar to GP|15215854|gb|AAK91471.1 At2g43170/F14B2.11
{Arabidopsis thaliana}, partial (34%)
Length = 1726
Score = 35.0 bits (79), Expect = 0.088
Identities = 52/244 (21%), Positives = 93/244 (37%), Gaps = 2/244 (0%)
Frame = +1
Query: 100 STLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATL 159
STL +I++S D+ + V K+ ++ LVN R+ + + +N T
Sbjct: 493 STLSDFQYIDSSGRDQGNNVRKKSQNLVVLVND----KERIVEVRQKAAVNREKFRNNT- 657
Query: 160 SRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELS 219
PG +YR G SS G+ R + ED + +E R++ +
Sbjct: 658 --PGGMYRPGS---HSSIGSYGDRYEEDRYANREEDRNGYGYGRE---REMGSRDDDRYN 813
Query: 220 PNGD-FEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHR 278
+GD + D Y G +G+ + +Y + R + D + D
Sbjct: 814 RDGDRYGRDYEERYGRDGYRDDDRGRSRSVDYNYDDTRSR------NSDRDRDFD----- 960
Query: 279 SSDDSENASENGDVSGTESADGEECSREEHEED-GDHDNKVESEGEAEGMADANDVEGDG 337
DD +++S + + + R+ E++ G + E+ GEA+ DVE
Sbjct: 961 --DDGQHSSRGSNAKVEDQSLEARLQRKLSEQNSGAPPSYEEAVGEAQSPVPERDVETSA 1134
Query: 338 ASLP 341
S P
Sbjct: 1135ESAP 1146
>TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-infected
erythrocyte surface antigen {Plasmodium falciparum},
partial (2%)
Length = 2007
Score = 35.0 bits (79), Expect = 0.088
Identities = 31/163 (19%), Positives = 68/163 (41%), Gaps = 5/163 (3%)
Frame = +1
Query: 175 SSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYAN 234
SSE + S + K+H ++E+ KS++E E+ +++N N
Sbjct: 49 SSEEKSHLENEESKETGESSEEKSHLENEENKDEEKSKQENEEIKDGEKIQQEN---EEN 219
Query: 235 AGLEAVHKGKDCNTSQHYQNRREEQICGVAGGEND-----DESDGSPHRSSDDSENASEN 289
E + + N + ++++E ++ GGE + +E + + ++ + +
Sbjct: 220 KDEEKSQQENEENKDEE-KSQQENELKKNEGGEKETGEITEEKSKQENEETSETNSKDKE 396
Query: 290 GDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADAND 332
+ S +D +E E HE+D + E+ G G + +D
Sbjct: 397 NEESNQNGSDAKEQVGENHEQDSKQGTE-ETNGTEGGEKEEHD 522
Score = 32.3 bits (72), Expect = 0.57
Identities = 29/139 (20%), Positives = 55/139 (38%), Gaps = 18/139 (12%)
Frame = +1
Query: 212 EREEGELSPNGDFEEDNFAVYANA----GLEAVHKGKDCNTS---QHYQNRRE--EQICG 262
E+E GE++ +E+ N+ E+ G D H Q+ ++ E+ G
Sbjct: 313 EKETGEITEEKSKQENEETSETNSKDKENEESNQNGSDAKEQVGENHEQDSKQGTEETNG 492
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECS---------REEHEEDGD 313
GGE ++ SSD+ E + + E+ G+ S ++ +
Sbjct: 493 TEGGEKEEHDKIKEDTSSDNQVQDGEKNNEAREENYSGDNASSAVVDNKSQESSNKTEEQ 672
Query: 314 HDNKVESEGEAEGMADAND 332
D K ++E E E ++N+
Sbjct: 673 FDKKEKNEFELESQKNSNE 729
Score = 29.6 bits (65), Expect = 3.7
Identities = 66/326 (20%), Positives = 113/326 (34%), Gaps = 17/326 (5%)
Frame = +1
Query: 24 TEDAVKAKKDSAKTGTASI-----------AKGDSSPGVGATVMSPKNTFEQSNSCKEWQ 72
TE K + D K T+S A+ ++ G A+ N ++S++ E Q
Sbjct: 493 TEGGEKEEHDKIKEDTSSDNQVQDGEKNNEAREENYSGDNASSAVVDNKSQESSNKTEEQ 672
