

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146570.9 + phase: 0
(475 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
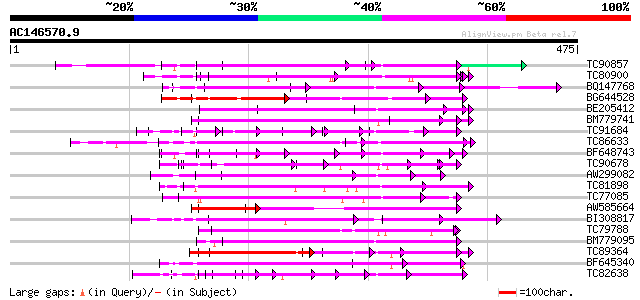
Score E
Sequences producing significant alignments: (bits) Value
TC90857 weakly similar to GP|16930691|gb|AAL32011.1 AT4g26540/M3... 136 2e-32
TC80900 weakly similar to PIR|G84652|G84652 probable receptor-li... 118 6e-27
BQ147768 weakly similar to PIR|H84632|H84 probable receptor-like... 115 5e-26
BG644528 weakly similar to GP|21391894|g systemin receptor SR160... 114 8e-26
BE205412 weakly similar to GP|14495542|gb receptor-like protein ... 113 2e-25
BM779741 weakly similar to GP|13872902|db putative protein kinas... 109 2e-24
TC91684 weakly similar to PIR|A96574|A96574 protein F12M16.30 [i... 78 3e-24
TC86633 weakly similar to SP|Q00874|D100_ARATH DNA-damage-repair... 108 4e-24
BF648743 weakly similar to PIR|H84632|H846 probable receptor-lik... 107 8e-24
TC90678 weakly similar to GP|9759550|dbj|BAB11152.1 receptor pro... 107 1e-23
AW299082 weakly similar to GP|10177183|d receptor protein kinase... 105 3e-23
TC81898 weakly similar to GP|20260576|gb|AAM13186.1 putative rec... 102 3e-22
TC77085 similar to GP|21536600|gb|AAM60932.1 putative disease re... 101 5e-22
AW585664 similar to PIR|T46033|T46 receptor protein kinase-like ... 100 9e-22
BI308817 weakly similar to PIR|B84659|B846 probable receptor-lik... 100 1e-21
TC79788 weakly similar to GP|21554189|gb|AAM63268.1 putative leu... 99 3e-21
BM779095 weakly similar to PIR|B84431|B8 probable receptor prote... 97 1e-20
TC89364 weakly similar to PIR|C96745|C96745 hypothetical protein... 97 1e-20
BF645340 weakly similar to GP|6016693|gb|A putative disease resi... 97 1e-20
TC82638 weakly similar to PIR|T00712|T00712 protein kinase homol... 95 5e-20
>TC90857 weakly similar to GP|16930691|gb|AAL32011.1 AT4g26540/M3E9_30
{Arabidopsis thaliana}, partial (22%)
Length = 829
Score = 136 bits (342), Expect = 2e-32
Identities = 85/222 (38%), Positives = 127/222 (56%)
Frame = +3
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G+IP+ G LKNL++L+L +N L G IP EIGN +L + S N+ +G+IP FG L
Sbjct: 9 LTGSIPTKLGNLKNLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIPKTFGNL 188
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR 276
+ L L LS N +SG +P LG + +++ +N + G + +E GNL NLTL+ L +N+
Sbjct: 189 TLLQELQLSVNQISGEIPAELGNCQQLTHVEIDNNLITGTIPSELGNLGNLTLLFLWHNK 368
Query: 277 LCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQ 336
L + +L +LE + LS N L G I ++ LQNL L L + L G+IP +
Sbjct: 369 LQGNIPSTLSNCQNLEAIDLSQNLLTGPIPKGIFQ-LQNLNKLLLLSNNLSGKIPSQIGN 545
Query: 337 LKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L ++NNITG + ++ L +LN L L N ++G I
Sbjct: 546 CSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRIEGII 671
Score = 120 bits (300), Expect = 1e-27
Identities = 72/200 (36%), Positives = 111/200 (55%)
Frame = +3
Query: 179 NGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLG 238
N LTG+IP ++GNL LK L+L NN G IP G L ++D S NS++G++P T G
Sbjct: 3 NSLTGSIPTKLGNLKNLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIPKTFG 182
Query: 239 RLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSN 298
L + +L LS N + G++ E GN + LT +++ NN + + L + +L + L +
Sbjct: 183 NLTLLQELQLSVNQISGEIPAELGNCQQLTHVEIDNNLITGTIPSELGNLGNLTLLFLWH 362
Query: 299 NPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKL 358
N L G+I + N QNL ++LS L G IP+ + QL+ L L L NN++G + ++
Sbjct: 363 NKLQGNIPS-TLSNCQNLEAIDLSQNLLTGPIPKGIFQLQNLNKLLLLSNNLSGKIPSQI 539
Query: 359 ETLPSLNALYLSGNNLKGEI 378
SL + NN+ G I
Sbjct: 540 GNCSSLIRFRANNNNITGFI 599
Score = 100 bits (248), Expect = 2e-21
Identities = 82/271 (30%), Positives = 127/271 (46%), Gaps = 2/271 (0%)
Frame = +3
Query: 39 QEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINIGPIHENSLPC 98
Q L TI +GN C S+ S + G T N+ + E L
Sbjct: 72 QNNLVGTIPSEIGN-----------CYQLSVIDASMNSITGSIPKTFGNLTLLQELQLSV 218
Query: 99 ANEKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPG 156
E EL + L + N N + +IP+ GN L ++ F +
Sbjct: 219 NQISGEIPAELGNCQQLTHVEIDN-----NLITGTIPSELGNLGNL-----TLLFLWHNK 368
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L GNIPST +NL+++ L +N LTG IP+ I L L +L+L NN SG IP G
Sbjct: 369 LQGNIPSTLSNCQNLEAIDLSQNLLTGPIPKGIFQLQNLNKLLLLSNNLSGKIPSQIGNC 548
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR 276
S L+ + N+++G +P +G L ++ LDL N +EG + + +NLT +DL +N
Sbjct: 549 SSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRIEGIIPEKISGCRNLTFLDLHSNY 728
Query: 277 LCCGLVLSLQEMNSLEEMVLSNNPLGGDIRT 307
+ L SL E+ SL+ + S+N + G +++
Sbjct: 729 IAGTLPDSLSELVSLQFLDFSDNMIEGALKS 821
Score = 85.1 bits (209), Expect = 5e-17
Identities = 56/158 (35%), Positives = 81/158 (50%)
Frame = +3
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
NKL +IP+ ++LE+I+ N L G IP L+NL L+LL N L+G IP
Sbjct: 363 NKLQGNIPSTLSN--CQNLEAIDLSQNL-LTGPIPKGIFQLQNLNKLLLLSNNLSGKIPS 533
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLD 247
+IGN L R + NN +G IP G L +L LDL N + G +P + ++ LD
Sbjct: 534 QIGNCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRIEGIIPEKISGCRNLTFLD 713
Query: 248 LSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSL 285
L N++ G L + L +L +D +N + L SL
Sbjct: 714 LHSNYIAGTLPDSLSELVSLQFLDFSDNMIEGALKSSL 827
Score = 46.2 bits (108), Expect = 3e-05
Identities = 42/149 (28%), Positives = 57/149 (38%), Gaps = 14/149 (9%)
Frame = +3
Query: 299 NPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKL 358
N L G I T K NL+NL L L L+G IP + +L + S N+ITG++
Sbjct: 3 NSLTGSIPT-KLGNLKNLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIPKTF 179
Query: 359 ETLPSLNALYLSGNNLKGEIQFSKG--------------FFGKLGRRFGAWSNPKLCYPF 404
L L L LS N + GEI G G + G N L + +
Sbjct: 180 GNLTLLQELQLSVNQISGEIPAELGNCQQLTHVEIDNNLITGTIPSELGNLGNLTLLFLW 359
Query: 405 ELMSTNNVPYGVKPCHQEEIHLVKSNAKT 433
N+P + C E + N T
Sbjct: 360 HNKLQGNIPSTLSNCQNLEAIDLSQNLLT 446
>TC80900 weakly similar to PIR|G84652|G84652 probable receptor-like protein
kinase [imported] - Arabidopsis thaliana, partial (24%)
Length = 1054
Score = 118 bits (295), Expect = 6e-27
Identities = 76/213 (35%), Positives = 121/213 (56%), Gaps = 1/213 (0%)
Frame = +2
Query: 167 VLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
++K L+ + L N L+ IP+ IGNLV L L L NN +G IP+ G L++L L L
Sbjct: 8 LMKRLKWIYLGYNNLSREIPKNIGNLVSLNHLNLVYNNLTGPIPESLGNLTNLQYLFLYL 187
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQ 286
N L+G +P ++ L +++ LDLS N+L G++ N NL+ L ++ L +N + ++
Sbjct: 188 NKLTGPIPKSIFNLKNLISLDLSDNYLSGEISNLVVNLQKLEILHLFSNNFTGKIPNTIT 367
Query: 287 EMNSLEEMVLSNNPLGGDI-RTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGL 345
+ L+ + L +N L G+I +TL N NL IL+LS+ L G+IP SL K L + L
Sbjct: 368 SLPHLQVLQLWSNKLTGEIPQTLGIHN--NLTILDLSSNNLTGKIPNSLCASKNLHKIIL 541
Query: 346 SDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
N++ G + L + +L + L NNL G++
Sbjct: 542 FSNSLKGEIPKGLTSCKTLERVRLQDNNLSGKL 640
Score = 117 bits (294), Expect = 7e-27
Identities = 80/227 (35%), Positives = 120/227 (52%)
Frame = +2
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G IP + G L NLQ L L N LTG IP+ I NL L L LS N SG I ++ L
Sbjct: 122 LTGPIPESLGNLTNLQYLFLYLNKLTGPIPKSIFNLKNLISLDLSDNYLSGEISNLVVNL 301
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR 276
L IL L N+ +G +P T+ L + L L N L G++ G NLT++DL +N
Sbjct: 302 QKLEILHLFSNNFTGKIPNTITSLPHLQVLQLWSNKLTGEIPQTLGIHNNLTILDLSSNN 481
Query: 277 LCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQ 336
L + SL +L +++L +N L G+I + + L + L + L G++P ++Q
Sbjct: 482 LTGKIPNSLCASKNLHKIILFSNSLKGEI-PKGLTSCKTLERVRLQDNNLSGKLPLEITQ 658
Query: 337 LKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKG 383
L ++ L +S N +G ++ + +PSL L L+ NN G++ S G
Sbjct: 659 LPQIYLLDISGNKFSGKINDRKWNMPSLQMLNLANNNFSGDLPNSFG 799
