

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
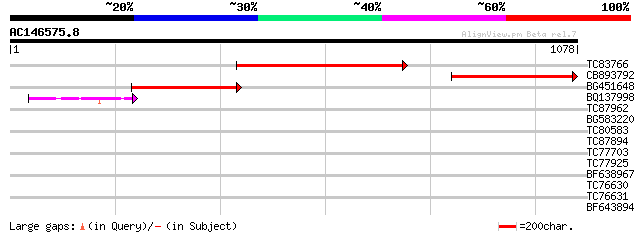
Query= AC146575.8 - phase: 0
(1078 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC83766 similar to PIR|T52638|T52638 exportin 1 [validated] - Ar... 647 0.0
CB893792 similar to PIR|T52638|T5 exportin 1 [validated] - Arabi... 480 e-135
BG451648 similar to PIR|T52638|T5 exportin 1 [validated] - Arabi... 411 e-115
BQ137998 GP|6165483|emb putative nuclear transport receptor {Sch... 43 7e-04
TC87962 similar to GP|14596039|gb|AAK68747.1 Unknown protein {Ar... 32 1.0
BG583220 similar to PIR|H86371|H863 hypothetical protein AAC9803... 32 1.7
TC80583 squalene monooxygenase 1 [Medicago truncatula] 30 3.8
TC87894 similar to GP|12005328|gb|AAG44394.1 unknown {Hevea bras... 30 5.0
TC77703 similar to GP|14334884|gb|AAK59620.1 unknown protein {Ar... 30 6.5
TC77925 similar to GP|18700163|gb|AAL77693.1 AT4g02720/T10P11_1 ... 30 6.5
BF638967 similar to GP|8777419|dbj mitotic checkpoint protein-li... 30 6.5
TC76630 homologue to GP|18033654|gb|AAL57201.1 putative nodule m... 29 8.6
TC76631 homologue to GP|18033654|gb|AAL57201.1 putative nodule m... 29 8.6
BF643894 29 8.6
>TC83766 similar to PIR|T52638|T52638 exportin 1 [validated] - Arabidopsis
thaliana, partial (30%)
Length = 981
Score = 647 bits (1670), Expect = 0.0
Identities = 325/326 (99%), Positives = 325/326 (99%)
Frame = +2
Query: 431 AKPEEVLIVEDENGNIVRETMKDSDVLVQYKIMRETLIYLSHLDHDDTEKQMLGKLSKQL 490
AKPEEVLIVEDENGNIVRETMKDSDVLVQYKIMRETLIYLSHLDHDDTEKQMLGKLSKQL
Sbjct: 2 AKPEEVLIVEDENGNIVRETMKDSDVLVQYKIMRETLIYLSHLDHDDTEKQMLGKLSKQL 181
Query: 491 SGKDWTWNNLNTLCWAIGSISGSMGEEQENRFLVMVIRDLLNLCEITKGKDNKAVIASNI 550
SGKDWTWNNLNTLCWAIGSISGSMGEEQENRFLVMVIRDLLNLCEITKGKDNKAVIASNI
Sbjct: 182 SGKDWTWNNLNTLCWAIGSISGSMGEEQENRFLVMVIRDLLNLCEITKGKDNKAVIASNI 361
Query: 551 MYVVGQYPRFLRAHWKFLKTVVNKLFEFMHETHPGVQDMACDTFLKIVQKCRRKFVITQV 610
MYVVGQYPRFLRAHWKFLKTVVNK FEFMHETHPGVQDMACDTFLKIVQKCRRKFVITQV
Sbjct: 362 MYVVGQYPRFLRAHWKFLKTVVNKWFEFMHETHPGVQDMACDTFLKIVQKCRRKFVITQV 541
Query: 611 GENEPFVSELLSGLSTTIADLEPHQIHTFYESVGSMIQAESDSQKRDEYLQRLMVLPNQK 670
GENEPFVSELLSGLSTTIADLEPHQIHTFYESVGSMIQAESDSQKRDEYLQRLMVLPNQK
Sbjct: 542 GENEPFVSELLSGLSTTIADLEPHQIHTFYESVGSMIQAESDSQKRDEYLQRLMVLPNQK 721
