

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146574.11 + phase: 0 /pseudo
(738 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
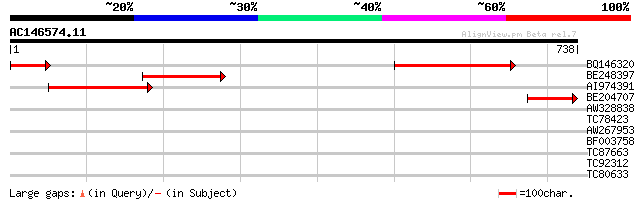
Score E
Sequences producing significant alignments: (bits) Value
BQ146320 303 2e-82
BE248397 221 8e-58
AI974391 weakly similar to PIR|F86194|F86 hypothetical protein [... 199 4e-51
BE204707 87 2e-17
AW328838 similar to GP|11559645|gb short integuments 1 {Arabidop... 39 0.009
TC78423 30 2.6
AW267953 similar to GP|21240638|gb hypothetical protein {Dictyos... 30 3.3
BF003758 similar to GP|15216030|em amino acid permease AAP4 {Vic... 30 4.4
TC87663 similar to GP|20258975|gb|AAM14203.1 unknown protein {Ar... 29 5.7
TC92312 weakly similar to GP|4567275|gb|AAD23688.1| expressed pr... 29 7.5
TC80633 weakly similar to GP|9954734|gb|AAG09087.1| Unknown Prot... 28 9.7
>BQ146320
Length = 473
Score = 303 bits (776), Expect = 2e-82
Identities = 151/157 (96%), Positives = 153/157 (97%)
Frame = +1
Query: 502 DKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEG 561
DKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEG
Sbjct: 1 DKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEG 180
Query: 562 GIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTILSQTALNALIR 621
GIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTILSQTALNALIR
Sbjct: 181 GIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTILSQTALNALIR 360
Query: 622 KRNALALQQRAIEDEMAVCNMKIHRWLAGEEDDFELK 658
K+NALAL QRAIEDEMAVCNMKIHRWLAG EDD +K
Sbjct: 361 KKNALALXQRAIEDEMAVCNMKIHRWLAGXEDDL*IK 471
Score = 68.2 bits (165), Expect = 1e-11
Identities = 30/52 (57%), Positives = 40/52 (76%)
Frame = +1
Query: 2 HIIEGGIKDESAFDKICVDATVENESIEKCTPIADNFNADFEKVRSFVDSKG 53
HI EGGIKD+SAFDKIC T EN+S+EKC A++ N + EK+++F+ SKG
Sbjct: 166 HITEGGIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKG 321
>BE248397
Length = 323
Score = 221 bits (563), Expect = 8e-58
Identities = 107/107 (100%), Positives = 107/107 (100%)
Frame = +1
Query: 174 CTCLDASKNVPNIEGWPVSKVAILLVDSKMENCLLRFCSTTDGVWSVIEKDVGSSDQISE 233
CTCLDASKNVPNIEGWPVSKVAILLVDSKMENCLLRFCSTTDGVWSVIEKDVGSSDQISE
Sbjct: 1 CTCLDASKNVPNIEGWPVSKVAILLVDSKMENCLLRFCSTTDGVWSVIEKDVGSSDQISE 180
Query: 234 DMNELKHTYQKRRVIQNPTKNGLNVDDDEFLQVGYSAVKEATGVNSN 280
DMNELKHTYQKRRVIQNPTKNGLNVDDDEFLQVGYSAVKEATGVNSN
Sbjct: 181 DMNELKHTYQKRRVIQNPTKNGLNVDDDEFLQVGYSAVKEATGVNSN 321
>AI974391 weakly similar to PIR|F86194|F86 hypothetical protein [imported] -
Arabidopsis thaliana, partial (3%)
Length = 421
Score = 199 bits (505), Expect = 4e-51
Identities = 92/135 (68%), Positives = 112/135 (82%)
Frame = +3
Query: 51 SKGKMGRSDVCPTADAVRAFVEHLVDPMLPAKASIRDAPSISQQQKVAKQVHSVVLLYNY 110
+K + SDVCPT DA++ ++HLVDP+LP KAS+RD P+ SQQQ +AKQVHSVVLLYNY
Sbjct: 15 NKNMVASSDVCPTEDAIQVGLQHLVDPLLPQKASVRDNPTTSQQQVIAKQVHSVVLLYNY 194
Query: 111 YHRKRHPELEFVAFKDFCKLIVGLRPALLIYMKFTQKPDQTDLVDVEQQLSLTEKAIASS 170
YHRK+HPELE++ F +FCKLIV LRP LL YMKF QK ++ +L+DVE+QLSLTEK I +
Sbjct: 195 YHRKQHPELEYLPFNEFCKLIVVLRPPLLAYMKFMQKINEVELIDVEKQLSLTEKGIMDA 374
Query: 171 CDICTCLDASKNVPN 185
CDIC CLDASKNVPN
Sbjct: 375 CDICNCLDASKNVPN 419
>BE204707
Length = 424
Score = 87.0 bits (214), Expect = 2e-17
Identities = 40/65 (61%), Positives = 48/65 (73%)
Frame = -3
Query: 674 LDGICHENNWILPTYSVSLSDGEFHATVRVKGVDFEYSCEGNTCPFPREARDSAAAQMLT 733
LDG+CHENNW+LPTY +S +G F A V V G F+ S EGN C P EAR+SAAA+ML
Sbjct: 422 LDGVCHENNWVLPTYRISQPNGGFQANVTVNGDKFKCSDEGNLCSTPLEARESAAAKMLN 243
