

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146588.2 + phase: 0
(305 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
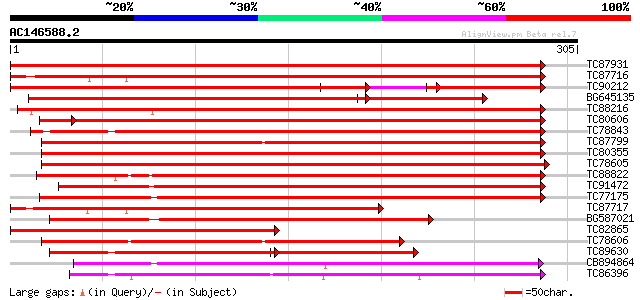
Score E
Sequences producing significant alignments: (bits) Value
TC87931 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Pru... 377 e-105
TC87716 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Pru... 356 5e-99
TC90212 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Pru... 243 8e-90
BG645135 weakly similar to PIR|D84713|D84 probable dioxygenase [... 230 1e-82
TC88216 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Pru... 288 2e-78
TC80606 similar to GP|5031283|gb|AAD38147.1| unknown {Prunus arm... 272 3e-76
TC78843 weakly similar to GP|8843897|dbj|BAA97423.1 1-aminocyclo... 274 4e-74
TC87799 weakly similar to PIR|C84713|C84713 probable dioxygenase... 269 9e-73
TC80355 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Pru... 259 9e-70
TC78605 similar to GP|5031283|gb|AAD38147.1| unknown {Prunus arm... 258 2e-69
TC88822 weakly similar to PIR|T06406|T06406 ripening protein E8 ... 243 9e-65
TC91472 weakly similar to PIR|S59548|S59548 1-aminocyclopropane-... 234 2e-62
TC77175 weakly similar to PIR|C84713|C84713 probable dioxygenase... 219 8e-58
TC87717 weakly similar to PIR|D86201|D86201 protein F12K11.6 [im... 219 1e-57
BG587021 weakly similar to PIR|D86201|D8 protein F12K11.6 [impor... 211 4e-55
TC82865 weakly similar to PIR|D86201|D86201 protein F12K11.6 [im... 194 3e-50
TC78606 similar to GP|5031283|gb|AAD38147.1| unknown {Prunus arm... 182 1e-46
TC89630 similar to PIR|D86201|D86201 protein F12K11.6 [imported]... 115 9e-43
CB894864 similar to PIR|D86201|D86 protein F12K11.6 [imported] -... 155 1e-38
TC86396 similar to GP|10178137|dbj|BAB11549. leucoanthocyanidin ... 152 2e-37
>TC87931 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Prunus
armeniaca}, partial (59%)
Length = 1323
Score = 377 bits (968), Expect = e-105
Identities = 178/288 (61%), Positives = 229/288 (78%)
Frame = +1
Query: 1 MEVKNSNLSHENDDSTYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPI 60
M V+NSN E+ D TYDR E+K FDDSK+GV+GL++ GV+KIPRMFY+ ++ I ++
Sbjct: 82 MVVENSNQLKESSDFTYDREAEVKAFDDSKVGVKGLVDSGVSKIPRMFYTRKLEISESTA 261
Query: 61 KNSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRR 120
+S L++PI+DLKDIHI+P++RVEVI+QIR+AC EWGFFQVINH IPI VLDE IDGI R
Sbjct: 262 SDSKLSIPIVDLKDIHINPAQRVEVIDQIRSACHEWGFFQVINHEIPIIVLDEMIDGICR 441
Query: 121 FHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFR 180
FHEQD +VRK+FY RD+KKK+ Y S L+ + ANWRD+ G +AP+P +E+P++ R
Sbjct: 442 FHEQDADVRKEFYTRDLKKKVSYYSNVRLFNGQGANWRDTFGFAIAPDPFNPDEVPQICR 621
Query: 181 DIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGA 240
DI++EYS+K+ LG TI EL SE LGL+PSYLKE EG+F GH+YP CPEPELTMG+
Sbjct: 622 DIVIEYSQKIKNLGFTIFELLSEALGLNPSYLKEFGCAEGLFTLGHFYPTCPEPELTMGS 801
Query: 241 STHTDPAFMTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+ HTD FMT++LQ+QL GLQVL +++W NV PVHGALVVNIGDLLQ+
Sbjct: 802 TAHTDSTFMTLLLQDQLGGLQVLHEDKWVNVPPVHGALVVNIGDLLQL 945
>TC87716 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Prunus
armeniaca}, partial (66%)
Length = 1297
Score = 356 bits (914), Expect = 5e-99
Identities = 179/292 (61%), Positives = 219/292 (74%), Gaps = 4/292 (1%)