Score = 112 bits (281), Expect = 2e-25
Identities = 80/222 (36%), Positives = 117/222 (52%)
Frame = +2
Query: 161 IPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLL 220
IP G L +L L L+ N LTG IP+ +GNL L+ L L N +G IP L +L+
Sbjct: 62 IPKNIGNLVSLNHLNLVYNNLTGPIPESLGNLTNLQYLFLYLNKLTGPIPKSIFNLKNLI 241
Query: 221 ILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCG 280
LDLS N LSG + + L + L L N GK+ N +L +L ++ L +N+L
Sbjct: 242 SLDLSDNYLSGEISNLVVNLQKLEILHLFSNNFTGKIPNTITSLPHLQVLQLWSNKLTGE 421
Query: 281 LVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKL 340
+ +L N+L + LS+N L G I + +NL + L + L GEIP+ L+ K L
Sbjct: 422 IPQTLGIHNNLTILDLSSNNLTGKIPNSLCAS-KNLHKIILFSNSLKGEIPKGLTSCKTL 598
Query: 341 RFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSK 382
+ L DNN++G L ++ LP + L +SGN G+I K
Sbjct: 599 ERVRLQDNNLSGKLPLEITQLPQIYLLDISGNKFSGKINDRK 724
Score = 103 bits (256), Expect = 2e-22
Identities = 91/269 (33%), Positives = 125/269 (45%), Gaps = 47/269 (17%)
Frame = +2
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G IP + LKNL SL L +N L+G I + NL KL+ L L NNF+G IP+ L
Sbjct: 194 LTGPIPKSIFNLKNLISLDLSDNYLSGEISNLVVNLQKLEILHLFSNNFTGKIPNTITSL 373
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNL--------- 267
L +L L N L+G +P TLG ++ LDLS N L GK+ N KNL
Sbjct: 374 PHLQVLQLWSNKLTGEIPQTLGIHNNLTILDLSSNNLTGKIPNSLCASKNLHKIILFSNS 553
Query: 268 -------------TL--MDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWEN 312
TL + L++N L L L + ++ + + +S N G I KW N
Sbjct: 554 LKGEIPKGLTSCKTLERVRLQDNNLSGKLPLEITQLPQIYLLDISGNKFSGKINDRKW-N 730
Query: 313 LQNLVILELSNMELIGEIPES----------LSQ-------------LKKLRFLGLSDNN 349
+ +L +L L+N G++P S LSQ L +L L L++NN
Sbjct: 731 MPSLQMLNLANNNFSGDLPNSFGGNKVEGLDLSQNQFSGYIQIGFKNLPELVQLKLNNNN 910
Query: 350 ITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+ G +L L +L LS N L GEI
Sbjct: 911 LFGKFPEELFQCNKLVSLDLSHNRLNGEI 997
Score = 74.3 bits (181), Expect = 9e-14
Identities = 54/165 (32%), Positives = 76/165 (45%)
Frame = +2
Query: 224 LSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVL 283
L N+LS +P +G L+S+ L+L +N L G + GNL NL + L N+L +
Sbjct: 35 LGYNNLSREIPKNIGNLVSLNHLNLVYNNLTGPIPESLGNLTNLQYLFLYLNKLTGPIPK 214
Query: 284 SLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFL 343
S+ NL+NL+ L+LS+ L GEI + L+KL L
Sbjct: 215 SIF-------------------------NLKNLISLDLSDNYLSGEISNLVVNLQKLEIL 319
Query: 344 GLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
L NN TG + + +LP L L L N L GEI + G L
Sbjct: 320 HLFSNNFTGKIPNTITSLPHLQVLQLWSNKLTGEIPQTLGIHNNL 454
Score = 67.0 bits (162), Expect = 1e-11
Identities = 54/187 (28%), Positives = 84/187 (44%), Gaps = 23/187 (12%)
Frame = +2
Query: 113 KHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQ 172
K+L I F+ N L IP G ++LE + + N L G +P L +
Sbjct: 518 KNLHKIILFS-----NSLKGEIPKGLTS--CKTLERVRLQDN-NLSGKLPLEITQLPQIY 673
Query: 173 SLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGG----------------- 215
L + N +G I N+ L+ L L+ NNFSG++P+ FGG
Sbjct: 674 LLDISGNKFSGKINDRKWNMPSLQMLNLANNNFSGDLPNSFGGNKVEGLDLSQNQFSGYI 853
Query: 216 ------LSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTL 269
L +L+ L L+ N+L G P L + ++ LDLSHN L G++ + + L L
Sbjct: 854 QIGFKNLPELVQLKLNNNNLFGKFPEELFQCNKLVSLDLSHNRLNGEIPEKLAKMPVLGL 1033
Query: 270 MDLRNNR 276
+D+ N+
Sbjct: 1034LDISENQ 1054
Score = 32.3 bits (72), Expect = 0.41
Identities = 18/47 (38%), Positives = 27/47 (57%)
Frame = +2
Query: 337 LKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKG 383
+K+L+++ L NN++ + + L SLN L L NNL G I S G
Sbjct: 11 MKRLKWIYLGYNNLSREIPKNIGNLVSLNHLNLVYNNLTGPIPESLG 151
>BQ147768 weakly similar to PIR|H84632|H84 probable receptor-like protein
kinase [imported] - Arabidopsis thaliana, partial (12%)
Length = 712
Score = 115 bits (287), Expect = 5e-26
Identities = 78/210 (37%), Positives = 116/210 (55%)
Frame = +3
Query: 137 GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLK 196
GN KL ++F N L G IP++ G + L+ L + N L G+IP E+G + L
Sbjct: 9 GNLSKLTH----LDFSYN-SLEGEIPNSLGNHRQLKYLDISNNNLNGSIPHELGFIKYLG 173
Query: 197 RLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGK 256
L LS N SG+IP G L L L + NSL G +P ++G L S+ L++S N+++G
Sbjct: 174 SLNLSTNRISGDIPPSLGNLVKLTHLVIYGNSLVGKIPPSIGNLRSLESLEISDNYIQGS 353
Query: 257 LLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNL 316
+ G LKNLT + L +NR+ + SL + LEE+ +SNN + G + + L+NL
Sbjct: 354 IPPRLGLLKNLTTLRLSHNRIKGEIPPSLGNLKQLEELDISNNNIQGFL-PFELGLLKNL 530
Query: 317 VILELSNMELIGEIPESLSQLKKLRFLGLS 346
L+LS+ L G +P SL L +L +L S
Sbjct: 531 TTLDLSHNRLNGNLPISLKNLTQLIYLNCS 620
Score = 109 bits (273), Expect = 2e-24
Identities = 79/218 (36%), Positives = 109/218 (49%)
Frame = +3
Query: 164 TFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILD 223
+ G L L L N L G IP +GN +LK L +S NN +G+IP G + L L+
Sbjct: 3 SLGNLSKLTHLDFSYNSLEGEIPNSLGNHRQLKYLDISNNNLNGSIPHELGFIKYLGSLN 182
Query: 224 LSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVL 283
LS N +SG +P +LG L+ + L + N L GK+ GNL+
Sbjct: 183 LSTNRISGDIPPSLGNLVKLTHLVIYGNSLVGKIPPSIGNLR------------------ 308
Query: 284 SLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFL 343
SLE + +S+N + G I + L+NL L LS+ + GEIP SL LK+L L
Sbjct: 309 ------SLESLEISDNYIQGSIPP-RLGLLKNLTTLRLSHNRIKGEIPPSLGNLKQLEEL 467
Query: 344 GLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFS 381
+S+NNI G L +L L +L L LS N L G + S
Sbjct: 468 DISNNNIQGFLPFELGLLKNLTTLDLSHNRLNGNLPIS 581
Score = 96.3 bits (238), Expect = 2e-20
Identities = 71/227 (31%), Positives = 108/227 (47%)
Frame = +3
Query: 236 TLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMV 295
+LG L + LD S+N LEG++ N GN + L +D+ NN L + L + L +
Sbjct: 3 SLGNLSKLTHLDFSYNSLEGEIPNSLGNHRQLKYLDISNNNLNGSIPHELGFIKYLGSLN 182
Query: 296 LSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLS 355
LS N + GDI NL L L + L+G+IP S+ L+ L L +SDN I G++
Sbjct: 183 LSTNRISGDIPP-SLGNLVKLTHLVIYGNSLVGKIPPSIGNLRSLESLEISDNYIQGSIP 359
Query: 356 PKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPFELMSTNNVPYG 415
P+L L +L L LS N +KGEI S G +L + +N + PFEL N
Sbjct: 360 PRLGLLKNLTTLRLSHNRIKGEIPPSLGNLKQLEELDISNNNIQGFLPFELGLLKN---- 527
Query: 416 VKPCHQEEIHLVKSNAKTEVINGDINHNSNFITSMGFSSCATSCFWW 462
L + +NG++ + +T + + +C+ F W
Sbjct: 528 ----------LTTLDLSHNRLNGNLPISLKNLTQLIYLNCSYKFFHW 638
Score = 85.1 bits (209), Expect = 5e-17
Identities = 53/124 (42%), Positives = 73/124 (58%)
Frame = +3
Query: 129 KLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQE 188
K+P SI GN SLES+E N + G+IP G+LKNL +L L N + G IP
Sbjct: 279 KIPPSI--GN----LRSLESLEISDNY-IQGSIPPRLGLLKNLTTLRLSHNRIKGEIPPS 437
Query: 189 IGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDL 248
+GNL +L+ L +S NN G +P G L +L LDLS N L+G LP++L L ++ L+
Sbjct: 438 LGNLKQLEELDISNNNIQGFLPFELGLLKNLTTLDLSHNRLNGNLPISLKNLTQLIYLNC 617
Query: 249 SHNF 252
S+ F
Sbjct: 618 SYKF 629
>BG644528 weakly similar to GP|21391894|g systemin receptor SR160
{Lycopersicon peruvianum}, partial (10%)
Length = 762
Score = 114 bits (285), Expect = 8e-26
Identities = 75/200 (37%), Positives = 106/200 (52%)
Frame = +2
Query: 153 SNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDI 212
+N L G+IP +NL S +N TG +P E+G L KL+RL++ N SG IPDI
Sbjct: 158 ANNQLNGSIPHGMKKFQNLISFSFEQNYFTGELPLELGTLKKLERLLIYQNRLSGEIPDI 337
Query: 213 FGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDL 272
FG ++L IL + N SG + ++GR + LDL N L G + E L LT + L
Sbjct: 338 FGNFTNLFILAIGNNQFSGRIHASIGRCKRLSFLDLRMNKLAGVIPMEIFQLSGLTTLYL 517
Query: 273 RNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPE 332
N L G + +M LE MV+S+N L G+I ++ L+ L+ ++ G IP
Sbjct: 518 HGNSL-NGSLPPQFKMEQLEAMVVSDNKLSGNIPKIEVNGLKTLM---MARNNFSGSIPN 685
Query: 333 SLSQLKKLRFLGLSDNNITG 352
SL L L L LS N++TG
Sbjct: 686 SLGDLPSLVTLDLSSNSLTG 745
Score = 62.8 bits (151), Expect = 3e-10
