

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146575.4 - phase: 0
(165 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
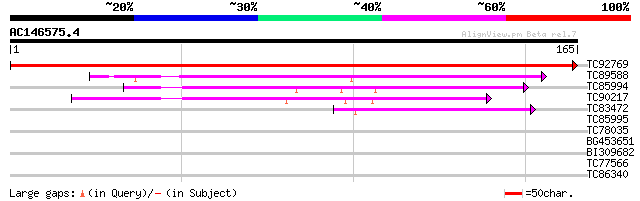
Score E
Sequences producing significant alignments: (bits) Value
TC92769 weakly similar to GP|5923663|gb|AAD56314.1| putative sig... 343 1e-95
TC89588 weakly similar to PIR|C86368|C86368 hypothetical protein... 74 3e-14
TC85994 similar to PIR|E84708|E84708 probable signal peptidase I... 56 7e-09
TC90217 similar to PIR|E84708|E84708 probable signal peptidase I... 44 3e-05
TC83472 similar to GP|9294054|dbj|BAB02011.1 contains similarity... 44 4e-05
TC85995 similar to PIR|E86203|E86203 probable signal peptidase [... 35 0.014
TC78035 similar to GP|8778540|gb|AAF79548.1| F22G5.9 {Arabidopsi... 28 1.7
BG453651 homologue to PIR|E84708|E847 probable signal peptidase ... 27 4.9
BI309682 weakly similar to GP|14532612|g unknown protein {Arabid... 26 6.4
TC77566 similar to GP|310580|gb|AAA34017.1|| protein kinase 2 {G... 26 6.4
TC86340 similar to GP|9294523|dbj|BAB02785.1 protein transport p... 26 6.4
>TC92769 weakly similar to GP|5923663|gb|AAD56314.1| putative signal
peptidase {Arabidopsis thaliana}, partial (93%)
Length = 634
Score = 343 bits (881), Expect = 1e-95
Identities = 165/165 (100%), Positives = 165/165 (100%)
Frame = +3
Query: 1 MGTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKL 60
MGTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKL
Sbjct: 87 MGTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKL 266
Query: 61 CLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNA 120
CLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNA
Sbjct: 267 CLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNA 446
Query: 121 ASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNTTPERLPSS 165
ASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNTTPERLPSS
Sbjct: 447 ASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNTTPERLPSS 581
>TC89588 weakly similar to PIR|C86368|C86368 hypothetical protein AAC98041.1
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1100
Score = 73.9 bits (180), Expect = 3e-14
Identities = 48/141 (34%), Positives = 72/141 (51%), Gaps = 8/141 (5%)
Frame = +1
Query: 24 VSDRYATVVPVR--GASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNF 81
V+++Y + PV+ G SM PT + T E++ K + GDIVI SP N
Sbjct: 316 VTNKYL-IDPVQTIGPSMLPTID-----VTPSLYLAERISPRFGKAAQGDIVILRSPRNP 477
Query: 82 KETHIKRIIALPGEWFV------NRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLG 135
+ KR++ L G+ Q+ + VP+GH W+EGDN S DS+++GPVP G
Sbjct: 478 RMCITKRLVGLEGDTITYVADPNKDDKQETVVVPKGHVWIEGDNKYKSNDSRNFGPVPYG 657
Query: 136 LVRGRVTHVVWPPQRIGAVKN 156
L+ R+ V P + G+ N
Sbjct: 658 LIESRLFWKVSPLKDFGSFWN 720
>TC85994 similar to PIR|E84708|E84708 probable signal peptidase I [imported]
- Arabidopsis thaliana, partial (64%)
Length = 973
Score = 55.8 bits (133), Expect = 7e-09
Identities = 45/146 (30%), Positives = 62/146 (41%), Gaps = 28/146 (19%)
Frame = +3
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFK-------ETHI 86
+ ASM PT D V EK K DIVIF +PS K + I
Sbjct: 75 IPSASMYPTLE------VGDRVLTEKFSFFFRKPDVSDIVIFKAPSWLKAYGFSSSDVFI 236
Query: 87 KRIIALPGE--------WFVNRHNQDV-------------LKVPEGHCWVEGDNAASSTD 125
KR++A G+ VN +D + VP+GH +V GDN S D
Sbjct: 237 KRVVAKAGDVVEVRDGKLLVNGVAEDEEFVLEPLAYELAPMVVPKGHVFVMGDNRNKSFD 416
Query: 126 SKSYGPVPLGLVRGRVTHVVWPPQRI 151
S ++GP+P+ + GR WPP ++
Sbjct: 417 SHNWGPLPIENIVGRSMFRYWPPSKV 494
>TC90217 similar to PIR|E84708|E84708 probable signal peptidase I [imported]
- Arabidopsis thaliana, partial (26%)
Length = 974
Score = 43.9 bits (102), Expect = 3e-05
Identities = 39/148 (26%), Positives = 64/148 (42%), Gaps = 26/148 (17%)
Frame = +3
Query: 19 LITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSP 78
L+ F + ++ + + +SM PT + D + +EK S DI+ F P
Sbjct: 282 LVVFLLWSTFSHLCYIPSSSMYPTLH------VGDRIIIEKASYYIRSPSIHDIITFRDP 443