Frame = +3
Query: 1 MEVKNSNLSHENDDSTYDRNDELKVFDDSKLGVRGLMERGV--TKIPRMFYSGEVNIIKN 58
ME++ SN S STYDR E+K F++SK GV+GL+E G+ TKIPRMF+S +N+ +
Sbjct: 15 MELEASNGS----SSTYDRTAEVKAFEESKTGVKGLLESGIWTTKIPRMFHSPNLNLNSD 182
Query: 59 PIK--NSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETID 116
+S +VPIIDL+DI+ DP EV+ +IR+ACKEWGFFQVINHGIP+ VLDE I
Sbjct: 183 HESEASSKFSVPIIDLQDINTDPCLHAEVLEKIRSACKEWGFFQVINHGIPVIVLDEMIS 362
Query: 117 GIRRFHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELP 176
GIRRFHEQD + RK FY RD KK+ Y S SL+ D +ANWRDS+ F++PNPP EE+P
Sbjct: 363 GIRRFHEQDADARKPFYTRDKSKKVRYFSNGSLFTDPAANWRDSLSFFVSPNPPDPEEIP 542
Query: 177 EVFRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPEL 236
+V RDI++EYSKKV LG TI ELFSE LGLHPSYLK+ G F+ HYYPPCPEPEL
Sbjct: 543 QVCRDIVIEYSKKVRALGLTIFELFSEALGLHPSYLKDLALDYGQFLLCHYYPPCPEPEL 722
Query: 237 TMGASTHTDPAFMTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
TMG S HTD FMTI+LQ+Q+ GLQVL NQW +V P+HG+LVVNIGDLLQ+
Sbjct: 723 TMGTSKHTDIDFMTILLQDQIGGLQVLHQNQWVHVPPLHGSLVVNIGDLLQL 878
>TC90212 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Prunus
armeniaca}, partial (70%)
Length = 1335
Score = 243 bits (621), Expect(2) = 8e-90
Identities = 115/194 (59%), Positives = 153/194 (78%)
Frame = +1
Query: 1 MEVKNSNLSHENDDSTYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPI 60
M VKN+N E DSTYDR E+K FDDS+ GV GL+E GV+KIPR F++G+++I +N
Sbjct: 43 MVVKNTNQLEEATDSTYDRKAEVKAFDDSRAGVNGLVESGVSKIPRFFHAGKLDIGENST 222
Query: 61 KNSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRR 120
+S L+VPI+DLKDIH +P++RV+VI+QIR+AC EWGFFQVINH IPI VLDE IDGIR
Sbjct: 223 CDSKLSVPIVDLKDIHNNPAQRVDVIHQIRSACHEWGFFQVINHEIPITVLDEMIDGIRS 402
Query: 121 FHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFR 180
FHEQD +VRK+FY RD+KKK++Y S L+ ++ANWRD+ G +AP+P K EE+P + R
Sbjct: 403 FHEQDADVRKEFYTRDLKKKVMYYSNVHLFSGQAANWRDTFGFAVAPHPFKPEEVPPICR 582
Query: 181 DIIMEYSKKVATLG 194
DI++EYS+K+ +G
Sbjct: 583 DIVIEYSQKIKGVG 624
Score = 104 bits (260), Expect(2) = 8e-90
Identities = 46/64 (71%), Positives = 54/64 (83%)
Frame = +3
Query: 225 GHYYPPCPEPELTMGASTHTDPAFMTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGD 284
GH YPPCPEPELTMG + HTD FMT++LQ+QL GLQVL ++W NV PVHGALVVN+GD
Sbjct: 717 GHCYPPCPEPELTMGTTKHTDSNFMTLLLQDQLGGLQVLHKDKWVNVPPVHGALVVNVGD 896
Query: 285 LLQV 288
LLQ+
Sbjct: 897 LLQL 908
Score = 45.1 bits (105), Expect = 3e-05
Identities = 28/65 (43%), Positives = 35/65 (53%)
Frame = +2
Query: 168 NPPKHEELPEVFRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHY 227
NP K + E+ + + +K LG TI EL SE LGL+ SYLKE N EG+FI G
Sbjct: 554 NPRKCHQYAEI---L*LNIHRK*RELGFTIFELLSEALGLNSSYLKELNCAEGLFIFGSL 724
Query: 228 YPPCP 232
P
Sbjct: 725 LSTVP 739
>BG645135 weakly similar to PIR|D84713|D84 probable dioxygenase [imported] -
Arabidopsis thaliana, partial (44%)
Length = 812
Score = 230 bits (586), Expect(2) = 1e-82
Identities = 108/184 (58%), Positives = 147/184 (79%)
Frame = +1
Query: 11 ENDDSTYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNVPII 70
E+ +STYDR E+K FDDSK GVRGL+E GV+KIPR+F++G+++I +N +S L+VPI+
Sbjct: 7 ESTNSTYDRKTEVKNFDDSKAGVRGLVEHGVSKIPRIFHTGKLDIGENSASDSKLSVPIV 186
Query: 71 DLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRK 130
