

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
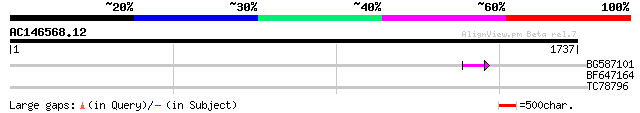
Query= AC146568.12 + phase: 0 /pseudo
(1737 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 62 2e-09
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 41 0.004
TC78796 similar to GP|15810563|gb|AAL07169.1 putative calreticul... 30 6.2
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 62.0 bits (149), Expect = 2e-09
Identities = 37/83 (44%), Positives = 48/83 (57%)
Frame = +3
Query: 1387 EPLRKSCKAVSGGLLFSRIAMIL*GGVTIAKEQVVFLRGMRCP*RESLKLNHLTVGALIL 1446
+P R+S K VSGGL S++ V A+++ + RGMRC S KL +LT G I
Sbjct: 156 KPFRRSNKLVSGGLRCSKMPTPSSLSVIHARDREISARGMRCHRTSSWKLRYLTCGGSIS 335
Query: 1447 WDLSLPLPLISIYLFVLTTSPNG 1469
WD S PL IS L +LTT P+G
Sbjct: 336 WDHSHPLTTISTSLLLLTTFPSG 404
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 41.2 bits (95), Expect = 0.004
Identities = 32/72 (44%), Positives = 39/72 (53%)
Frame = +2
Query: 1603 LIGPSKLLILIKL*LVRKDS*SSMS*KK*DLVRMRTPSFTKRELRSIMTRVW*GESFMLD 1662
L+G K L I K *S MS*+ *D MR P FTK+E ++ M V GES
Sbjct: 41 LLGLLKPLTWITKPPEEKGY*S*MS*RS*D*TPMRMPRFTKKEQKNGMIGV*SGESLERG 220
Query: 1663 NLFCCLIPG*NY 1674
+L+ L PG*NY
Sbjct: 221 SLYYYLTPG*NY 256
>TC78796 similar to GP|15810563|gb|AAL07169.1 putative calreticulin protein
{Arabidopsis thaliana}, partial (89%)
Length = 1605
Score = 30.4 bits (67), Expect = 6.2
Identities = 20/42 (47%), Positives = 26/42 (61%)
Frame = +2
Query: 396 KLMKMLWTRMKMLKMRK*LKKRVIKV*LKIKRRKKQREKKVR 437
KL K +W + LK RK LKKR K L KRR+K +EK+ +
Sbjct: 1106 KLWKNIWLIIGRLK-RKLLKKRKKKEGLGKKRRQKGQEKRAK 1228
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.369 0.165 0.601
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,841,917
Number of Sequences: 36976
Number of extensions: 770502
Number of successful extensions: 9274
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1973
Number of HSP's successfully gapped in prelim test: 283
Number of HSP's that attempted gapping in prelim test: 7139
Number of HSP's gapped (non-prelim): 2600
length of query: 1737
length of database: 9,014,727
effective HSP length: 110
effective length of query: 1627
effective length of database: 4,947,367
effective search space: 8049366109
effective search space used: 8049366109
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146568.12