Query: 73 TNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNA 132
+ K + L+S ++ +T ST+ N N S D+ N +A
Sbjct: 673 FDKKE--KNEFELESQKNSNETTESTDSTITQNSQGNESEKDQAQTENDTPKG-----SA 831
Query: 133 NVSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAI 192
+ S + EQ T ++ D S G D+T T+
Sbjct: 832 SESDEQKQEQEQNNTTKDDVQTT----------------DTSSQNGNDTTEKQNETS--- 954
Query: 193 IEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQ-H 251
ED + + S + E+ + GD + ++ + +D NT+
Sbjct: 955 -EDANSKK-EDSSALNTTPNNEDSKSGVAGDQADSTTTTSSS-------ETQDGNTNHGE 1107
Query: 252 YQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEE-----CSRE 306
Y++ E +G E ES GS + D+ + AS ++ T E+ S E
Sbjct: 1108YKDTTNENPEKNSGQEGTQES-GSSSNTFDNKDAASNKVQLTTTSDTSSEQKKDESSSAE 1284
Query: 307 EHEEDGDHDNKVESEGEAEGMADAND 332
E +DN + AND
Sbjct: 1285SKSESSQNDNANSGQSNTTSDESAND 1362
>TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {Arabidopsis
thaliana}, partial (39%)
Length = 883
Score = 34.7 bits (78), Expect = 0.12
Identities = 18/58 (31%), Positives = 29/58 (49%), Gaps = 3/58 (5%)
Frame = +3
Query: 257 EEQICGVAG---GENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEED 311
EE G G G+ +D+ + + SDD E+ ++GD E + EE +E E+D
Sbjct: 384 EEDYSGDEGEEEGDPEDDPEANGAGGSDDGEDDDDDGDEEDEEDGEDEEDEEDEEEDD 557
Score = 29.6 bits (65), Expect = 3.7
Identities = 22/74 (29%), Positives = 28/74 (37%), Gaps = 17/74 (22%)
Frame = +3
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGTE--------------SADGEECS---REEHE 309
+ DD+ D DD E +GD E S DGE+ EE E
Sbjct: 330 DEDDDDDDDVQDEDDDGEEEDYSGDEGEEEGDPEDDPEANGAGGSDDGEDDDDDGDEEDE 509
Query: 310 EDGDHDNKVESEGE 323
EDG+ + E E E
Sbjct: 510 EDGEDEEDEEDEEE 551
>TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed protein
{Arabidopsis thaliana}, partial (50%)
Length = 753
Score = 34.7 bits (78), Expect = 0.12
Identities = 27/96 (28%), Positives = 41/96 (42%), Gaps = 7/96 (7%)
Frame = +1
Query: 257 EEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEEC-------SREEHE 309
EE G ++DDE D DD + +GD E AD E+ ++ +
Sbjct: 283 EENKDGSETEDDDDEDDDDDVNDEDDDNDEDFSGD-EDDEDADPEDDPVPNGAGGSDDDD 459
Query: 310 EDGDHDNKVESEGEAEGMADANDVEGDGASLPYSER 345
ED D D+ +GE E D ++ E D P S++
Sbjct: 460 EDDDDDDDDNDDGEDE---DEDEEEDDDEDQPPSKK 558
Score = 30.4 bits (67), Expect = 2.2
Identities = 17/59 (28%), Positives = 27/59 (44%)
Frame = +1
Query: 280 SDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGA 338
S + EN D S TE D E+ + ++ED D+D + + E +D +GA
Sbjct: 262 SKGNHTCEENKDGSETEDDDDEDDDDDVNDEDDDNDEDFSGDEDDEDADPEDDPVPNGA 438
Score = 29.3 bits (64), Expect = 4.8
Identities = 26/108 (24%), Positives = 44/108 (40%)
Frame = +1
Query: 208 HFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGE 267