DLKDIHID ++RVEVI QI++AC GFFQVINH IPIDVLDE I GIR FHEQD +VRK
Sbjct: 187 DLKDIHIDQAQRVEVIEQIQSACHVGGFFQVINHDIPIDVLDEMIHGIRSFHEQDVDVRK 366
Query: 131 QFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKV 190
+FY RD+KKK++Y S +L+ ++ANWRD++G +AP+P K ELP + RD +++YS+K+
Sbjct: 367 EFYTRDLKKKVMYFSNGTLFSGQAANWRDTIGFAVAPHPFKPNELPSICRDSVIKYSQKI 546
Query: 191 ATLG 194
+G
Sbjct: 547 RDVG 558
Score = 94.4 bits (233), Expect(2) = 1e-82
Identities = 45/70 (64%), Positives = 52/70 (74%)
Frame = +2
Query: 188 KKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPA 247
KK LG I EL SE LGL YLKE N EG+FIQGHYYP CPEPELTMG + HTD +
Sbjct: 539 KK*GMLGFIIFELLSEALGLDRDYLKELNCCEGLFIQGHYYPACPEPELTMGTAKHTDTS 718
Query: 248 FMTIVLQEQL 257
F+T++LQ+QL
Sbjct: 719 FITLLLQDQL 748
>TC88216 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Prunus
armeniaca}, partial (68%)
Length = 1508
Score = 288 bits (736), Expect = 2e-78
Identities = 137/289 (47%), Positives = 200/289 (68%), Gaps = 5/289 (1%)
Frame = +2
Query: 5 NSNLSH--ENDDSTYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKN 62
N+NL + + YDR ELK+ D+SK GV+GL++ G+TK+P++F +++ N +
Sbjct: 164 NTNLEEIKQEHEHVYDRQKELKLLDESKEGVKGLVDAGLTKVPKIFIHDKIHEHNNKQTS 343
Query: 63 SM-LNVPIIDLKDI--HIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIR 119
S L++PIID + + S R+E+I +++ A ++WGFFQV+NHGIP VLDE IDG+
Sbjct: 344 STNLSIPIIDFGPLFTNTSSSSRLEIIEKVKHASEKWGFFQVVNHGIPSTVLDEMIDGVV 523
Query: 120 RFHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVF 179
RFHEQD E++K+FY+RD+ K+ + + LY + NWRDS+ C M P P ++LP V
Sbjct: 524 RFHEQDTEMKKKFYSRDITKRAYFNTNFDLYVTPAVNWRDSLSCVMGPQPLDPQDLPTVC 703
Query: 180 RDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMG 239
RDI ++YS V +G + EL SE LGL+ +YLK+ + EG+F+ HYYPPCPEPELT G
Sbjct: 704 RDITVKYSDYVNKVGMILLELLSEALGLNSNYLKDIDCAEGLFLISHYYPPCPEPELTFG 883
Query: 240 ASTHTDPAFMTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
S H+D +F T++LQ+QL GLQV NQW +V P+ GALV+N+GD++Q+
Sbjct: 884 TSAHSDSSFFTVLLQDQLGGLQVFHGNQWVDVTPIPGALVINLGDMMQL 1030
>TC80606 similar to GP|5031283|gb|AAD38147.1| unknown {Prunus armeniaca},
partial (58%)
Length = 890
Score = 272 bits (695), Expect(2) = 3e-76
Identities = 135/256 (52%), Positives = 176/256 (68%), Gaps = 1/256 (0%)
Frame = +2
Query: 34 RGLMERGVTKIPRMFYSGEVNIIKNPIKNS-MLNVPIIDLKDIHIDPSRRVEVINQIRTA 92
R L+ VT IPRMF+ + NS L +P IDL DIH DP+RR V+ +IR A
Sbjct: 98 RTLLMXSVTSIPRMFHHEFDKDSTSSSSNSHKLVIPSIDLVDIHQDPTRRKIVVEKIREA 277
Query: 93 CKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRD 152
+ WGFFQ++NHGI + VLDE +G+ RF EQD EV+++ Y RD K +VY S LY
Sbjct: 278 SETWGFFQIVNHGIEVSVLDEMKNGVVRFFEQDSEVKRELYTRDPVKPLVYNSNFDLYSS 457
Query: 153 KSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTISELFSETLGLHPSYL 212
+A WRD+ CFMAPN P E+LP V RDI++EY+K+V LG+ + EL SE LGL P+YL
Sbjct: 458 PAATWRDTFYCFMAPNSPNPEDLPSVCRDIMLEYTKQVMKLGNLLFELLSEALGLDPNYL 637
Query: 213 KERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQLAGLQVLRDNQWFNVA 272
E EG+ + HYYP CPEPELT+G + HTD F+T++LQ+ + GLQVLR+N W +V+
Sbjct: 638 NEMRCNEGLALVCHYYPSCPEPELTLGITKHTDNDFITVLLQDHIGGLQVLRENSWVDVS 817
Query: 273 PVHGALVVNIGDLLQV 288
PV GALV+NIGDLLQ+
Sbjct: 818 PVPGALVINIGDLLQL 865
Score = 30.8 bits (68), Expect(2) = 3e-76
Identities = 13/20 (65%), Positives = 16/20 (80%)
Frame = +3
