

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
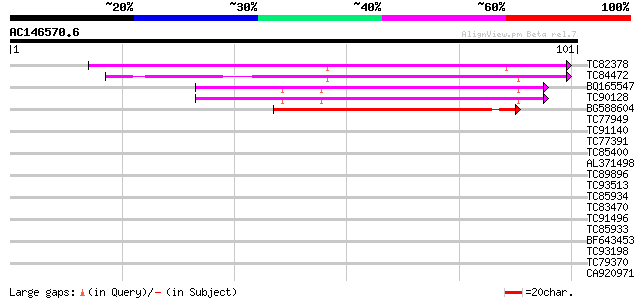
Query= AC146570.6 + phase: 0
(101 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC82378 weakly similar to GP|7770334|gb|AAF69704.1| F27J15.21 {A... 69 3e-13
TC84472 similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidop... 60 1e-10
BQ165547 similar to GP|7770334|gb|A F27J15.21 {Arabidopsis thali... 55 4e-09
TC90128 similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidop... 55 4e-09
BG588604 similar to PIR|G96739|G967 F14O23.12 [imported] - Arabi... 47 1e-06
TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {A... 29 0.33
TC91140 weakly similar to PIR|T46193|T46193 hypothetical protein... 28 0.74
TC77391 similar to GP|14719339|gb|AAK73157.1 putative protein ki... 27 0.97
TC85400 sucrose synthase 27 0.97
AL371498 similar to GP|10041548|em unnamed protein product {Arab... 27 1.3
TC89896 weakly similar to SP|P45068|FTSA_HAEIN Cell division pro... 26 2.2
TC93513 homologue to GP|20135918|emb|CAD29515. hypothetical prot... 26 2.8
TC85934 similar to GP|15292843|gb|AAK92792.1 putative CCR4-assoc... 25 4.8
TC83470 weakly similar to GP|16118437|gb|AAL12626.1 leucine-rich... 25 4.8
TC91496 similar to GP|13272465|gb|AAK17171.1 unknown protein {Ar... 25 4.8
TC85933 similar to GP|20197620|gb|AAD15397.2 putative CCR4-assoc... 25 4.8
BF643453 24 8.2
TC93198 24 8.2
TC79370 24 8.2
CA920971 similar to GP|13940608|gb| Putative cystathionine gamma... 24 8.2
>TC82378 weakly similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidopsis
thaliana}, partial (37%)
Length = 568
Score = 68.9 bits (167), Expect = 3e-13
Identities = 37/93 (39%), Positives = 56/93 (59%), Gaps = 7/93 (7%)
Frame = +2
Query: 15 ISLKTKKLEELSTSSKNIIKKIYKAIHCKILKMKKQEDDKC------LWKKTILMGEKCQ 68
+ LK + S K ++K I I KMKK++ K +W+KTILMG+KC+
Sbjct: 101 LKLKAAPMTSEVKSPKKLLKNISNKAMPFIEKMKKKKKGKLEWGDGGVWQKTILMGDKCE 280
Query: 69 PLEFPGAIFYDSEGNQLSE-PPRTPRSSPSSSF 100
PL+F G I+YD++G Q++E P R+PR++P F
Sbjct: 281 PLDFSGVIYYDNKGKQVNEFPLRSPRATPVPGF 379
>TC84472 similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidopsis
thaliana}, partial (25%)
Length = 751
Score = 60.5 bits (145), Expect = 1e-10
Identities = 38/87 (43%), Positives = 50/87 (56%), Gaps = 4/87 (4%)
Frame = +1
Query: 18 KTKKLEELSTSSKNIIKKIYKAIHCKILKMKKQEDDKC---LWKKTILMGEKCQPLEFPG 74
KTKKL LS S + + K + K KKQ D++ +W+K ILMG KC+PL+F G
Sbjct: 196 KTKKL--LSNMSNKALSQFGK-----MKKTKKQRDNELNDGVWQKEILMGGKCEPLDFSG 354