H E ++G + + D E+D+ + V+ D N +E G E
Sbjct: 274 HTCEENKDGSETEDDDDEDDD---------DDVNDEDDDN---------DEDFSGDEDDE 399
Query: 268 NDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHD 315
+ D D + S++ E+ D ++ DGE+ E+ EED D D
Sbjct: 400 DADPEDDPVPNGAGGSDDDDEDDDDDDDDNDDGEDED-EDEEEDDDED 540
>TC80629
Length = 847
Score = 34.7 bits (78), Expect = 0.12
Identities = 40/159 (25%), Positives = 65/159 (40%), Gaps = 5/159 (3%)
Frame = +1
Query: 137 SSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDT 196
SS + N ++ ASG + + R + L SS+ DS DTS + + + +
Sbjct: 175 SSTSLKKNKHSSKTKASGRSKSRKHKSKKVRRREVSLSSSDYDDSKSMDTSLSSSSEDRS 354
Query: 197 KAHRCHKESVGHFKSEREE-GELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQN- 254
K R KS +++ S + D ED ++YA + K K T + +
Sbjct: 355 KRKRERSRPRKDVKSRKKKVRRRSYSSDSSED--SLYAKKRKKVKRKDKHDETKEKSRRK 528
Query: 255 ---RREEQICGVAGGENDDESDGSPHRSSDDSENASENG 290
+RE + V+ G D G S+D+SE S G
Sbjct: 529 KKVKRETSVSSVSSGSRRDVQLGID--SNDNSEYESSRG 639
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 34.3 bits (77), Expect = 0.15
Identities = 32/125 (25%), Positives = 51/125 (40%)
Frame = +3
Query: 210 KSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGEND 269
K EGE + + EED+ G A +G+D +S+ G G N
Sbjct: 399 KQHDAEGEDGNDDEDEEDD------DGDGAFGEGEDELSSED----------GGGYGNNS 530
Query: 270 DESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMAD 329
+ S + A ENG+ E D ++ +E E+D D + E E E EG+ +
Sbjct: 531 NNKSNSKKAPEGGAGGADENGEEEDDEDGDDQDEDDDEDEDDDDEEEGGE-EDEEEGVDE 707
Query: 330 ANDVE 334
++ E
Sbjct: 708 EDNEE 722
Score = 32.3 bits (72), Expect = 0.57
Identities = 30/108 (27%), Positives = 44/108 (39%), Gaps = 12/108 (11%)
Frame = +3
Query: 242 KGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDD--SENASENGDVSGTES-- 297
KG SQ Q+ E + E DD+ DG+ D+ SE+ G+ S +S
Sbjct: 369 KGDSKTQSQDKQHDAEGEDGNDDEDEEDDDGDGAFGEGEDELSSEDGGGYGNNSNNKSNS 548
Query: 298 --------ADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDG 337
+E EE +EDGD ++ + E E D +D E G
Sbjct: 549 KKAPEGGAGGADENGEEEDDEDGDDQDEDDDEDE-----DDDDEEEGG 677
>TC85305 aquaporin
Length = 1221
Score = 33.9 bits (76), Expect = 0.20
Identities = 30/113 (26%), Positives = 46/113 (40%), Gaps = 10/113 (8%)
Frame = -3
Query: 237 LEAVHKGKDCNTSQHYQNRREEQI----CGVAGGENDDESDGSPHRS---SDDSENASEN 289
+ VH+ K + QH + C + D+ GS + S SDD + +E
Sbjct: 574 IHCVHQSKGHHNLQHQSWSNSNSL**SECWNRRSGDKDKEGGSNNGSEELSDDVNDTTEE 395
Query: 290 GDVS---GTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGAS 339
GDV+ GTE G + S + +GD + + E G+ D G G S
Sbjct: 394 GDVTTNEGTEGDSGVDMSTGDVSTNGDSNKESECMGDG-----GRDETGRGGS 251
>TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidopsis
thaliana}, partial (6%)
Length = 890
Score = 33.1 bits (74), Expect = 0.34