Query: 17 YDRNDELKVFDDSKLGVRGL 36
YDR ELK FD++K GV+GL
Sbjct: 45 YDRASELKAFDETKDGVKGL 104
>TC78843 weakly similar to GP|8843897|dbj|BAA97423.1
1-aminocyclopropane-1-carboxylate oxidase {Arabidopsis
thaliana}, partial (70%)
Length = 1240
Score = 274 bits (700), Expect = 4e-74
Identities = 135/280 (48%), Positives = 194/280 (69%), Gaps = 3/280 (1%)
Frame = +2
Query: 12 NDDSTYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNV-PII 70
N DST++ E K FD++K GV+GL++ GV KIP +F+ K I N+ NV P+I
Sbjct: 53 NLDSTFN---ERKAFDETKAGVKGLVDAGVEKIPSLFHHQPD---KYEIANNTSNVIPVI 214
Query: 71 DLKDI-HIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVR 129
DL DI + DPS ++ +++ AC+ GFFQV+NHGIP+ VL+E DG++RF+EQD E +
Sbjct: 215 DLDDIDNKDPSIHQGIVEKVKEACETLGFFQVVNHGIPLSVLEELNDGVKRFYEQDTEAK 394
Query: 130 KQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAP-NPPKHEELPEVFRDIIMEYSK 188
K FY RDM++ +Y S +Y + NW+DS GC +AP + K EE P V RDI++ Y K
Sbjct: 395 KSFYTRDMQRSFIYNSNVDIYSSPALNWKDSFGCNLAPPDTLKPEEFPVVCRDILLRYGK 574
Query: 189 KVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAF 248
+ LG+ + EL SE LGL+P++LK+ + EG+ HYYPPCPEPELT+G + H+D F
Sbjct: 575 HMMNLGTLLFELLSEALGLNPNHLKDMDCAEGLIALCHYYPPCPEPELTVGTTKHSDNDF 754
Query: 249 MTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+T++LQ+ + GLQVL D++W ++ PV GAL+VN+GDLLQ+
Sbjct: 755 LTVLLQDHIGGLQVLYDDKWIDITPVPGALIVNVGDLLQL 874
>TC87799 weakly similar to PIR|C84713|C84713 probable dioxygenase [imported]
- Arabidopsis thaliana, partial (65%)
Length = 1398
Score = 269 bits (688), Expect = 9e-73
Identities = 131/271 (48%), Positives = 181/271 (66%)
Frame = +1
Query: 18 DRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNVPIIDLKDIHI 77
+R ELK FD++KLGV+GL++ G+TKIPR+FY + K VP+IDL +I
Sbjct: 85 ERVKELKAFDETKLGVKGLVDAGITKIPRIFYQPRDSTKKASELGHTTIVPVIDLANIEK 264
Query: 78 DPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDM 137
DP R V+ IR A + +GFFQ++NHGIP+ L+E DG++ F EQD EV+K+FY R+
Sbjct: 265 DPFARKRVVESIRDASETFGFFQIVNHGIPVSTLEEMKDGVKSFFEQDSEVKKEFYTRE- 441
Query: 138 KKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTI 197
K ++ S +LY +A W+DS C MAP PPK E+LP V RDI++EY +V +G+ +
Sbjct: 442 HKPFMHYSNIALYTAPAATWKDSFLCNMAPIPPKPEDLPVVCRDILLEYLSQVKKVGTLL 621
Query: 198 SELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQL 257
EL SE LGL+ +YL + EG + GHYYPPCPEPELT+G S H D F+T++LQ+ +
Sbjct: 622 FELLSEALGLNSTYLTDIGCAEGFYAFGHYYPPCPEPELTLGTSKHVDVVFLTLLLQDHV 801
Query: 258 AGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
GLQ L + W +V P+ AL+VNIGD LQ+
Sbjct: 802 GGLQYLHRDMWIDVPPMPEALIVNIGDFLQL 894
>TC80355 weakly similar to GP|5031283|gb|AAD38147.1| unknown {Prunus
armeniaca}, partial (61%)
Length = 878
Score = 259 bits (662), Expect = 9e-70
Identities = 127/273 (46%), Positives = 180/273 (65%), Gaps = 2/273 (0%)
Frame = +2
Query: 18 DRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNV-PIIDLKDIH 76
+R ELK FDD+K GV+GL+++GV KIP +F+ K N+ ++ PIID +
Sbjct: 50 NRLHELKAFDDTKTGVKGLVDKGVVKIPTLFHHPLDKFGKATNSNNTQHIIPIIDFANFG 229
Query: 77 IDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRD 136
DP R E+ ++IR AC+ WGFFQV+NHGIP++VL++ +G+ RF EQD EV+K+ Y+RD
Sbjct: 230 NDPKTRHEITSKIREACETWGFFQVVNHGIPLNVLEDMRNGVIRFFEQDVEVKKELYSRD 409
Query: 137 MKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGST 196