Identities = 42/107 (39%), Positives = 65/107 (60%)
Frame = +2
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
NKL IP ++ L ++ N L G++P F ++ L+++V+ +N L+GNIP+
Sbjct: 452 NKLAGVIPMEIFQ--LSGLTTLYLHGN-SLNGSLPPQFK-MEQLEAMVVSDNKLSGNIPK 619
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLP 234
N LK L+++ NNFSG+IP+ G L L+ LDLS NSL+G +P
Sbjct: 620 IEVN--GLKTLMMARNNFSGSIPNSLGDLPSLVTLDLSSNSLTGPIP 754
Score = 47.8 bits (112), Expect = 9e-06
Identities = 36/136 (26%), Positives = 58/136 (42%), Gaps = 1/136 (0%)
Frame = +2
Query: 249 SHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNS-LEEMVLSNNPLGGDIRT 307
S+ L + N L ++ + +N L L S+ ++S L++ ++NN L G I
Sbjct: 11 SNTSLNFQFFESLRNSTQLQILMINDNNLTGELPSSVDYLSSNLQQFCVANNQLNGSIPH 190
Query: 308 LKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNAL 367
+ QNL+ GE+P L LKKL L + N ++G + +L L
Sbjct: 191 -GMKKFQNLISFSFEQNYFTGELPLELGTLKKLERLLIYQNRLSGEIPDIFGNFTNLFIL 367
Query: 368 YLSGNNLKGEIQFSKG 383
+ N G I S G
Sbjct: 368 AIGNNQFSGRIHASIG 415
>BE205412 weakly similar to GP|14495542|gb receptor-like protein kinase
INRPK1 {Ipomoea nil}, partial (7%)
Length = 627
Score = 113 bits (282), Expect = 2e-25
Identities = 70/209 (33%), Positives = 106/209 (50%), Gaps = 1/209 (0%)
Frame = +2
Query: 160 NIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDL 219
NI FG+ NL + L N L G + + G + L + +SGN SG IP LS L
Sbjct: 2 NITEAFGIHPNLSFISLSRNRLIGYLSPDWGKCISLTEMEMSGNKLSGKIPIDLNKLSKL 181
Query: 220 LILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCC 279
L L N +G +P +G + + L+LS N L G++ G L L ++DL +N
Sbjct: 182 QFLSLHSNEFTGNIPHEIGNISLLFMLNLSRNHLSGEIPKSIGRLAQLNIVDLSDNNFSG 361
Query: 280 GLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNL-VILELSNMELIGEIPESLSQLK 338
+ L N L M LS+N L G I + NL +L +L+LS+ L EIP++L +L
Sbjct: 362 SIPNELGNCNRLLSMNLSHNDLSGMI-PYELGNLYSLQSLLDLSSNNLSREIPQNLQKLA 538
Query: 339 KLRFLGLSDNNITGNLSPKLETLPSLNAL 367
L +S NN++G + ++PSL ++
Sbjct: 539 SLEIFNVSHNNLSGTIPQSFSSMPSLQSV 625
Score = 75.1 bits (183), Expect = 5e-14
Identities = 54/176 (30%), Positives = 88/176 (49%)
Frame = +2
Query: 208 NIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNL 267
NI + FG +L + LSRN L G L G+ IS+ ++++S N L GK+
Sbjct: 2 NITEAFGIHPNLSFISLSRNRLIGYLSPDWGKCISLTEMEMSGNKLSGKI---------- 151
Query: 268 TLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELI 327
+ L +++ L+ + L +N G+I + N+ L +L LS L
Sbjct: 152 --------------PIDLNKLSKLQFLSLHSNEFTGNIPH-EIGNISLLFMLNLSRNHLS 286
Query: 328 GEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKG 383
GEIP+S+ +L +L + LSDNN +G++ +L L ++ LS N+L G I + G
Sbjct: 287 GEIPKSIGRLAQLNIVDLSDNNFSGSIPNELGNCNRLLSMNLSHNDLSGMIPYELG 454
Score = 55.8 bits (133), Expect = 3e-08
Identities = 37/99 (37%), Positives = 54/99 (54%)
Frame = +2
Query: 290 SLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNN 349
+L + LS N L G + W +L +E+S +L G+IP L++L KL+FL L N
Sbjct: 32 NLSFISLSRNRLIGYLSP-DWGKCISLTEMEMSGNKLSGKIPIDLNKLSKLQFLSLHSNE 208
Query: 350 ITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
TGN+ ++ + L L LS N+L GEI S G +L
Sbjct: 209 FTGNIPHEIGNISLLFMLNLSRNHLSGEIPKSIGRLAQL 325
Score = 34.7 bits (78), Expect = 0.082
Identities = 21/59 (35%), Positives = 27/59 (45%)
Frame = +2
Query: 330 IPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
I E+ L F+ LS N + G LSP SL + +SGN L G+I KL
Sbjct: 5 ITEAFGIHPNLSFISLSRNRLIGYLSPDWGKCISLTEMEMSGNKLSGKIPIDLNKLSKL 181
>BM779741 weakly similar to GP|13872902|db putative protein kinase Xa21
{Oryza sativa (japonica cultivar-group)}, partial (11%)
Length = 815
Score = 109 bits (273), Expect = 2e-24
Identities = 75/215 (34%), Positives = 108/215 (49%), Gaps = 3/215 (1%)
Frame = +3
Query: 153 SNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDI 212
SN L G IP+ G+LK L+ L L N L G IP E+ N +K + L+ N G +P
Sbjct: 165 SNVNLHGEIPTQVGLLKGLRVLDLGNNNLQGEIPIELTNCTNIKVIRLALNKLIGRVPAY 344
Query: 213 FGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDL 272
FG + L L L N+L GT+P +LG L S+ KL N LEG + G L LT + L
Sbjct: 345 FGSMMQLTELSLGHNNLVGTIPSSLGNLSSLEKLSFLQNHLEGSIPYSLGRLSVLTWLSL 524
Query: 273 RNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRT---LKWENLQNLVILELSNMELIGE 329
N L + SL +++++ +S N L G I + L + NL+ I + ++
Sbjct: 525 SLNNLSGEIPHSLYNLSNIQIFSISGNNLFGSIPSNIDLVFPNLEQFFI---GSNQISAT 695
Query: 330 IPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSL 364
P S+S L +L+ ++ NNI G L L L L
Sbjct: 696 FPSSISNLTRLQVFDIAYNNINGALPLTLGRLNKL 800
Score = 99.8 bits (247), Expect = 2e-21
Identities = 68/231 (29%), Positives = 108/231 (46%), Gaps = 1/231 (0%)
Frame = +3
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G + S+ G L L+ L L L G IP ++G L L+ L L NN G IP ++
Sbjct: 111 GTLGSSLGNLTFLRMLNLSNVNLHGEIPTQVGLLKGLRVLDLGNNNLQGEIPIELTNCTN 290
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
+ ++ L+ N L G +P G ++ + +L L HN L G + + GNL +L + N L
Sbjct: 291 IKVIRLALNKLIGRVPAYFGSMMQLTELSLGHNNLVGTIPSSLGNLSSLEKLSFLQNHLE 470
Query: 279 CGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQL- 337
+ SL ++ L + LS N L G+I + NL N+ I +S L G IP ++ +
Sbjct: 471 GSIPYSLGRLSVLTWLSLSLNNLSGEIPHSLY-NLSNIQIFSISGNNLFGSIPSNIDLVF 647
Query: 338 KKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
L + N I+ + L L ++ NN+ G + + G KL
Sbjct: 648 PNLEQFFIGSNQISATFPSSISNLTRLQVFDIAYNNINGALPLTLGRLNKL 800
Score = 41.6 bits (96), Expect = 7e-04
Identities = 24/60 (40%), Positives = 31/60 (51%)
Frame = +3
Query: 319 LELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L L N G + SL L LR L LS+ N+ G + ++ L L L L NNL+GEI
Sbjct: 84 LHLENQTFGGTLGSSLGNLTFLRMLNLSNVNLHGEIPTQVGLLKGLRVLDLGNNNLQGEI 263
>TC91684 weakly similar to PIR|A96574|A96574 protein F12M16.30 [imported] -
Arabidopsis thaliana, partial (20%)
Length = 1085
Score = 78.2 bits (191), Expect(2) = 3e-24
Identities = 57/162 (35%), Positives = 87/162 (53%)
Frame = +1
Query: 107 PELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFG 166
P+L +L L+ I + N L +IP W L L ++ F N L G IP FG
Sbjct: 19 PDLVRLPFLQEIDL-----TLNYLNGTIPK-QWATL--KLVNVSFYGNR-LSGPIPKEFG 171
Query: 167 VLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
+ L+SLVL N L+GN+P E+G+L +++RL+LS NNF+G +P F L+ L +
Sbjct: 172 NITTLKSLVLEFNQLSGNLPPELGSLSQIERLLLSSNNFTGLLPATFAKLTALKQFRIGD 351
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLT 268
+ SG +P + I++ L + + L G + + LKNLT
Sbjct: 352 SQFSGAIPNFIQSWINLEMLTIRGSGLSGPIPSGISLLKNLT 477
Score = 72.8 bits (177), Expect = 3e-13
Identities = 44/141 (31%), Positives = 74/141 (52%)
Frame = +1
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G IP + LK L ++ N L+G IP+E GN+ LK LVL N SGN+P G L
Sbjct: 73 LNGTIPKQWATLK-LVNVSFYGNRLSGPIPKEFGNITTLKSLVLEFNQLSGNLPPELGSL 249
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR 276
S + L LS N+ +G LP T +L ++ + + + G + N + NL ++ +R +
Sbjct: 250 SQIERLLLSSNNFTGLLPATFAKLTALKQFRIGDSQFSGAIPNFIQSWINLEMLTIRGSG 429
Query: 277 LCCGLVLSLQEMNSLEEMVLS 297
L + + + +L ++ ++
Sbjct: 430 LSGPIPSGISLLKNLTDLTIT 492
Score = 58.9 bits (141), Expect = 4e-09
Identities = 42/150 (28%), Positives = 77/150 (51%)
Frame = +1
Query: 229 LSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEM 288
LSGTLP L RL + ++DL+ N+L G + ++ LK L + NRL + +
Sbjct: 1 LSGTLPPDLVRLPFLQEIDLTLNYLNGTIPKQWATLK-LVNVSFYGNRLSGPIPKEFGNI 177
Query: 289 NSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDN 348
+L+ +VL N L G++ + +L + L LS+ G +P + ++L L+ + D+
Sbjct: 178 TTLKSLVLEFNQLSGNLPP-ELGSLSQIERLLLSSNNFTGLLPATFAKLTALKQFRIGDS 354
Query: 349 NITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+G + +++ +L L + G+ L G I
Sbjct: 355 QFSGAIPNFIQSWINLEMLTIRGSGLSGPI 444
Score = 57.0 bits (136), Expect(2) = 3e-10
Identities = 33/79 (41%), Positives = 46/79 (57%)
Frame = +3
Query: 179 NGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLG 238