Query: 671 WMEIIGQARQNADFLKDQDVIRTVLNVLQTNTSVASSLGTYFLPQITLIFLDMLNVYRMY 730
WMEIIGQARQNADFLKDQDVIRTVLNVLQTNTSVASSLGTYFLPQITLIFLDMLNVYRMY
Sbjct: 722 WMEIIGQARQNADFLKDQDVIRTVLNVLQTNTSVASSLGTYFLPQITLIFLDMLNVYRMY 901
Query: 731 SELISKSIAEGTPFTSRTSYVKLLRS 756
SELISKSIAEGTPFTSRTSYVKLLRS
Sbjct: 902 SELISKSIAEGTPFTSRTSYVKLLRS 979
>CB893792 similar to PIR|T52638|T5 exportin 1 [validated] - Arabidopsis
thaliana, partial (22%)
Length = 873
Score = 480 bits (1235), Expect = e-135
Identities = 239/239 (100%), Positives = 239/239 (100%)
Frame = +3
Query: 840 ITKNFEDYPEHRLKFFSLLRAIATHCFPALICLSSQQLKFVMDSIIWAFRHTERNIAETG 899
ITKNFEDYPEHRLKFFSLLRAIATHCFPALICLSSQQLKFVMDSIIWAFRHTERNIAETG
Sbjct: 3 ITKNFEDYPEHRLKFFSLLRAIATHCFPALICLSSQQLKFVMDSIIWAFRHTERNIAETG 182
Query: 900 LNLLLEMLNKFQASEFCNQFYRTYFLTIEQEIFAVLTDTFHKPGFKLHVLVLQHLLCLAE 959
LNLLLEMLNKFQASEFCNQFYRTYFLTIEQEIFAVLTDTFHKPGFKLHVLVLQHLLCLAE
Sbjct: 183 LNLLLEMLNKFQASEFCNQFYRTYFLTIEQEIFAVLTDTFHKPGFKLHVLVLQHLLCLAE 362
Query: 960 SGALTEPLWDAATNSYPYPSNGAFVREYTIKLLSTSFPNMTAAEVTQFVNGLFESTNDLS 1019
SGALTEPLWDAATNSYPYPSNGAFVREYTIKLLSTSFPNMTAAEVTQFVNGLFESTNDLS
Sbjct: 363 SGALTEPLWDAATNSYPYPSNGAFVREYTIKLLSTSFPNMTAAEVTQFVNGLFESTNDLS 542
Query: 1020 TFKTHIRDFLIQSKEFSAQDNKDLYAEEAAAQRERERQRMLSIPGLIAPVELQDEMVDS 1078
TFKTHIRDFLIQSKEFSAQDNKDLYAEEAAAQRERERQRMLSIPGLIAPVELQDEMVDS
Sbjct: 543 TFKTHIRDFLIQSKEFSAQDNKDLYAEEAAAQRERERQRMLSIPGLIAPVELQDEMVDS 719
>BG451648 similar to PIR|T52638|T5 exportin 1 [validated] - Arabidopsis
thaliana, partial (17%)
Length = 634
Score = 411 bits (1057), Expect = e-115
Identities = 208/210 (99%), Positives = 208/210 (99%)
Frame = +3
Query: 232 IFESPLLETLLKFFTMQAYRNLTLQCLTEVASLQFGNFYDAQYVKMYNIFMVQLQSILPP 291
IFESPLLETLLKFFTMQAYRNLTLQCLTEVASLQFGNFYDAQYVKMYNIFMVQLQSILPP
Sbjct: 3 IFESPLLETLLKFFTMQAYRNLTLQCLTEVASLQFGNFYDAQYVKMYNIFMVQLQSILPP 182
Query: 292 TTNIPEAYAHGSSDEQAFIQNLALFFTLFFKVHIRILESTQENISALLLGLEYLINISYV 351
TTNIPEAYAHGSSDEQAFIQNLALFFTLFFKVHIRILESTQENISALLLGLEYLINISYV
Sbjct: 183 TTNIPEAYAHGSSDEQAFIQNLALFFTLFFKVHIRILESTQENISALLLGLEYLINISYV 362
Query: 352 DDTEVFKVCLDYWNTLVSELFQPHRSLENSAAAATNMMGSQVSLMPPGMVDGLGPQLLQR 411
DDTEVFKVCLDYWNTLVSELFQPHRSLENSAAAATNMMGSQVSLMPPGMVDGLGPQLLQR
Sbjct: 363 DDTEVFKVCLDYWNTLVSELFQPHRSLENSAAAATNMMGSQVSLMPPGMVDGLGPQLLQR 542