+ Y S +LY + NWRD+ C +APN PK ++LP V RD ++EY + V LG T
Sbjct: 410 PMRPFSYNSNFNLYSSSALNWRDTFVCSLAPNAPKPQDLPLVCRDELLEYGEYVTKLGMT 589
Query: 197 ISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIV-LQE 255
+ EL SE LGLHP++LK+ +GI HYYP CPEPELTMG + H+D F+T++ +
Sbjct: 590 LFELLSEALGLHPNHLKDMGCAKGILSLSHYYPACPEPELTMGTTKHSDSCFLTLLPSDD 769
Query: 256 QLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+ GLQV N+W ++ P+ A VN GDLLQ+
Sbjct: 770 HIGGLQVFHQNKWIDIPPIPDAPGVNGGDLLQL 868
>TC78605 similar to GP|5031283|gb|AAD38147.1| unknown {Prunus armeniaca},
partial (58%)
Length = 961
Score = 258 bits (660), Expect = 2e-69
Identities = 126/273 (46%), Positives = 175/273 (63%)
Frame = +3
Query: 18 DRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNVPIIDLKDIHI 77
+R ELK FD++KLGVRGL++ G+TK+PR+FY + K +P+IDL +I
Sbjct: 90 ERVKELKAFDETKLGVRGLVDAGITKVPRIFYQPPDSTKKASESGDTTTIPVIDLANILE 269
Query: 78 DPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDM 137
DP R V+ +R A + +GFFQ++NHGIP+ L+E DG+ F EQD EV+K++Y R+
Sbjct: 270 DPCARKRVVESVRDASEIFGFFQIVNHGIPVSTLNEMKDGVVSFFEQDSEVKKKYYTRER 449
Query: 138 KKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTI 197
K + + LY W+D+ C APNPPK E+LP V R+I++EY V +G+ +
Sbjct: 450 KPFVYNSNYNLLYTSDPITWKDTFLCNRAPNPPKPEDLPAVCRNILLEYLNHVMKVGTLV 629
Query: 198 SELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQL 257
EL SE LGL+P+YL + EG+ GHYYP CPEPELT+G H D F+T++LQ+ +
Sbjct: 630 FELLSEALGLNPTYLIDIGCAEGLSAFGHYYPSCPEPELTIGTVKHADIDFITVLLQDHI 809
Query: 258 AGLQVLRDNQWFNVAPVHGALVVNIGDLLQVNL 290
GLQVL + W +V P+ ALVVNIGD L V L
Sbjct: 810 GGLQVLHKDMWVDVPPIPEALVVNIGDFLPVYL 908
>TC88822 weakly similar to PIR|T06406|T06406 ripening protein E8 homolog -
tomato, partial (31%)
Length = 1112
Score = 243 bits (619), Expect = 9e-65
Identities = 120/276 (43%), Positives = 175/276 (62%), Gaps = 2/276 (0%)
Frame = +3
Query: 15 STYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNI--IKNPIKNSMLNVPIIDL 72
S YDR +K FD+SK+GV+GL + G+ IP++F + + IK+ K + ++P+IDL
Sbjct: 42 SGYDRVKAVKEFDESKMGVKGLSDSGIKTIPQIFIHPQQTLSDIKSTSKQTT-SIPLIDL 218
Query: 73 KDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQF 132
++ P+ R +I QI+ A K WGFFQ+INH + + LD TI+ I FH Q E + ++
Sbjct: 219 SNV-ASPNHRPNIIKQIKEAAKTWGFFQIINHNVNVSSLDNTINAIESFHNQPHETKSKY 395
Query: 133 YNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVAT 192
Y RD + ++Y S LYR +A+W DS+ +MAP PK EELPE+ R ++E+ A
Sbjct: 396 YKRDEGRGVMYASNNDLYRTNAASWHDSLQAWMAPEAPKAEELPEICRKEMVEWDFHAAG 575
Query: 193 LGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIV 252
+ + EL SE LGL LKE ++ E I GH YP CP+P+LTMG + HTDP +T++
Sbjct: 576 VAEVVFELLSEGLGLERGKLKELSFCETRVIVGHCYPYCPQPDLTMGITPHTDPCAITLL 755
Query: 253 LQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
LQ Q+ GLQV + +W +V PV G ++VNIGD LQ+
Sbjct: 756 LQNQVPGLQVKYEGEWVDVNPVQGGIIVNIGDFLQI 863
>TC91472 weakly similar to PIR|S59548|S59548
1-aminocyclopropane-1-carboxylate oxidase homolog (clone
2A6) - Arabidopsis thaliana, partial (47%)
Length = 967
Score = 234 bits (598), Expect = 2e-62
Identities = 113/263 (42%), Positives = 170/263 (63%), Gaps = 1/263 (0%)
Frame = +1
Query: 27 DDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSML-NVPIIDLKDIHIDPSRRVEV 85
D+SK GV+GL++ G+TK+PR F + ++ K+ +S L +VP+ID + RR+E+