Identities = 20/71 (28%), Positives = 37/71 (51%), Gaps = 6/71 (8%)
Frame = +1
Query: 268 NDDESDGS-----PHRSSDDSENASENGDVSGTESADGEECSREEHEEDG-DHDNKVESE 321
N+D+ DGS S +SE+++ + SG++S + +E E+ EE D K+ +
Sbjct: 400 NEDKDDGSEGESESESSESESESSTSSSSTSGSDSDEDDEDDAEDEEEGQISDDEKMVTW 579
Query: 322 GEAEGMADAND 332
A+G D ++
Sbjct: 580 SAADGDLDDDE 612
>TC80809 similar to GP|22136852|gb|AAM91770.1 unknown protein {Arabidopsis
thaliana}, partial (11%)
Length = 681
Score = 33.1 bits (74), Expect = 0.34
Identities = 28/104 (26%), Positives = 50/104 (47%), Gaps = 13/104 (12%)
Frame = +3
Query: 244 KDCNTSQHYQNRREE-QICGVAGGENDDESDGSPH------RSSDDSENASENGD----- 291
KD +TS+ + Q + G ++D SDG + SDD +N ++N D
Sbjct: 123 KDDSTSESEEEEAPPLQHLELDGSDSDLSSDGDDQLADDFLQGSDDEDNDNDNDDEGSDL 302
Query: 292 VSGTESADGEECSREEHEEDGDHDNK-VESEGEAEGMADANDVE 334
SG+ S + E+ E EED +K ++ + E + A A++++
Sbjct: 303 ESGSGSDEDEDSDSESGEEDIARKSKLIDRKRERDSQAAADELQ 434
>TC89612 similar to GP|6934300|gb|AAF31706.1| unknown {Euphorbia esula},
complete
Length = 1504
Score = 32.7 bits (73), Expect = 0.44
Identities = 30/97 (30%), Positives = 40/97 (40%), Gaps = 2/97 (2%)
Frame = +3
Query: 254 NRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGD 313
++ ++ GV G NDD+ D+ N NGD G E DG+ EE G
Sbjct: 264 SKSDQPSIGVNDGGNDDDGG--------DNFNGGGNGDDGGGEEEDGD-------EEFGP 398
Query: 314 HDN--KVESEGEAEGMADANDVEGDGASLPYSERFLL 348
N V E EA G+ +D+E E FLL
Sbjct: 399 LLNFEAVMKEAEARGVKLPSDMEEAARVTGIREMFLL 509
>TC86473 similar to PIR|E84828|E84828 probable WD-40 repeat protein
[imported] - Arabidopsis thaliana, partial (68%)
Length = 2654
Score = 32.7 bits (73), Expect = 0.44
Identities = 26/96 (27%), Positives = 47/96 (48%), Gaps = 19/96 (19%)
Frame = +2
Query: 268 NDDESDGSPHRS-----SDDSENASENGDVSGTESADGEECSREEHEE------------ 310
+DD+SD S S S+D ++++++ D + +D + S E EE
Sbjct: 131 DDDDSDDSVEVSDYDSISEDEDSSADSIDDDDSSDSDSDSKSELEDEEGASPTTHHNASD 310
Query: 311 DGDHDNKVESEGEAEGMADANDV--EGDGASLPYSE 344
D D ++ E ++EG ++++D+ EG GA SE
Sbjct: 311 DSDSQDEEEDVVDSEGGSESSDLHQEGGGAESDSSE 418
Score = 32.3 bits (72), Expect = 0.57
Identities = 24/70 (34%), Positives = 35/70 (49%), Gaps = 3/70 (4%)
Frame = +2
Query: 267 ENDDESDGSP---HRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGE 323
E +DE SP H +SDDS++ E DV +S G E S + H+E G ES+
Sbjct: 257 ELEDEEGASPTTHHNASDDSDSQDEEEDV--VDSEGGSE-SSDLHQEGGG----AESDSS 415
Query: 324 AEGMADANDV 333
+ +A N +
Sbjct: 416 EDEVAPRNTI 445
>TC86635 weakly similar to GP|9758600|dbj|BAB09233.1
gene_id:MDF20.10~pir||T08929~similar to unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1963
Score = 32.3 bits (72), Expect = 0.57
Identities = 32/151 (21%), Positives = 62/151 (40%), Gaps = 9/151 (5%)
Frame = +2