NG PQ + N+ L +LVL N SG +P+ G L++L ++DLS N LSG +PV+
Sbjct: 501 NGSDSPFPQ-LQNMSNLSKLVLRNCNISGALPEYLGKLTNLKVIDLSNNKLSGQIPVSFD 677
Query: 239 RLISVLKLDLSHNFLEGKL 257
L ++ L LS N L G L
Sbjct: 678 GLQNMYLLFLSGNQLSGSL 734
Score = 52.0 bits (123), Expect(2) = 3e-24
Identities = 26/77 (33%), Positives = 43/77 (55%)
Frame = +3
Query: 302 GGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETL 361
G D + +N+ NL L L N + G +PE L +L L+ + LS+N ++G + + L
Sbjct: 504 GSDSPFPQLQNMSNLSKLVLRNCNISGALPEYLGKLTNLKVIDLSNNKLSGQIPVSFDGL 683
Query: 362 PSLNALYLSGNNLKGEI 378
++ L+LSGN L G +
Sbjct: 684 QNMYLLFLSGNQLSGSL 734
Score = 47.4 bits (111), Expect = 1e-05
Identities = 31/89 (34%), Positives = 52/89 (57%)
Frame = +3
Query: 263 NLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELS 322
N+ NL+ + LRN + L L ++ +L+ + LSNN L G I + ++ LQN+ +L LS
Sbjct: 534 NMSNLSKLVLRNCNISGALPEYLGKLTNLKVIDLSNNKLSGQI-PVSFDGLQNMYLLFLS 710
Query: 323 NMELIGEIPESLSQLKKLRFLGLSDNNIT 351
+L G +P+ ++ K ++ LS NN T
Sbjct: 711 GNQLSGSLPDWIA---KPDYVDLSYNNFT 788
Score = 44.7 bits (104), Expect = 8e-05
Identities = 32/101 (31%), Positives = 48/101 (46%), Gaps = 6/101 (5%)
Frame = +3
Query: 117 AISFFN-CFQSPNKLPVSIPTGNWEKLAESLESIEFRS-----NPGLIGNIPSTFGVLKN 170
A SF+N F+ P+ L + + L+++ S N + G +P G L N
Sbjct: 438 ANSFWNFTFEEPH*LDYYLTLNGSDSPFPQLQNMSNLSKLVLRNCNISGALPEYLGKLTN 617
Query: 171 LQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPD 211
L+ + L N L+G IP L + L LSGN SG++PD
Sbjct: 618 LKVIDLSNNKLSGQIPVSFDGLQNMYLLFLSGNQLSGSLPD 740
Score = 25.4 bits (54), Expect(2) = 3e-10
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = +1
Query: 145 SLESIEFRSNPGLIGNIPSTFGVLKNLQSLVL 176
+LE + R + GL G IPS +LKNL L +
Sbjct: 397 NLEMLTIRGS-GLSGPIPSGISLLKNLTDLTI 489
>TC86633 weakly similar to SP|Q00874|D100_ARATH DNA-damage-repair/toleration
protein DRT100 precursor. [Mouse-ear cress] {Arabidopsis
thaliana}, partial (93%)
Length = 1407
Score = 108 bits (270), Expect = 4e-24
Identities = 96/338 (28%), Positives = 141/338 (41%), Gaps = 5/338 (1%)
Frame = +3
Query: 52 NSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINI-----GPIHENSLPCANEKLEFK 106
NSW+G D GVSC+ W VT IN+ PI +N + E
Sbjct: 159 NSWSGYDC------CRGWHGVSCNPTT--WRVTDINLRGDSEDPIFQNLTHSGDMTGEIS 314
Query: 107 PELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFG 166
PE+ +L L ++ +W+ ++ G IPS
Sbjct: 315 PEVCKLDEL----------------TTLVVADWKSIS---------------GEIPSCIT 401
Query: 167 VLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
L +L+ L L N ++GNIP IG L L L L+ N SG IP +S L+ LDL+
Sbjct: 402 SLSSLRILDLTGNKISGNIPGNIGKLQHLTVLNLADNAISGEIPMSIVRISGLMHLDLAG 581
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQ 286
N +SG LP +G+L + + S N L G + + + L +DL NR+ + +
Sbjct: 582 NQISGELPSDIGKLRRLSRALFSRNQLTGSIPDSVLKMNRLADLDLSMNRITGSIPARIG 761
Query: 287 EMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLS 346
+M L + L N + G I + N + IL LS G IP+ L LS
Sbjct: 762 KMRVLSTLKLDGNSMTGQIPSTLLSNT-GMGILNLSRNGFEGTIPDVFGSKSYFMVLDLS 938
Query: 347 DNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGF 384
N +TG + L + + L +S N+L G I F
Sbjct: 939 FNKLTGRIPGSLSSAKFMGHLDISNNHLCGTIPIGSPF 1052
Score = 80.5 bits (197), Expect = 1e-15
Identities = 59/175 (33%), Positives = 85/175 (47%)
Frame = +3
Query: 125 QSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGN 184
Q +LP I G +L+ +L F N L G+IP + + L L L N +TG+
Sbjct: 585 QISGELPSDI--GKLRRLSRAL----FSRNQ-LTGSIPDSVLKMNRLADLDLSMNRITGS 743
Query: 185 IPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVL 244
IP IG + L L L GN+ +G IP + + IL+LSRN GT+P G +
Sbjct: 744 IPARIGKMRVLSTLKLDGNSMTGQIPSTLLSNTGMGILNLSRNGFEGTIPDVFGSKSYFM 923
Query: 245 KLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNN 299
LDLS N L G++ + K + +D+ NN L CG + + L+ SNN
Sbjct: 924 VLDLSFNKLTGRIPGSLSSAKFMGHLDISNNHL-CGTIPIGSPFDHLDAASFSNN 1085
Score = 48.9 bits (115), Expect = 4e-06
Identities = 33/109 (30%), Positives = 58/109 (52%)
Frame = +3
Query: 282 VLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLR 341
V L E+ +L +V + G+I + +L +L IL+L+ ++ G IP ++ +L+ L
Sbjct: 321 VCKLDELTTL--VVADWKSISGEIPSCI-TSLSSLRILDLTGNKISGNIPGNIGKLQHLT 491
Query: 342 FLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGR 390
L L+DN I+G + + + L L L+GN + GE+ G +L R
Sbjct: 492 VLNLADNAISGEIPMSIVRISGLMHLDLAGNQISGELPSDIGKLRRLSR 638
>BF648743 weakly similar to PIR|H84632|H846 probable receptor-like protein
kinase [imported] - Arabidopsis thaliana, partial (9%)
Length = 648
Score = 107 bits (268), Expect = 8e-24
Identities = 73/207 (35%), Positives = 112/207 (53%), Gaps = 2/207 (0%)
Frame = +1
Query: 168 LKNLQSLVLLENGLTGN-IPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
L NL L+L N +G IP EIG L L L + +N G+IP G L++L +DLS+
Sbjct: 4 LTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSK 183
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNF-LEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSL 285
NSLSG +P T+G L + L LS+N + G + + N+ +LT++ N L + S+
Sbjct: 184 NSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSI 363
Query: 286 QEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGL 345
Q + +L+E+ L N L G I + + L+NL+ L L + L G IP S+ L L+ L +
Sbjct: 364 QNLVNLKELALDINHLSGSIPSTIGD-LKNLIKLYLGSNNLSGPIPASIGNLINLQVLSV 540
Query: 346 SDNNITGNLSPKLETLPSLNALYLSGN 372
+NN+TG + + L L + N
Sbjct: 541 QENNLTGTIPASIGNLKWLTVFEVXTN 621
Score = 102 bits (255), Expect = 2e-22
Identities = 73/195 (37%), Positives = 101/195 (51%), Gaps = 2/195 (1%)
Frame = +1
Query: 191 NLVKLKRLVLSGNNFSGN-IPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLS 249
NL L L+L GNN+SG IP G L++LL L + +++L G++P +G L ++ +DLS
Sbjct: 1 NLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLS 180
Query: 250 HNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVL-SLQEMNSLEEMVLSNNPLGGDIRTL 308
N L G + GNL L + L NN G + SL M+SL + N L G I
Sbjct: 181 KNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPD- 357
Query: 309 KWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALY 368
+NL NL L L L G IP ++ LK L L L NN++G + + L +L L
Sbjct: 358 SIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLS 537
Query: 369 LSGNNLKGEIQFSKG 383
+ NNL G I S G
Sbjct: 538 VQENNLTGTIPASIG 582
Score = 97.1 bits (240), Expect = 1e-20
Identities = 68/191 (35%), Positives = 98/191 (50%), Gaps = 1/191 (0%)
Frame = +1
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G IP G L NL L + ++ L G+IPQEIG L L + LS N+ SG IP+ G LS
Sbjct: 52 GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSK 231
Query: 219 LLILDLSRNS-LSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
L L LS N+ +SG +P +L + S+ L + L G + + NL NL + L N L
Sbjct: 232 LDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHL 411
Query: 278 CCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQL 337
+ ++ ++ +L ++ L +N L G I NL NL +L + L G IP S+ L
Sbjct: 412 SGSIPSTIGDLKNLIKLYLGSNNLSGPI-PASIGNLINLQVLSVQENNLTGTIPASIGNL 588
Query: 338 KKLRFLGLSDN 348
K L + N
Sbjct: 589 KWLTVFEVXTN 621
Score = 90.5 bits (223), Expect = 1e-18
Identities = 57/153 (37%), Positives = 83/153 (53%), Gaps = 2/153 (1%)
Frame = +1
Query: 126 SPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTG 183
S N L IP GN KL +++ +N + G IP + + +L L GL+G
Sbjct: 178 SKNSLSGGIPETIGNLSKL----DTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSG 345
Query: 184 NIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISV 243
+IP I NLV LK L L N+ SG+IP G L +L+ L L N+LSG +P ++G LI++
Sbjct: 346 SIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINL 525
Query: 244 LKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR 276
L + N L G + GNLK LT+ ++ N+
Sbjct: 526 QVLSVQENNLTGTIPASIGNLKWLTVFEVXTNK 624
Score = 85.5 bits (210), Expect = 4e-17
Identities = 58/168 (34%), Positives = 81/168 (47%), Gaps = 25/168 (14%)
Frame = +1