Sbjct: 1 DESKTGVKGLLDSGITKLPRFFIHPKESLPKHDATSSCLFHVPVIDFTGY--EECRRLEI 174
Query: 86 INQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDMKKKIVYLS 145
+N+I+ A + WGFFQ++NH +P+ V+D+ + I+ FHEQ EV+K++Y+RD K K+ Y S
Sbjct: 175 LNEIKEASETWGFFQMVNHCVPVCVMDDMLKAIKEFHEQPSEVKKEWYSRDHKVKVRYFS 354
Query: 146 TTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTISELFSETL 205
L K+ANWRD++ P + P V R+ +++Y K + L +SEL SE L
Sbjct: 355 NGDLLVAKAANWRDTIIFDFHDGPLDPQAYPLVCREAVIKYMKHILKLKEILSELMSEAL 534
Query: 206 GLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQLAGLQVLRD 265
GL YL + + HYYP CPEPELT GA+ H+DP+ +TI+LQ+ + GLQVL
Sbjct: 535 GLKRDYLANIECMKSETVVCHYYPTCPEPELTFGATKHSDPSSLTILLQDTIGGLQVLHQ 714
Query: 266 NQWFNVAPVHGALVVNIGDLLQV 288
N W ++ P+HGALV NIGD +Q+
Sbjct: 715 NHWVDINPIHGALVANIGDFMQL 783
>TC77175 weakly similar to PIR|C84713|C84713 probable dioxygenase [imported]
- Arabidopsis thaliana, partial (40%)
Length = 1425
Score = 219 bits (559), Expect = 8e-58
Identities = 112/274 (40%), Positives = 167/274 (60%), Gaps = 2/274 (0%)
Frame = +3
Query: 17 YDRNDELKVFDDSKLGVRGLMERGVTKIPRMF-YSGEVNIIKNPIKN-SMLNVPIIDLKD 74
YDR +K FD++K GV+GL++ G+ IP F + E+ P + +P IDL
Sbjct: 153 YDRIKAVKEFDETKSGVKGLIDSGIKTIPSFFIHPPEILSDLTPRSDFPQPEIPTIDLSA 332
Query: 75 IHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYN 134
+H R V+ Q+R+A GFFQVINHG+ +++ I +++FHEQ + RK+ Y
Sbjct: 333 VH---HSRAAVVEQLRSAASTVGFFQVINHGVAPELMRSVIGAMKKFHEQPADERKKVYC 503
Query: 135 RDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLG 194
R+M + Y+S L+ K+A+WRD++ M P P + +E+PEV R +ME+ K+V +G
Sbjct: 504 REMGTGVSYISNVDLFASKAASWRDTLQIKMGPVPAEEKEIPEVCRKEVMEWDKEVVRVG 683
Query: 195 STISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQ 254
+ L SE LGL L E +G + GHYYP CP+P LT+G ++H DP +T++LQ
Sbjct: 684 DILLGLLSEGLGLGEERLTELGLSQGRVMVGHYYPFCPQPNLTVGLNSHADPGALTVLLQ 863
Query: 255 EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+ + GLQV + W NV P+ GALV+NIGDLLQ+
Sbjct: 864 DHIGGLQVRTQHGWINVKPLDGALVINIGDLLQI 965
>TC87717 weakly similar to PIR|D86201|D86201 protein F12K11.6 [imported] -
Arabidopsis thaliana, partial (6%)
Length = 653
Score = 219 bits (558), Expect = 1e-57
Identities = 113/210 (53%), Positives = 150/210 (70%), Gaps = 9/210 (4%)
Frame = +1
Query: 1 MEVKNSNLSHENDDSTYDRNDELKVFDDSKLGVRGLMERG--VTKIPRMFYSGEVNIIKN 58
ME++ SN S STY+R E+K FD+SK GV+GL+E G VTKIPRMF+ + ++ N
Sbjct: 31 MELEASNSS---TTSTYNRIAEVKAFDESKAGVKGLVESGTCVTKIPRMFHFPKSSLKNN 201
Query: 59 PIK-------NSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVL 111
+ +S L VPIIDL+D++ +P VEV+++IR+ACKEWGFFQVINHGIP+ VL
Sbjct: 202 THETILQIDSSSKLCVPIIDLQDMNTNPCLHVEVVDKIRSACKEWGFFQVINHGIPVSVL 381
Query: 112 DETIDGIRRFHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPK 171
DE IRRFHEQ+ + RK FY RD KK+ Y S +L+RD +ANWRD++ F +P+PP
Sbjct: 382 DEMTSAIRRFHEQEVDARKPFYTRDTSKKVRYFSNGTLFRDPAANWRDTIAFFTSPDPPN 561
Query: 172 HEELPEVFRDIIMEYSKKVATLGSTISELF 201
EE+P+V RDI++EYSKKV LG TI EL+
Sbjct: 562 PEEIPQVCRDIVIEYSKKVRALGLTIFELY 651
>BG587021 weakly similar to PIR|D86201|D8 protein F12K11.6 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 663
Score = 211 bits (536), Expect = 4e-55
Identities = 108/209 (51%), Positives = 144/209 (68%), Gaps = 2/209 (0%)
Frame = +2