Query: 75 AIFYDSEGNQLSEPP-RTPRSSPSSSF 100
I YD G Q E P R+PR+ P S +
Sbjct: 355 VIHYDINGRQTGEVPLRSPRAIPFSRY 435
>BQ165547 similar to GP|7770334|gb|A F27J15.21 {Arabidopsis thaliana},
partial (32%)
Length = 649
Score = 55.1 bits (131), Expect = 4e-09
Identities = 32/75 (42%), Positives = 43/75 (56%), Gaps = 12/75 (16%)
Frame = -1
Query: 34 KKIYKAIHCKILKM---KKQEDDK--------CLWKKTILMGEKCQPLEFPGAIFYDSEG 82
KK+ I K L KKQ +++ +W+K ILMG KC+PL+F G I+YD G
Sbjct: 469 KKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDING 290
Query: 83 NQLSEPP-RTPRSSP 96
Q E P R+PR+SP
Sbjct: 289 KQTREVPIRSPRASP 245
>TC90128 similar to GP|7770334|gb|AAF69704.1| F27J15.21 {Arabidopsis
thaliana}, partial (32%)
Length = 716
Score = 55.1 bits (131), Expect = 4e-09
Identities = 32/75 (42%), Positives = 43/75 (56%), Gaps = 12/75 (16%)
Frame = +3
Query: 34 KKIYKAIHCKILKM---KKQEDDK--------CLWKKTILMGEKCQPLEFPGAIFYDSEG 82
KK+ I K L KKQ +++ +W+K ILMG KC+PL+F G I+YD G
Sbjct: 255 KKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDING 434
Query: 83 NQLSEPP-RTPRSSP 96
Q E P R+PR+SP
Sbjct: 435 KQTREVPIRSPRASP 479
>BG588604 similar to PIR|G96739|G967 F14O23.12 [imported] - Arabidopsis
thaliana, partial (32%)
Length = 568
Score = 47.0 bits (110), Expect = 1e-06
Identities = 21/44 (47%), Positives = 31/44 (69%)
Frame = +3
Query: 48 KKQEDDKCLWKKTILMGEKCQPLEFPGAIFYDSEGNQLSEPPRT 91
++Q ++ +W+K ILMG KCQ +F G I YDS+GN ++ P RT
Sbjct: 168 EQQREEDSIWQKNILMGGKCQLPDFSGVIIYDSDGNVVN-PART 296
>TC77949 weakly similar to GP|11762218|gb|AAG40387.1 AT4g27520 {Arabidopsis
thaliana}, partial (34%)
Length = 1297
Score = 28.9 bits (63), Expect = 0.33
Identities = 24/76 (31%), Positives = 35/76 (45%), Gaps = 13/76 (17%)
Frame = +3
Query: 37 YKAIHCKILKMKKQEDDKCLWKKTILM---GEKCQPLEFPGAIFYDSEGNQLSE------ 87
YK +L++KK++ +KC I GE L+ G ++ S +Q E
Sbjct: 294 YKKGSDSVLEVKKEDYEKCNKTNPIKKFEDGETEFTLDRAGPFYFISGKDQNCENGQKLT 473
Query: 88 ----PPRTPRSSPSSS 99
PRTP+SSPS S
Sbjct: 474 LVVISPRTPKSSPSPS 521
>TC91140 weakly similar to PIR|T46193|T46193 hypothetical protein T8H10.170
- Arabidopsis thaliana, partial (9%)
Length = 1067
Score = 27.7 bits (60), Expect = 0.74
Identities = 13/39 (33%), Positives = 25/39 (63%), Gaps = 1/39 (2%)
Frame = +3
Query: 18 KTKKLEELS-TSSKNIIKKIYKAIHCKILKMKKQEDDKC 55
K +LE+L+ T+ + + + K+ +C+IL++ K DKC
Sbjct: 879 KCSRLEQLNWTTDRQDVYRSVKSSYCQILQLYK*RPDKC 995
>TC77391 similar to GP|14719339|gb|AAK73157.1 putative protein kinase {Oryza
sativa}, partial (82%)
Length = 1753