Query: 412 RQLYAGPVSKLRMLMICRMAKPEEVLIVED 441
RQLYAGPVSKL MLMICRMAKP EVLIVED
Sbjct: 543 RQLYAGPVSKLXMLMICRMAKPXEVLIVED 632
>BQ137998 GP|6165483|emb putative nuclear transport receptor
{Schizosaccharomyces pombe}, partial (11%)
Length = 656
Score = 42.7 bits (99), Expect = 7e-04
Identities = 41/212 (19%), Positives = 90/212 (42%), Gaps = 6/212 (2%)
Frame = +3
Query: 37 ADQILRELQNNPDMWLQVMHILQNTQ-NLNTKFFALQVLEGVIKYRWNALPVEQRDGMKN 95
A + L + Q + + W + +L+++ + K FA L+G I Y + +P EQ
Sbjct: 6 AHEFLEKFQKSQEAWTTTLAMLESSSVDAAAKLFAATTLKGKIVYDLHQIPREQ------ 167
Query: 96 FISDVIVQLSRNEASFRTERLYVNKLNIILVQILKHEWPARWRNFIPDLVSAAKTSETIC 155
+ ++ + RN F + + + L + W++ + D+V A +
Sbjct: 168 -LPELRASIMRNLTVFHAGPKPIRLTLCVCLANLAIQM-TEWKDVLTDIVHALGSDPATL 341
Query: 156 ENCMAILKLLSEEV-----FDFSRGEMTQLKIKELKQSLNSEFQLIHELCLFVLSVSQRT 210
+ L++L EEV + E+T ++ ++ + +L+ + ++
Sbjct: 342 PCVLDFLRVLPEEVTHGRKIALTEHELTMRTVELIEDNAEQALKLLVTYATSSPAAARNP 521
Query: 211 ELIRATLSTLHAFLSWIPLGYIFESPLLETLL 242
+L L+ + +++ IPL I SPLL ++
Sbjct: 522 QL----LNCITSWMREIPLDSILNSPLLAVII 605
>TC87962 similar to GP|14596039|gb|AAK68747.1 Unknown protein {Arabidopsis
thaliana}, partial (90%)
Length = 1268
Score = 32.3 bits (72), Expect = 1.0
Identities = 14/38 (36%), Positives = 25/38 (64%)
Frame = -1
Query: 847 YPEHRLKFFSLLRAIATHCFPALICLSSQQLKFVMDSI 884
+P R+KF +LL+ + HC +LIC+S++ +M +I
Sbjct: 899 FPRDRIKFTNLLKMVKNHC--SLICMSTRSNDRIMKNI 792
>BG583220 similar to PIR|H86371|H863 hypothetical protein AAC98032.1
[imported] - Arabidopsis thaliana, partial (21%)
Length = 634
Score = 31.6 bits (70), Expect = 1.7
Identities = 39/160 (24%), Positives = 67/160 (41%), Gaps = 6/160 (3%)
Frame = +3
Query: 316 FFTLFFKVHIRILES---TQENISALLLGLEYLINISYVDDTEVFKVCLDYWNTLVSELF 372
F L F VH ILES +++S + + + +LI DD ++++S +
Sbjct: 54 FKLLVFAVHAFILESGFVRVDHVSGMAISISHLI-----DDMSS--------SSMISLRY 194
Query: 373 QPHRSLENSAAAATNMMGSQVSLMPPGMVDGLGPQLLQRRQLYAGPVSKLRMLMI--CRM 430
L N ++ + N+ + LG + LY S + M+ + CR
Sbjct: 195 TLPEILTNGSSHSLNLK-----------IQTLGHFVNVYGSLYDDVGSSVHMVYLDKCRF 341
Query: 431 AKPEEVLIVEDENGNIVRETMKDSD-VLVQYKIMRETLIY 469
AKP E L++ + N D D VL +KI+ + L +
Sbjct: 342 AKPLEFLLMSNSESNACVNVDGDKDEVLELWKIVNDRLAF 461
>TC80583 squalene monooxygenase 1 [Medicago truncatula]