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNN------------ 204
L+G+IP G L NL + L +N L+G IP+ IGNL KL LVLS N
Sbjct: 118 LVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWN 297
Query: 205 -------------FSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHN 251
SG+IPD L +L L L N LSG++P T+G L +++KL L N
Sbjct: 298 MSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSN 477
Query: 252 FLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNN 299
L G + GNL NL ++ ++ N L + S+ + L + N
Sbjct: 478 NLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVXTN 621
Score = 64.3 bits (155), Expect = 1e-10
Identities = 35/78 (44%), Positives = 45/78 (56%)
Frame = +1
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G+IPST G LKNL L L N L+G IP IGNL+ L+ L + NN +G IP G L
Sbjct: 409 LSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNL 588
Query: 217 SDLLILDLSRNSLSGTLP 234
L + ++ N S +P
Sbjct: 589 KWLTVFEVXTNKPSWRIP 642
Score = 43.9 bits (102), Expect = 1e-04
Identities = 34/84 (40%), Positives = 44/84 (51%)
Frame = +1
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
N L SIP+ + ++L + SN L G IP++ G L NLQ L + EN LTG IP
Sbjct: 403 NHLSGSIPSTIGD--LKNLIKLYLGSN-NLSGPIPASIGNLINLQVLSVQENNLTGTIPA 573
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPD 211
IGNL L + N S IP+
Sbjct: 574 SIGNLKWLTVFEVXTNKPSWRIPN 645
>TC90678 weakly similar to GP|9759550|dbj|BAB11152.1 receptor protein
kinase-like protein {Arabidopsis thaliana}, partial
(48%)
Length = 1392
Score = 107 bits (267), Expect = 1e-23
Identities = 77/239 (32%), Positives = 122/239 (50%), Gaps = 2/239 (0%)
Frame = +2
Query: 142 LAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENG-LTGNIPQEIGNLVKLKRLVL 200
+ ESL+ + + L+ +P+ FG L L+ L L N L IP+++G L LK+L+L
Sbjct: 512 IEESLKCLTWEVTCFLV-MVPNVFGNLTKLEVLDLSMNPYLVSEIPEDVGELGNLKQLLL 688
Query: 201 SGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTL-GRLISVLKLDLSHNFLEGKLLN 259
G++F G +P+ GL L LDLS N+L+G + TL L++++ D+S N L G N
Sbjct: 689 QGSSFQGEVPESLKGLISLTHLDLSENNLTGEVSKTLVSSLMNLVSFDVSQNKLLGSFPN 868
Query: 260 EFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVIL 319
K L + L NR + S E SLE + NN GD + + +L + ++
Sbjct: 869 GLCKGKGLINLSLHTNRFTGLIPNSTSECKSLERFQVQNNGFSGDFPIVLF-SLPKIKLI 1045
Query: 320 ELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
N G+IPES+S+ +L + L +N + G + L + SL S N+ GE+
Sbjct: 1046RGENNRFTGKIPESISEAVQLEQVQLDNNLLDGKIPSGLGFVKSLYRFSASLNHFYGEL 1222
Score = 85.1 bits (209), Expect = 5e-17
Identities = 73/265 (27%), Positives = 114/265 (42%), Gaps = 48/265 (18%)
Frame = +2
Query: 146 LESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNF 205
LE ++ NP L+ IP G L NL+ L+L + G +P+ + L+ L L LS NN
Sbjct: 596 LEVLDLSMNPYLVSEIPEDVGELGNLKQLLLQGSSFQGEVPESLKGLISLTHLDLSENNL 775
Query: 206 SGNIPD-IFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNL 264
+G + + L +L+ D+S+N L G+ P L + ++ L L N G + N
Sbjct: 776 TGEVSKTLVSSLMNLVSFDVSQNKLLGSFPNGLCKGKGLINLSLHTNRFTGLIPNSTSEC 955
Query: 265 KNLTLMDLRNN------------------------RLCCGLVLSLQEMNSLEEMVLSNNP 300
K+L ++NN R + S+ E LE++ L NN
Sbjct: 956 KSLERFQVQNNGFSGDFPIVLFSLPKIKLIRGENNRFTGKIPESISEAVQLEQVQLDNNL 1135
Query: 301 LGGDIRT--------LKWENLQN---------------LVILELSNMELIGEIPESLSQL 337
L G I + ++ N + I+ LS+ L G IP+ L +
Sbjct: 1136LDGKIPSGLGFVKSLYRFSASLNHFYGELPPNFCDSPVMSIVNLSHNSLSGSIPQ-LKKC 1312
Query: 338 KKLRFLGLSDNNITGNLSPKLETLP 362
KKL L L+DN++TG + L LP
Sbjct: 1313KKLVSLSLADNSLTGXIPNSLAELP 1387
Score = 68.2 bits (165), Expect = 7e-12
Identities = 62/212 (29%), Positives = 95/212 (44%)
Frame = +2
Query: 126 SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
S NKL S P G + + L ++ +N G IP++ K+L+ + NG +G+
Sbjct: 836 SQNKLLGSFPNGLCK--GKGLINLSLHTNR-FTGLIPNSTSECKSLERFQVQNNGFSGDF 1006
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
P + +L K+K + N F+G IP+ L + L N L G +P LG + S+ +
Sbjct: 1007 PIVLFSLPKIKLIRGENNRFTGKIPESISEAVQLEQVQLDNNLLDGKIPSGLGFVKSLYR 1186
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
S N G+L F C V+S+ + LS+N L G I
Sbjct: 1187 FSASLNHFYGELPPNF----------------CDSPVMSI--------VNLSHNSLSGSI 1294
Query: 306 RTLKWENLQNLVILELSNMELIGEIPESLSQL 337
LK + LV L L++ L G IP SL++L
Sbjct: 1295 PQLK--KCKKLVSLSLADNSLTGXIPNSLAEL 1384
Score = 61.2 bits (147), Expect = 8e-10
Identities = 39/98 (39%), Positives = 55/98 (55%)
Frame = +3
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSL 229
NLQSL L+G+I I +L L L L+ N F+ IP S L L+LS N +
Sbjct: 246 NLQSL-----NLSGDISSSICDLPSLSYLNLANNIFNQPIPLHLSQCSSLKSLNLSNNLI 410
Query: 230 SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNL 267
GT+P + + +S+ LDLS N +EG + + G+LKNL
Sbjct: 411 WGTIPSQISQFVSLSVLDLSRNHIEGNIPDSLGSLKNL 524
Score = 49.3 bits (116), Expect = 3e-06
Identities = 35/96 (36%), Positives = 46/96 (47%)
Frame = +3
Query: 145 SLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNN 204
S+ S+ +S L G+I S+ L +L L L N IP + LK L LS N
Sbjct: 231 SVTSVNLQSL-NLSGDISSSICDLPSLSYLNLANNIFNQPIPLHLSQCSSLKSLNLSNNL 407
Query: 205 FSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRL 240
G IP L +LDLSRN + G +P +LG L
Sbjct: 408 IWGTIPSQISQFVSLSVLDLSRNHIEGNIPDSLGSL 515
Score = 43.9 bits (102), Expect = 1e-04
Identities = 36/124 (29%), Positives = 56/124 (45%)
Frame = +3
Query: 241 ISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNP 300
+SV ++L L G + + +L +L+ ++L NN + L L + +SL+ + LSNN
Sbjct: 228 LSVTSVNLQSLNLSGDISSSICDLPSLSYLNLANNIFNQPIPLHLSQCSSLKSLNLSNN- 404
Query: 301 LGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLET 360
L W G IP +SQ L L LS N+I GN+ L +
Sbjct: 405 -------LIW-----------------GTIPSQISQFVSLSVLDLSRNHIEGNIPDSLGS 512
Query: 361 LPSL 364
L +L
Sbjct: 513 LKNL 524
Score = 32.7 bits (73), Expect = 0.31
Identities = 24/81 (29%), Positives = 37/81 (45%)
Frame = +3
Query: 115 LKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSL 174
L ++S+ N + P+ + SL+S+ SN + G IPS +L L
Sbjct: 297 LPSLSYLNLANNIFNQPIPLHLSQ----CSSLKSLNL-SNNLIWGTIPSQISQFVSLSVL 461
Query: 175 VLLENGLTGNIPQEIGNLVKL 195
L N + GNIP +G+L L
Sbjct: 462 DLSRNHIEGNIPDSLGSLKNL 524
>AW299082 weakly similar to GP|10177183|d receptor protein kinase-like
protein {Arabidopsis thaliana}, partial (16%)
Length = 565
Score = 105 bits (263), Expect = 3e-23
Identities = 59/185 (31%), Positives = 104/185 (55%)
Frame = +2
Query: 156 GLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGG 215
GL+G IP+ G +L+++ L N L+G IP +G+L++L+ ++S NN SG+IP
Sbjct: 11 GLVGAIPNEIGNCSSLRNIDLSLNSLSGTIPLSLGSLLELEEFMISDNNVSGSIPATLSN 190
Query: 216 LSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
+L L + N LSG +P +G+L ++L N LEG + + GN L +DL N
Sbjct: 191 AENLQQLQVDTNQLSGLIPPEIGKLSNLLVFFAWQNQLEGSIPSSLGNCSKLQALDLSRN 370
Query: 276 RLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLS 335
L + L ++ +L +++L +N + G I + + + ++L+ L L N + G IP+++
Sbjct: 371 SLTGSIPSRLFQLQNLTKLLLISNDISGSIPS-EIGSCKSLIRLRLGNNRITGSIPKTIR 547
Query: 336 QLKKL 340
L+ L
Sbjct: 548 NLRNL 562
Score = 105 bits (261), Expect = 5e-23
Identities = 68/188 (36%), Positives = 101/188 (53%)
Frame = +2
Query: 178 ENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTL 237
+NGL G IP EIGN L+ + LS N+ SG IP G L +L +S N++SG++P TL
Sbjct: 5 QNGLVGAIPNEIGNCSSLRNIDLSLNSLSGTIPLSLGSLLELEEFMISDNNVSGSIPATL 184
Query: 238 GRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLS 297
++ +L + N L G + E G L NL + N+L + SL + L+ + LS
Sbjct: 185 SNAENLQQLQVDTNQLSGLIPPEIGKLSNLLVFFAWQNQLEGSIPSSLGNCSKLQALDLS 364
Query: 298 NNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPK 357
N L G I + ++ LQNL L L + ++ G IP + K L L L +N ITG++
Sbjct: 365 RNSLTGSIPSRLFQ-LQNLTKLLLISNDISGSIPSEIGSCKSLIRLRLGNNRITGSIPKT 541
Query: 358 LETLPSLN 365
+ L +LN
Sbjct: 542 IRNLRNLN 565
Score = 90.1 bits (222), Expect = 2e-18
Identities = 64/174 (36%), Positives = 92/174 (52%), Gaps = 1/174 (0%)