Score = 27.3 bits (59), Expect = 0.97
Identities = 16/50 (32%), Positives = 29/50 (58%), Gaps = 2/50 (4%)
Frame = -2
Query: 17 LKTKKLEELSTSSKNIIKKIYKAIHCKILKMKKQ--EDDKCLWKKTILMG 64
++TKK EE K++ +Y + K L+ KK+ +++ C W+ I+MG
Sbjct: 213 VETKK*EENKIK*KHLFFIVY*SFDEKDLRKKKKGSKEEVC*WRLLIMMG 64
>TC85400 sucrose synthase
Length = 3133
Score = 27.3 bits (59), Expect = 0.97
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = -3
Query: 45 LKMKKQEDDKCLWKKTILMGEKCQPLEFPGAI 76
+K+++ D+K W K LMG K P + P I
Sbjct: 3026 IKLEENMDNKKAWLKIELMGAKHPPFKLPSYI 2931
>AL371498 similar to GP|10041548|em unnamed protein product {Arabidopsis
thaliana}, partial (18%)
Length = 458
Score = 26.9 bits (58), Expect = 1.3
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = +1
Query: 53 DKCLWKKTILMGEKCQ 68
D CLWKKT MG C+
Sbjct: 217 DLCLWKKTERMGSACK 264
>TC89896 weakly similar to SP|P45068|FTSA_HAEIN Cell division protein ftsA.
{Haemophilus influenzae}, partial (4%)
Length = 757
Score = 26.2 bits (56), Expect = 2.2
Identities = 16/43 (37%), Positives = 25/43 (57%)
Frame = -3
Query: 6 INKAIISYIISLKTKKLEELSTSSKNIIKKIYKAIHCKILKMK 48
+ K IISY+I L+ K L+ L S N ++ A+ KI K++
Sbjct: 146 LKKIIISYLIGLRRKILKTLLRSQIN*LQLRKIAVTRKIKKLE 18
>TC93513 homologue to GP|20135918|emb|CAD29515. hypothetical protein
{Trifolium repens}, partial (16%)
Length = 601
Score = 25.8 bits (55), Expect = 2.8
Identities = 21/55 (38%), Positives = 26/55 (47%)
Frame = -2
Query: 2 IINKINKAIISYIISLKTKKLEELSTSSKNIIKKIYKAIHCKILKMKKQEDDKCL 56
IIN I ++I L L L +S+K I +A H KILK DKCL
Sbjct: 330 IINVIGWCK*QFLIILTENLLTFLGSSTKCI----NEAAHNKILKTAMDGIDKCL 178
>TC85934 similar to GP|15292843|gb|AAK92792.1 putative CCR4-associated
factor {Arabidopsis thaliana}, partial (50%)
Length = 721
Score = 25.0 bits (53), Expect = 4.8
Identities = 7/27 (25%), Positives = 18/27 (65%)
Frame = +1
Query: 35 KIYKAIHCKILKMKKQEDDKCLWKKTI 61
+IY+ I+ K+++ K + +WK+++
Sbjct: 274 RIYRKIYRKVIRFKSERYGTIIWKRSL 354
>TC83470 weakly similar to GP|16118437|gb|AAL12626.1 leucine-rich repeat
receptor-like kinase F21M12.36 {Arabidopsis thaliana},
partial (8%)
Length = 632
Score = 25.0 bits (53), Expect = 4.8
Identities = 13/40 (32%), Positives = 24/40 (59%)
Frame = +2
Query: 7 NKAIISYIISLKTKKLEELSTSSKNIIKKIYKAIHCKILK 46
NK I+S++ KT+ E+ + + I ++YK CK+L+
Sbjct: 14 NKDIVSWVHG-KTRSKEKFMSVVDSRIPEMYKEEACKVLR 130
>TC91496 similar to GP|13272465|gb|AAK17171.1 unknown protein {Arabidopsis
thaliana}, partial (31%)
Length = 672
Score = 25.0 bits (53), Expect = 4.8