Query: 22 ELKVFDDSKLGVRGLMERGVTKIPRMFYSG-EVNIIKNPI-KNSMLNVPIIDLKDIHIDP 79
+++ FDDSK GV+G+++ G+TKIP +F+ E + +P ++S +PIIDL+ I
Sbjct: 50 QVQAFDDSKTGVKGILDSGITKIPSIFHVNLEKSTQNSPTHESSFTTIPIIDLQHI---- 217
Query: 80 SRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDMKK 139
+RVE+++QI+ A K WGFFQVINHGIP+ VLDE I+GI RFHEQD E +K FY+RD K
Sbjct: 218 -KRVELVDQIKNASKNWGFFQVINHGIPLHVLDEMINGICRFHEQDAEAKKPFYSRDKDK 394
Query: 140 KIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTISE 199
K+ Y S L+RD +ANWRD++ PN P EELP V RDI+ EYSK+V LG TI E
Sbjct: 395 KVRYFSNGKLFRDMAANWRDTIAFAANPNLPNPEELPAVCRDIVAEYSKQVKALGVTIFE 574
Query: 200 LFSETLGLHPSYLKERNYFEGIFIQGHYY 228
L SE LGL+ SYLK + ++I G YY
Sbjct: 575 LLSEVLGLNLSYLK*MECAKALYIMGQYY 661
>TC82865 weakly similar to PIR|D86201|D86201 protein F12K11.6 [imported] -
Arabidopsis thaliana, partial (2%)
Length = 547
Score = 194 bits (494), Expect = 3e-50
Identities = 91/145 (62%), Positives = 116/145 (79%)
Frame = +2
Query: 1 MEVKNSNLSHENDDSTYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPI 60
M VKN+N E +STYDR E+K FDDSK+GV+GL+E GV+KIPRMF++G++ I ++
Sbjct: 107 MVVKNTNQLEETTNSTYDREAEVKAFDDSKVGVKGLVESGVSKIPRMFHTGKMEISEDSA 286
Query: 61 KNSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRR 120
S L++PIIDLKDIH +P+RRVEVI QI++AC EWGFFQ+INH IPI+VLDE IDGI +
Sbjct: 287 SESKLSIPIIDLKDIHNNPARRVEVIGQIQSACHEWGFFQIINHEIPINVLDEMIDGISK 466
Query: 121 FHEQDPEVRKQFYNRDMKKKIVYLS 145
FHEQD VRK FY RD+ KK++Y S
Sbjct: 467 FHEQDANVRKDFYTRDLNKKVMYYS 541
>TC78606 similar to GP|5031283|gb|AAD38147.1| unknown {Prunus armeniaca},
partial (43%)
Length = 630
Score = 182 bits (462), Expect = 1e-46
Identities = 91/195 (46%), Positives = 130/195 (66%)
Frame = +3
Query: 18 DRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNVPIIDLKDIHI 77
+R LK FD++KLGV+GL++ G+TKIPRMFY + ++ + +P IDL +I
Sbjct: 24 ERIKTLKAFDETKLGVKGLVDAGITKIPRMFYHPPDHTNESGDATNY-TIPFIDLANIDK 200
Query: 78 DPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDM 137
DP R V+ +R A + +GFFQ++NHGIP+ L+E DG+ F EQD EV+K+FY R+
Sbjct: 201 DPCVRKRVVESVRDASETFGFFQIVNHGIPVSTLNEMKDGVVSFFEQDSEVKKEFYTRE- 377
Query: 138 KKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTI 197
++ +Y S +LY +W+D+ C MAPNPPK E+LP V RDI++EY +V LG+ +
Sbjct: 378 QRPFMYNSNFNLYTSAPTSWKDTFLCNMAPNPPKPEDLPAVIRDILVEYLNQVMKLGTIL 557
Query: 198 SELFSETLGLHPSYL 212
EL SE LGL+PSYL
Sbjct: 558 LELLSEALGLNPSYL 602
>TC89630 similar to PIR|D86201|D86201 protein F12K11.6 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 680
Score = 115 bits (287), Expect(2) = 9e-43
Identities = 59/126 (46%), Positives = 83/126 (65%), Gaps = 2/126 (1%)
Frame = +3
Query: 22 ELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNV-PIIDLKDI-HIDP 79
E K FD++K V+G ++ G+ KIP F+ KN I + +V PIIDL DI + DP
Sbjct: 81 ERKAFDETKASVKGFVDEGIKKIPSFFHHQPD---KNEIAYNNCHVLPIIDLADIDNKDP 251
Query: 80 SRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDMKK 139
++I++++ AC+ WGFFQV+NHGIP+++L+E DGI+RFHEQDPEV+K Y D K
Sbjct: 252 IIHQKIIDKLKEACETWGFFQVVNHGIPLNILEELKDGIKRFHEQDPEVKKDLYTTDPKG 431
Query: 140 KIVYLS 145
Y S
Sbjct: 432 PFTYNS 449
Score = 76.3 bits (186), Expect(2) = 9e-43
Identities = 35/80 (43%), Positives = 52/80 (64%)
Frame = +1
Query: 141 IVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTISEL 200