Length = 1830
Score = 30.4 bits (67), Expect = 3.8
Identities = 15/32 (46%), Positives = 20/32 (61%)
Frame = +1
Query: 177 MTQLKIKELKQSLNSEFQLIHELCLFVLSVSQ 208
M QLK+ K+ + FQ +H L LFV+ VSQ
Sbjct: 673 MVQLKVYSTKRKMLRNFQHVHLLPLFVMVVSQ 768
>TC87894 similar to GP|12005328|gb|AAG44394.1 unknown {Hevea brasiliensis},
partial (26%)
Length = 997
Score = 30.0 bits (66), Expect = 5.0
Identities = 19/59 (32%), Positives = 30/59 (50%), Gaps = 5/59 (8%)
Frame = -1
Query: 826 PHIFEAVFQCTLEMITKNFEDYP-----EHRLKFFSLLRAIATHCFPALICLSSQQLKF 879
P +F + TL + ++YP EH + +SLL++I HCF L + QL+F
Sbjct: 880 PTVFPGIISTTLI----HHQEYPYHNKLEHSQRAYSLLKSILNHCF-----LETTQLEF 731
>TC77703 similar to GP|14334884|gb|AAK59620.1 unknown protein {Arabidopsis
thaliana}, partial (77%)
Length = 2410
Score = 29.6 bits (65), Expect = 6.5
Identities = 47/208 (22%), Positives = 88/208 (41%), Gaps = 22/208 (10%)
Frame = +3
Query: 708 LGTYFLPQITLIFLDM-----LNVYRMYSELISKSIAEGTPFTSRTSYVKLLRSVKRETL 762
LG F+ + LD ++VYRM ELI S+A+ + ++Y++ L + T
Sbjct: 915 LGVDFMSENVRAILDQAGFNEVDVYRMSDELIGCSLADA---AATSTYMEYLEPASKST- 1082
Query: 763 KLIETFLDKAEDQPQIGKQFVPP-------MMDPVLGDYARNVPDARESEVLSLFATIIN 815
L +++ + + VP ++ +L +A+ VPD LS++ +
Sbjct: 1083 SLHVIYINTKLETKAYAHELVPTITCTSSNVIQTILQAFAQ-VPD------LSIWYGPDS 1241
Query: 816 KYKASMIEDIPHIFEAVFQCTLEMITKNFEDYPEH----------RLKFFSLLRAIATHC 865
A+++E +F+ + T E + +PEH RL +F I H
Sbjct: 1242 YMGANIVE----LFQQMTVMTDEEVA---AIHPEHNVDSIKSLLPRLHYFQDGSCIVHHL 1400
Query: 866 FPALICLSSQQLKFVMDSIIWAFRHTER 893
F + +++ VM S++ R+ ER
Sbjct: 1401 FGNEVVEKIKEIVLVMHSLLPILRYLER 1484
>TC77925 similar to GP|18700163|gb|AAL77693.1 AT4g02720/T10P11_1
{Arabidopsis thaliana}, partial (47%)
Length = 1538
Score = 29.6 bits (65), Expect = 6.5
Identities = 29/97 (29%), Positives = 39/97 (39%), Gaps = 10/97 (10%)
Frame = -1
Query: 848 PEHRLKFFSLLRAIATHC--FPALICLSSQQLK------FVMDSIIWAFRHTERNIAETG 899
P+ R++FF LR C F LI L L+ + I+ A R T RNI T
Sbjct: 593 PQFRIRFFRFLRFRLRFCFLFNLLILLHINYLRNFFRFTIIFKIIVLASRRTRRNIPNTV 414
Query: 900 LNLLLEMLNKFQASEFCNQFYRTY--FLTIEQEIFAV 934
L +L A F Q + + F + I AV
Sbjct: 413 LQRFPHLLPLQSAILFITQSLQFFIRFRLVTPRIVAV 303
>BF638967 similar to GP|8777419|dbj mitotic checkpoint protein-like
{Arabidopsis thaliana}, partial (26%)