Identities = 23/72 (31%), Positives = 32/72 (43%)
Frame = -3
Query: 26 STSSKNIIKKIYKAIHCKILKMKKQEDDKCLWKKTILMGEKCQPLEFPGAIFYDSEGNQL 85
S ++K+I+ KI C I K DD KKT M E + GAI +
Sbjct: 394 SDATKHIMAKIKAVSTCSIPNSKYCNDDIVEEKKT--MKEHVAAVT-SGAIPT*RRSGLI 224
Query: 86 SEPPRTPRSSPS 97
S PP P++ P+
Sbjct: 223 SIPPPIPKTPPN 188
>TC85933 similar to GP|20197620|gb|AAD15397.2 putative CCR4-associated
factor {Arabidopsis thaliana}, partial (94%)
Length = 1362
Score = 25.0 bits (53), Expect = 4.8
Identities = 7/27 (25%), Positives = 18/27 (65%)
Frame = +1
Query: 35 KIYKAIHCKILKMKKQEDDKCLWKKTI 61
+IY+ I+ K+++ K + +WK+++
Sbjct: 295 RIYRRIYRKVIRFKSERYGTIIWKRSL 375
>BF643453
Length = 478
Score = 24.3 bits (51), Expect = 8.2
Identities = 17/60 (28%), Positives = 29/60 (48%), Gaps = 6/60 (10%)
Frame = -3
Query: 18 KTKKLEELSTSSKNIIKKIYKAIHCKILKMKKQEDD------KCLWKKTILMGEKCQPLE 71
K K L + + N K+ + + LK+K + + K WKK++L GEK + L+
Sbjct: 209 KYKTLNQQNYQRLN*TSKLTETLILTNLKIKTR*QNN*TIERKFEWKKSVLGGEKIEELD 30
>TC93198
Length = 676
Score = 24.3 bits (51), Expect = 8.2
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = -3
Query: 35 KIYKAIHCKILKMKKQEDDKCLWKKTIL 62
K YK I ++L K +KCL+ KTIL
Sbjct: 440 KKYKGIC*RVLHRIKDGLNKCLYVKTIL 357
>TC79370
Length = 1557
Score = 24.3 bits (51), Expect = 8.2
Identities = 13/44 (29%), Positives = 20/44 (44%)
Frame = +1
Query: 16 SLKTKKLEELSTSSKNIIKKIYKAIHCKILKMKKQEDDKCLWKK 59
SLK L +L ++Y A C+I E+ KC W++
Sbjct: 1024 SLKCVGLGDLIYVFNEDYHRMYPACVCEIRSRGGGENSKCYWRR 1155
>CA920971 similar to GP|13940608|gb| Putative cystathionine gamma synthase
(O-succinylhomoserine (thiol)-lyase) {Oryza sativa},
partial (3%)
Length = 730
Score = 24.3 bits (51), Expect = 8.2
Identities = 9/14 (64%), Positives = 12/14 (85%)
Frame = +1
Query: 86 SEPPRTPRSSPSSS 99
S PPR+P SSP++S
Sbjct: 628 STPPRSPNSSPTTS 669
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.132 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,271,692
Number of Sequences: 36976
Number of extensions: 41631
Number of successful extensions: 213
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 211
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 213
length of query: 101
length of database: 9,014,727
effective HSP length: 77
effective length of query: 24
effective length of database: 6,167,575
effective search space: 148021800
effective search space used: 148021800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 50 (23.9 bits)
Medicago: description of AC146570.6