++ + L + NWRD+ CF+AP+ PK E+ P V RDI++EYSK++ LG + EL
Sbjct: 439 LIIVILVGLSNSPALNWRDTFICFLAPDTPKPEDFPLVCRDILIEYSKQMMNLGFLLFEL 618
Query: 201 FSETLGLHPSYLKERNYFEG 220
SE LGL+P++LK+ EG
Sbjct: 619 LSEALGLNPNHLKDMGCVEG 678
>CB894864 similar to PIR|D86201|D86 protein F12K11.6 [imported] - Arabidopsis
thaliana, partial (4%)
Length = 786
Score = 155 bits (393), Expect = 1e-38
Identities = 96/265 (36%), Positives = 144/265 (54%), Gaps = 12/265 (4%)
Frame = +1
Query: 35 GLMERGVTKIPRMF-YSGEVNIIKNPIKN-SMLNVPIIDLKDIHIDPSRRVEVINQIRTA 92
GL++ G+ IP F + E+ P + +P IDL +H R V+ Q+R+A
Sbjct: 1 GLIDSGIKTIPSFFIHPPEILSDLTPRSDFPQPEIPTIDLSAVH---HSRAAVVEQLRSA 171
Query: 93 CKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEV-RKQFYNRDMKKKIVYLSTT-SLY 150
GFFQVINHG+ +++ I +++FHEQ + R+ R +YL+ SL
Sbjct: 172 ASTVGFFQVINHGVAPELMRSVIGAMKKFHEQPADRGRRCTVGRWELVFPIYLTLICSLL 351
Query: 151 RDKSANWRDSVGCFMAPN--------PPKHEELPEVFRDIIMEYSKKVATLGSTISELFS 202
+ + R ++ N P + +E+PEV R +ME+ K+V +G + L S
Sbjct: 352 KLPAGVTRFRFENYLRENSHY*DGTCPTEEKEIPEVCRKEVMEWDKEVVRVGDILLGLLS 531
Query: 203 ETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQLAGLQV 262
E LGL L E +G + GHYYP CP+P LT+G ++H DP +T++LQ+ + GLQV
Sbjct: 532 EGLGLGEERLTELGLSQGRVMVGHYYPFCPQPNLTVGLNSHADPGALTVLLQDHIGGLQV 711
Query: 263 LRDNQWFNVAPVHGALVVNIGDLLQ 287
+ W NV P+ GALV+NIGDLLQ
Sbjct: 712 RTQHGWINVKPLDGALVINIGDLLQ 786
>TC86396 similar to GP|10178137|dbj|BAB11549. leucoanthocyanidin
dioxygenase-like protein {Arabidopsis thaliana}, partial
(82%)
Length = 1326
Score = 152 bits (384), Expect = 2e-37
Identities = 95/273 (34%), Positives = 141/273 (50%), Gaps = 17/273 (6%)
Frame = +2
Query: 33 VRGLMERGVTKIPRMFYSGEVNIIKNPIKNSM--------LNVPIIDLKDIHI-DPSRRV 83
V+ L E G+T IP + + P K + +N+P+IDL+ + DP R
Sbjct: 137 VQALAESGLTSIPSCYIKPRS---QRPTKTTFATQNDHDHINIPVIDLEHLSSEDPVLRE 307
Query: 84 EVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRDMKKKIVY 143
V+ ++ AC+EWGFFQV+NHGI ++++ + R F EV+++F N + Y
Sbjct: 308 TVLKRVSEACREWGFFQVVNHGISHELMESAKEVWREFFNLPLEVKEEFANSPSTYE-GY 484
Query: 144 LSTTSLYRDKSANWRDSVGCFMAP----NPPKHEELPEVFRDIIMEYSKKVATLGSTISE 199
S + + +W D P N K P R II EY ++V LG + E
Sbjct: 485 GSRLGVKKGAILDWSDYFFLHSMPPSLRNQAKWPATPSSLRKIIAEYGEEVVKLGGRMLE 664
Query: 200 LFSETLGLHPSYLKERNYFE---GIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQ 256
L S LGL YL E G ++ ++YP CP+P+LT+G S H+DP MTI+L +
Sbjct: 665 LMSTNLGLKEDYLMNAFGGENELGACLRVNFYPKCPQPDLTLGLSPHSDPGGMTILLPDD 844
Query: 257 -LAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
++GLQV + N W V PV A ++NIGD +QV
Sbjct: 845 FVSGLQVRKGNDWITVKPVPNAFIINIGDQIQV 943
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.139 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,224,915
Number of Sequences: 36976
Number of extensions: 125070
Number of successful extensions: 877
Number of sequences better than 10.0: 168
Number of HSP's better than 10.0 without gapping: 734
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 738
length of query: 305
length of database: 9,014,727
effective HSP length: 96
effective length of query: 209
effective length of database: 5,465,031
effective search space: 1142191479
effective search space used: 1142191479
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146588.2