Query: 79 S-----NFKETHIKRIIALPGEW--------FVNRHNQD-------------VLKVPEGH 112
+ N IKR++A G+ +VN Q+ + VP+GH
Sbjct: 444 TQHSGDNTDVIFIKRVVAKEGDTVEVHHGGLYVNGVAQEEDFVVEKPTYTTKLTYVPKGH 623
Query: 113 CWVEGDNAASSTDSKSYGPVPLGLVRGR 140
+V GDN +S DS +GP+P+ + GR
Sbjct: 624 VYVLGDNRNNSYDSHIWGPLPMKNIVGR 707
>TC83472 similar to GP|9294054|dbj|BAB02011.1 contains similarity to signal
peptidase (leader peptidase)~gene_id:MOB24.17
{Arabidopsis thaliana}, partial (23%)
Length = 468
Score = 43.5 bits (101), Expect = 4e-05
Identities = 23/62 (37%), Positives = 33/62 (53%), Gaps = 3/62 (4%)
Frame = +3
Query: 95 EWFVN---RHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRI 151
E F+N ++ +VPE +V GDN +S DS +GP+P + GR WPP +I
Sbjct: 6 EKFINEQPKYEMKPTRVPENSVFVMGDNRNNSYDSHVWGPLPAKNIIGRSVLRYWPPNKI 185
Query: 152 GA 153
A
Sbjct: 186 AA 191
>TC85995 similar to PIR|E86203|E86203 probable signal peptidase [imported] -
Arabidopsis thaliana, partial (61%)
Length = 1571
Score = 35.0 bits (79), Expect = 0.014
Identities = 36/124 (29%), Positives = 48/124 (38%), Gaps = 28/124 (22%)
Frame = +2
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFK-------ETHI 86
+ ASM PT D V EK K DI IF +PS K + I
Sbjct: 980 IPSASMYPTLE------VGDRVLTEKFSFFFTKPDVSDIAIFKAPSWLKAYGFSSSDVFI 1141
Query: 87 KRIIALPGE--------WFVNRHNQDV-------------LKVPEGHCWVEGDNAASSTD 125
KR++A G+ VN +D + VP+GH +V GDN S D
Sbjct: 1142 KRVVAKAGDVVEVRDGKLLVNGVAEDEEFVLEPLAYELAPMVVPKGHVFVMGDNRNKSFD 1321
Query: 126 SKSY 129
S ++
Sbjct: 1322 SHNW 1333
>TC78035 similar to GP|8778540|gb|AAF79548.1| F22G5.9 {Arabidopsis
thaliana}, partial (24%)
Length = 2694
Score = 28.1 bits (61), Expect = 1.7
Identities = 14/35 (40%), Positives = 20/35 (57%), Gaps = 2/35 (5%)
Frame = +3
Query: 99 NRHNQDVLKVPEGHCWVEGD--NAASSTDSKSYGP 131
N+H DVL++P C V D N++SS + K P
Sbjct: 426 NQHPLDVLEIPNSTCCVTTDSGNSSSSNELKPLSP 530
>BG453651 homologue to PIR|E84708|E847 probable signal peptidase I [imported]
- Arabidopsis thaliana, partial (18%)
Length = 621
Score = 26.6 bits (57), Expect = 4.9
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +2
Query: 108 VPEGHCWVEGDNAASSTDSKSY 129
VP+GH +V GDN S DS ++
Sbjct: 23 VPKGHVFVMGDNRNKSFDSHNW 88
>BI309682 weakly similar to GP|14532612|g unknown protein {Arabidopsis
thaliana}, partial (16%)
Length = 805
Score = 26.2 bits (56), Expect = 6.4
Identities = 17/48 (35%), Positives = 27/48 (55%), Gaps = 9/48 (18%)
Frame = -2
Query: 54 YVFVE-KLCLDKFKFSHGDIVIFSSPS--------NFKETHIKRIIAL 92
Y+F + K CL+ F+ SH + SSPS N K+T +K+++ L
Sbjct: 690 YIFQQSKHCLE-FRCSHDHFAVTSSPSFLSIFEG*NTKQTQLKQLLNL 550
>TC77566 similar to GP|310580|gb|AAA34017.1|| protein kinase 2 {Glycine
max}, partial (88%)
Length = 1502
Score = 26.2 bits (56), Expect = 6.4
Identities = 22/61 (36%), Positives = 30/61 (49%)
Frame = -3
Query: 48 NSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLK 107
N FT YVF +KF I +FS PSN K + K + L + VN N++ LK
Sbjct: 189 NLFTPLYVFES----NKFP----SIFVFSQPSNTKISRTKFLQWLISFFHVNPLNKNSLK 34
Query: 108 V 108
+
Sbjct: 33 L 31
>TC86340 similar to GP|9294523|dbj|BAB02785.1 protein transport protein
Sec23 {Arabidopsis thaliana}, partial (96%)
Length = 2988
Score = 26.2 bits (56), Expect = 6.4
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +1
Query: 119 NAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNTTP 159
N++SST S+ P G+ R+T VWP ++ + K P
Sbjct: 94 NSSSSTMSEMANTDPEGIDGVRMTWNVWPRSKVESSKCVIP 216
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,416,276
Number of Sequences: 36976
Number of extensions: 74200
Number of successful extensions: 333
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 331
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 333
length of query: 165
length of database: 9,014,727
effective HSP length: 89
effective length of query: 76
effective length of database: 5,723,863
effective search space: 435013588
effective search space used: 435013588
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146575.4