Frame = +2
Query: 119 SFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFR-SNPGLIGNIPSTFGVLKNLQSLVLL 177
S N S N L +IP L LE EF S+ + G+IP+T +NLQ L +
Sbjct: 53 SLRNIDLSLNSLSGTIPLS----LGSLLELEEFMISDNNVSGSIPATLSNAENLQQLQVD 220
Query: 178 ENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTL 237
N L+G IP EIG L L N G+IP G S L LDLSRNSL+G++P L
Sbjct: 221 TNQLSGLIPPEIGKLSNLLVFFAWQNQLEGSIPSSLGNCSKLQALDLSRNSLTGSIPSRL 400
Query: 238 GRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSL 291
+L ++ KL L N + G + +E G+ K+L + L NNR+ + +++ + +L
Sbjct: 401 FQLQNLTKLLLISNDISGSIPSEIGSCKSLIRLRLGNNRITGSIPKTIRNLRNL 562
>TC81898 weakly similar to GP|20260576|gb|AAM13186.1 putative receptor-like
protein kinase {Arabidopsis thaliana}, partial (29%)
Length = 1103
Score = 102 bits (254), Expect = 3e-22
Identities = 83/265 (31%), Positives = 127/265 (47%), Gaps = 22/265 (8%)
Frame = +3
Query: 146 LESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNF 205
L S++ +N I +PS F L +L+SL L N ++G++ IGN L+ LS N+F
Sbjct: 288 LHSLDLSNNK--ITTLPSDFWSLTSLKSLNLSSNHISGSLTNNIGNFGLLENFDLSKNSF 461
Query: 206 SGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGN-L 264
S IP+ L L +L L N ++P + + S++ +DLS N L G L + FG+
Sbjct: 462 SDEIPEALSSLVSLKVLKLDHNMFVRSIPSGILKCQSLVSIDLSSNQLSGTLPHGFGDAF 641
Query: 265 KNLTLMDLRNNRLCCGL-----VLSLQEMN----------------SLEEMVLSNNPLGG 303
L ++L N + G+ + S+ +N LE + LS N G
Sbjct: 642 PKLRTLNLAENNIYGGVSNFSRLKSIVSLNISGNSFQGSIIEVFVLKLEALDLSRNQFQG 821
Query: 304 DIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPS 363
I +K+ N +LV L+LS +L GEI ++L+ L+ L L+ N + PK+E L
Sbjct: 822 HISQVKY-NWSHLVYLDLSENQLSGEIFQNLNNSMNLKHLSLACNRFSRQKFPKIEMLLG 998
Query: 364 LNALYLSGNNLKGEIQFSKGFFGKL 388
L L LS +L G I G L
Sbjct: 999 LEYLNLSKTSLVGHIPDEISHLGDL 1073
Score = 80.5 bits (197), Expect = 1e-15
Identities = 78/273 (28%), Positives = 117/273 (42%), Gaps = 48/273 (17%)
Frame = +3
Query: 126 SPNKLPVSIPTGNWEKLAESLESIEFRSN--PGLIGNIPSTFGVLKNLQSLVLLENGLTG 183
S NK+ ++P+ W SL+S+ SN G + N FG+L+N L +N +
Sbjct: 306 SNNKI-TTLPSDFWS--LTSLKSLNLSSNHISGSLTNNIGNFGLLENFD---LSKNSFSD 467
Query: 184 NIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLG----- 238
IP+ + +LV LK L L N F +IP L+ +DLS N LSGTLP G
Sbjct: 468 EIPEALSSLVSLKVLKLDHNMFVRSIPSGILKCQSLVSIDLSSNQLSGTLPHGFGDAFPK 647
Query: 239 -------------------RLISVLKLDLSHNFLEGKLLNEFG----------------- 262
RL S++ L++S N +G ++ F
Sbjct: 648 LRTLNLAENNIYGGVSNFSRLKSIVSLNISGNSFQGSIIEVFVLKLEALDLSRNQFQGHI 827
Query: 263 -----NLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLV 317
N +L +DL N+L + +L +L+ + L+ N + K E L L
Sbjct: 828 SQVKYNWSHLVYLDLSENQLSGEIFQNLNNSMNLKHLSLACNRFSRQ-KFPKIEMLLGLE 1004
Query: 318 ILELSNMELIGEIPESLSQLKKLRFLGLSDNNI 350
L LS L+G IP+ +S L L L LS N++
Sbjct: 1005YLNLSKTSLVGHIPDEISHLGDLNALDLSMNHL 1103
>TC77085 similar to GP|21536600|gb|AAM60932.1 putative disease resistance
protein {Arabidopsis thaliana}, partial (88%)
Length = 1662
Score = 101 bits (252), Expect = 5e-22
Identities = 71/214 (33%), Positives = 111/214 (51%), Gaps = 7/214 (3%)
Frame = +2
Query: 142 LAESLESIEFRSNPGLI------GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKL 195
++ SL ++F LI G P L NL+ + + N L+G IPQ IG++ +L
Sbjct: 326 ISPSLSKLKFLDGIYLINLLKISGPFPDFLFKLPNLKYIYIENNTLSGPIPQNIGSMNQL 505
Query: 196 KRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEG 255
+ L N F+G IP L+ L L L N L+GT+PV+L L ++ L L N L G
Sbjct: 506 EAFSLQENKFTGPIPSSISALTKLTQLKLGNNFLTGTIPVSLKNLTNLTYLSLQGNQLSG 685
Query: 256 KLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEM-NSLEEMVLSNNPLGGDIRTLKWENLQ 314
+ + F +LKNL ++ L +N+ + LS+ + +L + L +N L G I + +
Sbjct: 686 NIPDIFTSLKNLIILQLSHNKFSGNIPLSISSLYPTLRYLELGHNSLSGKIPDFLGK-FK 862
Query: 315 NLVILELSNMELIGEIPESLSQLKKLRFLGLSDN 348
L L+LS + G +P+S + L K+ L LSDN
Sbjct: 863 ALDTLDLSKNQFKGTVPKSFANLTKIFNLDLSDN 964
Score = 94.7 bits (234), Expect = 7e-20
Identities = 88/279 (31%), Positives = 120/279 (42%), Gaps = 29/279 (10%)
Frame = +2
Query: 129 KLPVSIPTGNWEKLAESLESIEFRSNPG--LIGNIPSTFGVLKNLQSLVLLENGLTGNIP 186
KL + TG ++L ++ + S G L GNIP F LKNL L L N +GNIP
Sbjct: 587 KLGNNFLTGTIPVSLKNLTNLTYLSLQGNQLSGNIPDIFTSLKNLIILQLSHNKFSGNIP 766
Query: 187 QEIGNLVK-LKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
I +L L+ L L N+ SG IPD G L LDLS+N GT+P + L +
Sbjct: 767 LSISSLYPTLRYLELGHNSLSGKIPDFLGKFKALDTLDLSKNQFKGTVPKSFANLTKIFN 946
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL---------------------CCGLVLS 284
LDLS NFL N+K + +DL N CG+ +
Sbjct: 947 LDLSDNFLVDPF--PVMNVKGIESLDLSRNMFHLKEIPKWVATSPIIYSLKLAHCGIKMK 1120
Query: 285 LQEMNSLEEMV-----LSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKK 339
L + LE LS N + G L N +I + L+ ESL +
Sbjct: 1121LDDWKPLETFFYDYIDLSGNEISGSAVGLL--NKTEYLIEFRGSENLLKFDLESLKFGNR 1294
Query: 340 LRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L++L LS N + G ++ + + LN Y N L GEI
Sbjct: 1295LKYLDLSHNLVFGKVTKSVVGIQKLNVSY---NRLCGEI 1402
>AW585664 similar to PIR|T46033|T46 receptor protein kinase-like protein -
Arabidopsis thaliana, partial (4%)
Length = 640
Score = 100 bits (250), Expect = 9e-22
Identities = 71/221 (32%), Positives = 105/221 (47%), Gaps = 1/221 (0%)
Frame = +1
Query: 159 GNIPSTFGVLK-NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLS 217
G +P+ G L L L L N ++G IP+E+GNLV L L + N+F G IP FG
Sbjct: 34 GCLPNFVGNLSFQLSELYLGGNEISGKIPEELGNLVNLTLLSMGHNHFEGIIPANFGKFQ 213
Query: 218 DLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
+ LDL +N LSG +P +G L + L + N LEG
Sbjct: 214 SMQRLDLRQNKLSGDIPYFIGNLSQLFDLHMEENMLEG---------------------- 327
Query: 278 CCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQL 337
+ LS+ E L+ + LS N L G I + L+LS L G +P+ + L
Sbjct: 328 --NIPLSIGECQMLQYLNLSQNNLQGAIPLEIFSIFSLTTGLDLSQNSLSGSLPDEVGLL 501
Query: 338 KKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
K + L +S+N+++G++ + SL L+L GN+L G I
Sbjct: 502 KNIHKLDVSENHLSGDIPITIGECISLEYLHLQGNSLHGTI 624
Score = 60.1 bits (144), Expect = 2e-09
Identities = 50/182 (27%), Positives = 75/182 (40%), Gaps = 1/182 (0%)
Frame = +1
Query: 198 LVLSGNNFSGNIPDIFGGLS-DLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGK 256
L L+ NNF G +P+ G LS L L L N +SG +P LG L+++ L + HN EG
Sbjct: 7 LSLAANNFGGCLPNFVGNLSFQLSELYLGGNEISGKIPEELGNLVNLTLLSMGHNHFEGI 186
Query: 257 LLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNL 316
+ FG K++++Q L
Sbjct: 187 IPANFG----------------------------------------------KFQSMQRL 228
Query: 317 VILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKG 376
+L +L G+IP + L +L L + +N + GN+ + L L LS NNL+G
Sbjct: 229 ---DLRQNKLSGDIPYFIGNLSQLFDLHMEENMLEGNIPLSIGECQMLQYLNLSQNNLQG 399
Query: 377 EI 378
I
Sbjct: 400 AI 405
Score = 50.4 bits (119), Expect = 1e-06
Identities = 25/58 (43%), Positives = 35/58 (60%)
Frame = +1
Query: 153 SNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
S L G++P G+LKN+ L + EN L+G+IP IG + L+ L L GN+ G IP
Sbjct: 454 SQNSLSGSLPDEVGLLKNIHKLDVSENHLSGDIPITIGECISLEYLHLQGNSLHGTIP 627
>BI308817 weakly similar to PIR|B84659|B846 probable receptor-like protein
kinase ERECTA [imported] - Arabidopsis thaliana,
partial (6%)
Length = 622
Score = 100 bits (249), Expect = 1e-21
Identities = 70/198 (35%), Positives = 97/198 (48%)
Frame = +2
Query: 167 VLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
+LKNL L L N G IP + NL +L+ L +S NN G +P L +L LDLS
Sbjct: 47 LLKNLTFLYLSYNKFKGEIPSSLENLKQLEDLDISYNNLKGQLPPELWLLKNLTFLDLSY 226
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQ 286
N G +P +LG L + L +S+N++EG + E LKN+ DL NNR
Sbjct: 227 NMFKGEIPSSLGNLTQLEDLYISNNYIEGHIPFELVFLKNMITFDLSNNR---------- 376
Query: 287 EMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLS 346
L ++ S+N L G + N + L +L +S+ + G IP L LK L L LS
Sbjct: 377 ----LTDLDFSSNYLKGQV-----GNPKQLQLLNISHNNIQGSIPLELGFLKNLTILDLS 529