Query: 197 KAHRCHKESVGHF---------KSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCN 247
K RC+KE + F KS +E ++ DF V +A V K+
Sbjct: 602 KLDRCNKEKLLEFCDVLDITITKSTTKEDIIAKLTDF-----LVAPHATTTVVLADKE-- 760
Query: 248 TSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREE 307
S + R ++ +GG + G + ++DS + + ++GEE + E
Sbjct: 761 -SSKKRRRTVKRGTPRSGGASTSRGSGKRQKKNEDSSGVEKKSTTDTEDESEGEEKNE-E 934
Query: 308 HEEDGDHDNKVESEGEAEGMADANDVEGDGA 338
++++ ++D +SE E ++ D G+
Sbjct: 935 NDDEPENDIPEKSEDETPQKSEREDKSDSGS 1027
>BQ750916 weakly similar to SP|P03186|TEGU Large tegument protein. [strain
B95-8 Human herpesvirus 4] {Epstein-barr virus},
partial (0%)
Length = 703
Score = 32.0 bits (71), Expect = 0.75
Identities = 31/107 (28%), Positives = 42/107 (38%), Gaps = 23/107 (21%)
Frame = +2
Query: 254 NRREEQICGVAGGENDDESDGSPHRSS------DDSENASENGD-------VSGTESA-- 298
NR+ + GG D DGS +S DDS N + +G G+ +A
Sbjct: 188 NRKSRNLRSRKGGPRDAREDGSVRNASRKSRLNDDSTNGARDGQRNRPGSHKGGSRNARE 367
Query: 299 DGEECSREE--------HEEDGDHDNKVESEGEAEGMADANDVEGDG 337
DG E R++ H DG + + G AEG A N E G
Sbjct: 368 DGRERRRQQGRQGRLKHHRTDGSGNRESGDLGAAEGGA-GNPAEDGG 505
>TC89226 weakly similar to GP|9759519|dbj|BAB10985.1
gb|AAF16763.1~gene_id:MLN1.10~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (5%)
Length = 1822
Score = 32.0 bits (71), Expect = 0.75
Identities = 43/221 (19%), Positives = 85/221 (38%), Gaps = 9/221 (4%)
Frame = +1
Query: 111 SIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQS---NGRTNINNASGLAATLSRPGYVYR 167
S P ++ + +K+ + Q + +S R S NG N L+ + + G
Sbjct: 1075 SAPLKIVQSSKKRRRLNQDKGRDKKLSKRTSHSKRDNGHHNFEVTENLSQRIKQQG---- 1242
Query: 168 EGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEED 227
+G G + R A+ + + H+ S RE+ + D++++
Sbjct: 1243 QGSQGQTGGRGPRTIRKRREEKRAVEDLSLGHKAANNSSN---IGREQSRILDE-DWDDE 1410
Query: 228 NFAVYANAGLEAVHKGKDCNTSQHYQNRR----EEQICGVAGGENDDES--DGSPHRSSD 281
+ +EA ++ N + ++ + V G+ + E +G+P+R +
Sbjct: 1411 KASPMRPIQMEAADMSSSSEEIEYDDNAQAVESDDNVEAVEYGQGNWEIGFNGTPNRWNR 1590
Query: 282 DSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEG 322
D SE DV +E + E EE+G + V S+G
Sbjct: 1591 DLVGMSEEEDVEASEDDNDNGIEGNEEEEEGS-EADVMSQG 1710
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,714,151
Number of Sequences: 36976
Number of extensions: 232723
Number of successful extensions: 1161
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 1104
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1133
length of query: 637
length of database: 9,014,727
effective HSP length: 102
effective length of query: 535
effective length of database: 5,243,175
effective search space: 2805098625
effective search space used: 2805098625
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146588.5