Length = 675
Score = 29.6 bits (65), Expect = 6.5
Identities = 35/158 (22%), Positives = 69/158 (43%), Gaps = 16/158 (10%)
Frame = +3
Query: 628 IADLEPHQIHTFYESVGSMIQAESDSQKRDEY-------LQRLMVLPNQKWMEI------ 674
+A+L+ + S + + ES K+DEY L L V+ N++ EI
Sbjct: 45 LANLKNGEAGDESSSANPIQELESSLMKKDEYIKELESTLNELRVVNNRQHEEIKILNEK 224
Query: 675 -IGQARQNADFLKDQDVIRTVLNVLQTNTSVA--SSLGTYFLPQITLIFLDMLNVYRMYS 731
+AR+ ++ D +R+ +++L+ SS T L + + +D N +
Sbjct: 225 LHNEARRIKSLERESDRLRSEISLLEAKLGHGDFSSANTKVLRMVNTLSVD--NEAKQTI 398
Query: 732 ELISKSIAEGTPFTSRTSYVKLLRSVKRETLKLIETFL 769
E++ + + + V+ L+S ET KL+E ++
Sbjct: 399 EVLQNELQK---TKEKLKAVEELKSQSGETGKLVENYI 503
>TC76630 homologue to GP|18033654|gb|AAL57201.1 putative nodule membrane
protein {Medicago sativa}, complete
Length = 2605
Score = 29.3 bits (64), Expect = 8.6
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = -1
Query: 896 AETGLNLLLEMLNKFQASEFCNQFYRTYFLTIEQ 929
A+TGLNLL + F S+ CN + ++F +E+
Sbjct: 1657 AQTGLNLLYHSPHAFTHSKLCNFPWNSHFFMMEE 1556
>TC76631 homologue to GP|18033654|gb|AAL57201.1 putative nodule membrane
protein {Medicago sativa}, partial (43%)
Length = 948
Score = 29.3 bits (64), Expect = 8.6
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = -1
Query: 896 AETGLNLLLEMLNKFQASEFCNQFYRTYFLTIEQ 929
A+TGLNLL + F S+ CN + ++F +E+
Sbjct: 681 AQTGLNLLYHSPHAFTHSKLCNFPWNSHFFMMEE 580
>BF643894
Length = 591
Score = 29.3 bits (64), Expect = 8.6
Identities = 13/36 (36%), Positives = 22/36 (61%), Gaps = 6/36 (16%)
Frame = +2
Query: 193 FQLIHELCLFVLS------VSQRTELIRATLSTLHA 222
FQ++HELC+F LS Q + ++ + +TLH+
Sbjct: 167 FQILHELCVFFLSQICSYITQQSNQKVQPSCNTLHS 274
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.136 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,974,473
Number of Sequences: 36976
Number of extensions: 405051
Number of successful extensions: 2133
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 2113
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2132
length of query: 1078
length of database: 9,014,727
effective HSP length: 106
effective length of query: 972
effective length of database: 5,095,271
effective search space: 4952603412
effective search space used: 4952603412
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146575.8