Query: 347 DNNITGNLSPKLETLPSL 364
N + GN S + L L
Sbjct: 530 HNRLNGNFSIFVSNLTQL 583
Score = 88.6 bits (218), Expect = 5e-18
Identities = 66/196 (33%), Positives = 101/196 (50%), Gaps = 6/196 (3%)
Frame = +2
Query: 103 LEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIP 162
+ + PEL+ LK+L + + S NK IP+ + E L + LE ++ N L G +P
Sbjct: 23 VSYHPELWLLKNLTFL-----YLSYNKFKGEIPS-SLENLKQ-LEDLDISYN-NLKGQLP 178
Query: 163 STFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLIL 222
+LKNL L L N G IP +GNL +L+ L +S N G+IP L +++
Sbjct: 179 PELWLLKNLTFLDLSYNMFKGEIPSSLGNLTQLEDLYISNNYIEGHIPFELVFLKNMITF 358
Query: 223 DLSRNSL------SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR 276
DLS N L S L +G + L++SHN ++G + E G LKNLT++DL +NR
Sbjct: 359 DLSNNRLTDLDFSSNYLKGQVGNPKQLQLLNISHNNIQGSIPLELGFLKNLTILDLSHNR 538
Query: 277 LCCGLVLSLQEMNSLE 292
L + + + L+
Sbjct: 539 LNGNFSIFVSNLTQLQ 586
Score = 59.7 bits (143), Expect = 2e-09
Identities = 41/100 (41%), Positives = 55/100 (55%)
Frame = +2
Query: 313 LQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
L+NL L LS + GEIP SL LK+L L +S NN+ G L P+L L +L L LS N
Sbjct: 50 LKNLTFLYLSYNKFKGEIPSSLENLKQLEDLDISYNNLKGQLPPELWLLKNLTFLDLSYN 229
Query: 373 NLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPFELMSTNNV 412
KGEI S G +L + + + + PFEL+ N+
Sbjct: 230 MFKGEIPSSLGNLTQLEDLYISNNYIEGHIPFELVFLKNM 349
Score = 28.1 bits (61), Expect = 7.7
Identities = 21/59 (35%), Positives = 29/59 (48%)
Frame = +2
Query: 356 PKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPFELMSTNNVPY 414
P+L L +L LYLS N KGEI S +L +++N K P EL N+ +
Sbjct: 35 PELWLLKNLTFLYLSYNKFKGEIPSSLENLKQLEDLDISYNNLKGQLPPELWLLKNLTF 211
>TC79788 weakly similar to GP|21554189|gb|AAM63268.1 putative leucine-rich
repeat disease resistance protein {Arabidopsis
thaliana}, partial (52%)
Length = 1484
Score = 99.4 bits (246), Expect = 3e-21
Identities = 81/245 (33%), Positives = 114/245 (46%), Gaps = 26/245 (10%)
Frame = +3
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G++ L L +L L +N G+IP I +L LK L L N+FSG IP L
Sbjct: 357 GSLTPLISKLTQLITLDLSDNNFFGSIPSSISSLSNLKTLTLRSNSFSGPIPPSIISLKS 536
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
L LDLS NSL+G+LP +L LI++ ++DLS N L G + NL L ++ N L
Sbjct: 537 LESLDLSHNSLTGSLPNSLNSLINLHRIDLSFNKLAGSIPKLPPNLLELA---IKANSLS 707
Query: 279 CGL-VLSLQEMNSLEEMVLSNNPLGGDIRT-------LKWENL----------------- 313
L + + N LE + LS N L G I T L+ NL
Sbjct: 708 GPLQKTTFEGSNQLEVVELSENALTGTIETWFLLLSSLQQVNLANNSFTGIQISKPARGV 887
Query: 314 -QNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
NLV L L + G P +L+ L FL + N++ GN+ + + S+ L+L GN
Sbjct: 888 ESNLVALNLGFNRIQGYAPANLAAYPLLSFLSIRHNSLRGNIPLEYGQIKSMKRLFLDGN 1067
Query: 373 NLKGE 377
G+
Sbjct: 1068FFDGK 1082
Score = 94.0 bits (232), Expect = 1e-19
Identities = 71/209 (33%), Positives = 104/209 (48%), Gaps = 2/209 (0%)
Frame = +3
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSL 229
++ + L +G++ I L +L L LS NNF G+IP LS+L L L NS
Sbjct: 318 HINQITLDPASYSGSLTPLISKLTQLITLDLSDNNFFGSIPSSISSLSNLKTLTLRSNSF 497
Query: 230 SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMN 289
SG +P ++ L S+ LDLSHN L G L N +L NL +DL N+L G + L
Sbjct: 498 SGPIPPSIISLKSLESLDLSHNSLTGSLPNSLNSLINLHRIDLSFNKL-AGSIPKLPP-- 668
Query: 290 SLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNN 349
+L E+ + N L G ++ +E L ++ELS L G I L L+ + L++N+
Sbjct: 669 NLLELAIKANSLSGPLQKTTFEGSNQLEVVELSENALTGTIETWFLLLSSLQQVNLANNS 848
Query: 350 ITG--NLSPKLETLPSLNALYLSGNNLKG 376
TG P +L AL L N ++G
Sbjct: 849 FTGIQISKPARGVESNLVALNLGFNRIQG 935
Score = 40.8 bits (94), Expect = 0.001
Identities = 21/52 (40%), Positives = 25/52 (47%)
Frame = +3
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
G P+ L L + N L GNIP E G + +KRL L GN F G P
Sbjct: 933 GYAPANLAAYPLLSFLSIRHNSLRGNIPLEYGQIKSMKRLFLDGNFFDGKPP 1088
>BM779095 weakly similar to PIR|B84431|B8 probable receptor protein kinase
[imported] - Arabidopsis thaliana, partial (15%)
Length = 721
Score = 97.4 bits (241), Expect = 1e-20
Identities = 77/227 (33%), Positives = 114/227 (49%), Gaps = 5/227 (2%)
Frame = +3
Query: 157 LIGNIPSTFGVLKN---LQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIF 213
L G IP F ++N LQSL L N LTG+I + L +L LSGNN S + P
Sbjct: 39 LTGPIPDNF--IQNSDKLQSLDLSFNNLTGSISDIKIDCKSLLQLDLSGNNLSDSFPISL 212
Query: 214 GGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNL-KNLTLMDL 272
+ L L+L+ N +SG +P LG+L + LDLSHN + G + +E N+ +L + L
Sbjct: 213 SNCTSLKSLNLASNFISGGIPKALGQLNKLQSLDLSHNQITGWIPSELSNVCGSLLELKL 392
Query: 273 RNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPE 332
N + + L+ + +SNN + G++ +L +L L L N + + P
Sbjct: 393 SFNNITGSVPFGFNSCTWLQLVDISNNNMTGELPESVIRSLGSLQELRLGNNAISMKFPS 572
Query: 333 SLSQLKKLRFLGLSDNNITGNLSPKL-ETLPSLNALYLSGNNLKGEI 378
S+S LKKLR + S N I G++ L SL L + N + GEI
Sbjct: 573 SISSLKKLRIVDFSSNKIYGSIPRDLCPGAGSLEELRMPDNLITGEI 713
Score = 95.5 bits (236), Expect = 4e-20
Identities = 69/203 (33%), Positives = 107/203 (51%), Gaps = 3/203 (1%)
Frame = +3
Query: 179 NGLTGNIPQE-IGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTL 237
+ LTG IP I N KL+ L LS NN +G+I DI LL LDLS N+LS + P++L
Sbjct: 33 HNLTGPIPDNFIQNSDKLQSLDLSFNNLTGSISDIKIDCKSLLQLDLSGNNLSDSFPISL 212
Query: 238 GRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEM-NSLEEMVL 296
S+ L+L+ NF+ G + G L L +DL +N++ + L + SL E+ L
Sbjct: 213 SNCTSLKSLNLASNFISGGIPKALGQLNKLQSLDLSHNQITGWIPSELSNVCGSLLELKL 392
Query: 297 SNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPES-LSQLKKLRFLGLSDNNITGNLS 355
S N + G + + + L ++++SN + GE+PES + L L+ L L +N I+
Sbjct: 393 SFNNITGSV-PFGFNSCTWLQLVDISNNNMTGELPESVIRSLGSLQELRLGNNAISMKFP 569
Query: 356 PKLETLPSLNALYLSGNNLKGEI 378
+ +L L + S N + G I
Sbjct: 570 SSISSLKKLRIVDFSSNKIYGSI 638
Score = 36.6 bits (83), Expect = 0.022
Identities = 25/57 (43%), Positives = 31/57 (53%), Gaps = 1/57 (1%)
Frame = +3
Query: 326 LIGEIPESLSQLK-KLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFS 381
L G IP++ Q KL+ L LS NN+TG++S SL L LSGNNL S
Sbjct: 39 LTGPIPDNFIQNSDKLQSLDLSFNNLTGSISDIKIDCKSLLQLDLSGNNLSDSFPIS 209
>TC89364 weakly similar to PIR|C96745|C96745 hypothetical protein T9N14.3
[imported] - Arabidopsis thaliana, partial (45%)
Length = 1903
Score = 97.1 bits (240), Expect = 1e-20
Identities = 56/147 (38%), Positives = 81/147 (55%)
Frame = +1
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G + S G NL +VL+ N +G +P EIG LV L++L LS NNFSG+IP G L
Sbjct: 106 GEVSSEIGYSTNLSEIVLMNNKFSGKVPSEIGKLVNLEKLYLSNNNFSGDIPREIGLLKQ 285
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
L L L NSL+G +P LG ++ L+L+ N L G + N + +L ++L N+L
Sbjct: 286 LSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIPNSVSLMSSLNSLNLSRNKLT 465
Query: 279 CGLVLSLQEMNSLEEMVLSNNPLGGDI 305
+ +L++M L + S N L G I
Sbjct: 466 GTIPDNLEKM-KLSSVDFSQNSLSGGI 543
Score = 79.0 bits (193), Expect = 4e-15
Identities = 46/105 (43%), Positives = 65/105 (61%)
Frame = +1
Query: 151 FRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
+ SN G+IP G+LK L +L L EN LTG IP+E+G+ +L L L+ N+ SGNIP
Sbjct: 226 YLSNNNFSGDIPREIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIP 405
Query: 211 DIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEG 255
+ +S L L+LSRN L+GT+P L ++ + +D S N L G
Sbjct: 406 NSVSLMSSLNSLNLSRNKLTGTIPDNLEKM-KLSSVDFSQNSLSG 537
Score = 78.2 bits (191), Expect = 6e-15
Identities = 56/166 (33%), Positives = 81/166 (48%), Gaps = 2/166 (1%)
Frame = +1
Query: 168 LKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRN 227
L N + + L N +G + EIG L +VL N FSG +P G L +L L LS N
Sbjct: 61 LPNAKIIDLGFNNFSGEVSSEIGYSTNLSEIVLMNNKFSGKVPSEIGKLVNLEKLYLSNN 240
Query: 228 SLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQE 287
+ SG +P +G L + L L N L G + E G+ L ++L N L + S+
Sbjct: 241 NFSGDIPREIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIPNSVSL 420