Query: 734 KFRSM 738
KF+SM
Sbjct: 242 KFKSM 228
>AW328838 similar to GP|11559645|gb short integuments 1 {Arabidopsis
thaliana}, partial (4%)
Length = 532
Score = 38.5 bits (88), Expect = 0.009
Identities = 32/101 (31%), Positives = 46/101 (44%), Gaps = 13/101 (12%)
Frame = +2
Query: 645 HRWLAGEEDDFELKL-----ESVIEGCNGTWLRK-LDGICHENNWILPTYSVSLSDGEFH 698
H LA ++ E K+ E + N T+ R+ L+ IC NW +P Y G H
Sbjct: 2 HEALAALKEKEESKIQEKNDEKETKNGNQTFTRQTLNDICLRRNWPMPFYRCVSEGGPAH 181
Query: 699 A-----TVRVKGVDFEYS--CEGNTCPFPREARDSAAAQML 732
A VRV D ++ C G P ++A+DSAA +L
Sbjct: 182 AKRFTFAVRVNTTDKGWTDECVGEAMPSVKKAKDSAAVLLL 304
>TC78423
Length = 624
Score = 30.4 bits (67), Expect = 2.6
Identities = 10/33 (30%), Positives = 19/33 (57%)
Frame = -3
Query: 509 SKNKECQKHTANTLHISEDQKIENPSVQHHSNE 541
++ +CQ N H+ +D + NP +Q+H N+
Sbjct: 172 NQTHQCQVTNLNPNHVQDDHFLRNPCLQNHQND 74
>AW267953 similar to GP|21240638|gb hypothetical protein {Dictyostelium
discoideum}, partial (1%)
Length = 569
Score = 30.0 bits (66), Expect = 3.3
Identities = 20/76 (26%), Positives = 34/76 (44%)
Frame = +3
Query: 524 ISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEGGIKDQSAFDKICAGTTFENDSV 583
+S + KIE P + SNE P K EK+ + A +K+ +SV
Sbjct: 57 VSAEVKIETPEAETKSNELNEPIKLEKSSET--------VVVPPAEEKVIEVVLTNAESV 212
Query: 584 EKCILNANSSNKNLEK 599
+ ++ A SS +++ K
Sbjct: 213 DNKVVEATSSIEDISK 260
>BF003758 similar to GP|15216030|em amino acid permease AAP4 {Vicia faba var.
minor}, partial (31%)
Length = 554
Score = 29.6 bits (65), Expect = 4.4
Identities = 12/28 (42%), Positives = 18/28 (63%)
Frame = -2
Query: 530 IENPSVQHHSNECTRPSKAEKAVSKRKH 557
I NPS ++H+N + PSK + S R+H
Sbjct: 340 ISNPSKENHNNRTSNPSKLSNSPS*RQH 257
>TC87663 similar to GP|20258975|gb|AAM14203.1 unknown protein {Arabidopsis
thaliana}, partial (40%)
Length = 1166
Score = 29.3 bits (64), Expect = 5.7
Identities = 14/45 (31%), Positives = 24/45 (53%)
Frame = +3
Query: 494 KEDWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHH 538
K+D ++DVD VL + EC K ++N H + + +P + H
Sbjct: 915 KQDMNVDVDPPSVLVTDVSECYKRSSNKQHHANLLQRASPEIAIH 1049
>TC92312 weakly similar to GP|4567275|gb|AAD23688.1| expressed protein
{Arabidopsis thaliana}, partial (29%)
Length = 1319
Score = 28.9 bits (63), Expect = 7.5
Identities = 13/24 (54%), Positives = 17/24 (70%)
Frame = -3
Query: 323 LIESFRGPLLKKSSSSWTITSVVE 346
+I FRG LKK+S SWT+ +VE
Sbjct: 1092 IISCFRGFKLKKTSPSWTLGHLVE 1021
>TC80633 weakly similar to GP|9954734|gb|AAG09087.1| Unknown Protein
{Arabidopsis thaliana}, partial (8%)
Length = 833
Score = 28.5 bits (62), Expect = 9.7
Identities = 15/39 (38%), Positives = 22/39 (55%)
Frame = +2
Query: 72 EHLVDPMLPAKASIRDAPSISQQQKVAKQVHSVVLLYNY 110
E+L + + SIR+ PSI +Q KV KQ ++ NY
Sbjct: 476 ENLDEVLAAESLSIREVPSIDKQLKVNKQFLDIIP*ENY 592
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,303,865
Number of Sequences: 36976
Number of extensions: 337984
Number of successful extensions: 2049
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1511
Number of HSP's successfully gapped in prelim test: 81
Number of HSP's that attempted gapping in prelim test: 517
Number of HSP's gapped (non-prelim): 1624
length of query: 738
length of database: 9,014,727
effective HSP length: 103
effective length of query: 635
effective length of database: 5,206,199
effective search space: 3305936365
effective search space used: 3305936365
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146574.11