Query: 288 MNSLEEMVLSNNPLGGDIRTLKWENLQNLVI--LELSNMELIGEIP 331
M+SL + LS N L G I +NL+ + + ++ S L G IP
Sbjct: 421 MSSLNSLNLSRNKLTGTIP----DNLEKMKLSSVDFSQNSLSGGIP 546
Score = 68.9 bits (167), Expect = 4e-12
Identities = 45/133 (33%), Positives = 71/133 (52%)
Frame = +1
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
+DL N G++ +E G NL+ + L NN+ + + ++ +LE++ LSNN GDI
Sbjct: 79 IDLGFNNFSGEVSSEIGYSTNLSEIVLMNNKFSGKVPSEIGKLVNLEKLYLSNNNFSGDI 258
Query: 306 RTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLN 365
+ L+ L L L L G IP+ L +L L L+ N+++GN+ + + SLN
Sbjct: 259 PR-EIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIPNSVSLMSSLN 435
Query: 366 ALYLSGNNLKGEI 378
+L LS N L G I
Sbjct: 436 SLNLSRNKLTGTI 474
Score = 53.5 bits (127), Expect = 2e-07
Identities = 40/126 (31%), Positives = 60/126 (46%)
Frame = +1
Query: 263 NLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELS 322
+L N ++DL N + + +L E+VL NN G + + + L NL L LS
Sbjct: 58 SLPNAKIIDLGFNNFSGEVSSEIGYSTNLSEIVLMNNKFSGKVPS-EIGKLVNLEKLYLS 234
Query: 323 NMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSK 382
N G+IP + LK+L L L +N++TG + +L L L L+ N+L G I S
Sbjct: 235 NNNFSGDIPREIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIPNSV 414
Query: 383 GFFGKL 388
L
Sbjct: 415 SLMSSL 432
>BF645340 weakly similar to GP|6016693|gb|A putative disease resistance
protein {Arabidopsis thaliana}, partial (7%)
Length = 655
Score = 97.1 bits (240), Expect = 1e-20
Identities = 72/209 (34%), Positives = 106/209 (50%), Gaps = 1/209 (0%)
Frame = +2
Query: 126 SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
S NKL ++ NW L E+ S+ N L +I L L L N L GN+
Sbjct: 29 SNNKLNGTV--SNW--LLETSRSLNLSQN--LFTSIDQISRNSDQLGDLDLSFNLLVGNL 190
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
I NL L+ L L NNF+GNIP L L ILDL N+ GTLP + ++
Sbjct: 191 SVSICNLSSLEFLNLGHNNFTGNIPQCLANLPSLQILDLQMNNFYGTLPNNFSKSSKLIT 370
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
L+L+ N LEG + +NL +++LRNN++ + LQ + L+ +VL +N L G I
Sbjct: 371 LNLNDNQLEGYFPKSLSHCENLQVLNLRNNKMEDKFPVWLQTLQYLKVLVLRDNKLHGHI 550
Query: 306 RTLKWEN-LQNLVILELSNMELIGEIPES 333
LK + +LVI ++S+ G +P++
Sbjct: 551 ANLKIRHPFPSLVIFDISSNNFTGPLPKA 637
Score = 52.0 bits (123), Expect = 5e-07
Identities = 39/117 (33%), Positives = 51/117 (43%), Gaps = 22/117 (18%)
Frame = +2
Query: 288 MNSLEEMVLSNNPLGGDIRTLKWENLQNLVI--------------------LELSNMELI 327
+ LE + LSNN L G + E ++L + L+LS L+
Sbjct: 2 LGKLESLDLSNNKLNGTVSNWLLETSRSLNLSQNLFTSIDQISRNSDQLGDLDLSFNLLV 181
Query: 328 GEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI--QFSK 382
G + S+ L L FL L NN TGN+ L LPSL L L NN G + FSK
Sbjct: 182 GNLSVSICNLSSLEFLNLGHNNFTGNIPQCLANLPSLQILDLQMNNFYGTLPNNFSK 352
>TC82638 weakly similar to PIR|T00712|T00712 protein kinase homolog F22O13.7
- Arabidopsis thaliana, partial (27%)
Length = 1074
Score = 95.1 bits (235), Expect = 5e-20
Identities = 54/137 (39%), Positives = 76/137 (55%)
Frame = +2
Query: 165 FGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDL 224
FG L +L+ +++ N G IP+E GNL KLK L L+ N G IPD G L L + L
Sbjct: 632 FGKLSSLEYMIIGYNEFEGGIPKEFGNLTKLKYLDLAEGNVGGEIPDELGKLKLLNTVFL 811
Query: 225 SRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLS 284
+NS G +P +G + S++ LDLS N L G + E LKNL L++ N+L +
Sbjct: 812 YKNSFEGKIPTNIGNMTSLVLLDLSDNMLSGNIPAEISQLKNLQLLNFMRNKLSGPVPSG 991
Query: 285 LQEMNSLEEMVLSNNPL 301
L ++ LE + L NN L
Sbjct: 992 LGDLPQLEVLELWNNSL 1042
Score = 84.0 bits (206), Expect = 1e-16
Identities = 46/119 (38%), Positives = 67/119 (55%)
Frame = +2
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G IP FG L L+ L L E + G IP E+G L L + L N+F G IP G ++
Sbjct: 686 GGIPKEFGNLTKLKYLDLAEGNVGGEIPDELGKLKLLNTVFLYKNSFEGKIPTNIGNMTS 865
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
L++LDLS N LSG +P + +L ++ L+ N L G + + G+L L +++L NN L
Sbjct: 866 LVLLDLSDNMLSGNIPAEISQLKNLQLLNFMRNKLSGPVPSGLGDLPQLEVLELWNNSL 1042
Score = 76.6 bits (187), Expect = 2e-14
Identities = 46/147 (31%), Positives = 76/147 (51%)
Frame = +2
Query: 190 GNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLS 249
G L L+ +++ N F G IP FG L+ L LDL+ ++ G +P LG+L + + L
Sbjct: 635 GKLSSLEYMIIGYNEFEGGIPKEFGNLTKLKYLDLAEGNVGGEIPDELGKLKLLNTVFLY 814
Query: 250 HNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLK 309
N EGK+ GN+ +L L+DL +N L + + ++ +L+ + N L G + +
Sbjct: 815 KNSFEGKIPTNIGNMTSLVLLDLSDNMLSGNIPAEISQLKNLQLLNFMRNKLSGPVPS-G 991
Query: 310 WENLQNLVILELSNMELIGEIPESLSQ 336
+L L +LEL N L +P+ Q
Sbjct: 992 LGDLPQLEVLELWNNSLSRPLPKDPRQ 1072
Score = 69.7 bits (169), Expect = 2e-12
Identities = 42/99 (42%), Positives = 53/99 (53%)
Frame = +2
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G IP G LK L ++ L +N G IP IGN+ L L LS N SGNIP L +
Sbjct: 758 GEIPDELGKLKLLNTVFLYKNSFEGKIPTNIGNMTSLVLLDLSDNMLSGNIPAEISQLKN 937
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKL 257
L +L+ RN LSG +P LG L + L+L +N L L
Sbjct: 938 LQLLNFMRNKLSGPVPSGLGDLPQLEVLELWNNSLSRPL 1054
Score = 57.8 bits (138), Expect = 9e-09
Identities = 35/107 (32%), Positives = 55/107 (50%)
Frame = +1
Query: 171 LQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLS 230
++ L L L+G++ EI +L L L L N F ++ L+ L LD+S+N +
Sbjct: 214 VEKLNLSHMNLSGSVSNEIQSLKSLTFLNLCCNGFESSLSKHITNLTSLKSLDVSQNFFT 393
Query: 231 GTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
G P+ LG+ +L L+ S N G L + GN+ +L +DLR L
Sbjct: 394 GGFPLGLGKASELLTLNASSNNFSGFLPEDLGNISSLETLDLRGKLL 534
Score = 50.8 bits (120), Expect = 1e-06
Identities = 50/193 (25%), Positives = 81/193 (41%), Gaps = 5/193 (2%)
Frame = +2
Query: 196 KRLVLSGNNFSGNIPDIFGGLSDLLILDLSRN----SLSGTLPVTLGRLISVLKLDLSHN 251
+ L+L + F G+IP +S ++ R+ S + G+L S+ + + +N
Sbjct: 506 RHLILEASFFEGSIPK---SISKFE*SEVPRSIWE*SHREKSRLRFGKLSSLEYMIIGYN 676
Query: 252 FLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRT-LKW 310
EG + EFGNL L +DL + GG+I L
Sbjct: 677 EFEGGIPKEFGNLTKLKYLDLAEGNV------------------------GGEIPDELGK 784
Query: 311 ENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLS 370
L N V L ++ E G+IP ++ + L L LSDN ++GN+ ++ L +L L
Sbjct: 785 LKLLNTVFLYKNSFE--GKIPTNIGNMTSLVLLDLSDNMLSGNIPAEISQLKNLQLLNFM 958
Query: 371 GNNLKGEIQFSKG 383
N L G + G
Sbjct: 959 RNKLSGPVPSGLG 997
Score = 48.5 bits (114), Expect = 5e-06
Identities = 39/108 (36%), Positives = 57/108 (52%), Gaps = 1/108 (0%)
Frame = +2
Query: 104 EFKPELFQLKHLKAISFF-NCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIP 162
E EL +LK L + + N F+ K+P +I GN SL ++ N L GNIP
Sbjct: 761 EIPDELGKLKLLNTVFLYKNSFEG--KIPTNI--GNMT----SLVLLDLSDNM-LSGNIP 913
Query: 163 STFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
+ LKNLQ L + N L+G +P +G+L +L+ L L N+ S +P
Sbjct: 914 AEISQLKNLQLLNFMRNKLSGPVPSGLGDLPQLEVLELWNNSLSRPLP 1057
Score = 44.7 bits (104), Expect = 8e-05
Identities = 30/91 (32%), Positives = 47/91 (50%), Gaps = 2/91 (2%)
Frame = +1
Query: 136 TGNWEKLAESLESIEFRSN--PGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLV 193
+G+ +SL+S+ F + G ++ L +L+SL + +N TG P +G
Sbjct: 247 SGSVSNEIQSLKSLTFLNLCCNGFESSLSKHITNLTSLKSLDVSQNFFTGGFPLGLGKAS 426
Query: 194 KLKRLVLSGNNFSGNIPDIFGGLSDLLILDL 224
+L L S NNFSG +P+ G +S L LDL
Sbjct: 427 ELLTLNASSNNFSGFLPEDLGNISSLETLDL 519
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,789,005
Number of Sequences: 36976
Number of extensions: 227693
Number of successful extensions: 2654
Number of sequences better than 10.0: 277
Number of HSP's better than 10.0 without gapping: 1717
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2245
length of query: 475
length of database: 9,014,727
effective HSP length: 100
effective length of query: 375
effective length of database: 5,317,127
effective search space: 1993922625
effective search space used: 1993922625
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146570.9


