

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.9 + phase: 0
(80 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
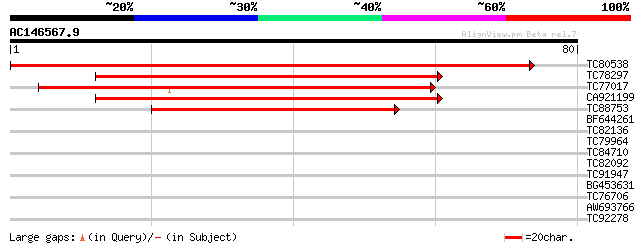
Score E
Sequences producing significant alignments: (bits) Value
TC80538 homologue to PIR|D96797|D96797 Sm-like protein [imported... 142 2e-35
TC78297 homologue to GP|9758886|dbj|BAB09440.1 U6 snRNA-associat... 40 1e-04
TC77017 homologue to PIR|T00420|T00420 probable small nuclear ri... 40 1e-04
CA921199 similar to PIR|T51950|T51 DNA-directed RNA polymerase (... 40 1e-04
TC88753 similar to PIR|T10586|T10586 small nuclear ribonucleopro... 38 6e-04
BF644261 homologue to GP|20196966|gb| putative small nuclear rib... 37 0.001
TC82136 similar to GP|22202729|dbj|BAC07386. putative snRNP prot... 37 0.001
TC79964 homologue to PIR|B84453|B84453 probable snRNP splicing f... 34 0.012
TC84710 homologue to PIR|B84453|B84453 probable snRNP splicing f... 33 0.026
TC82092 similar to PIR|H86164|H86164 hypothetical protein AAC721... 31 0.076
TC91947 similar to GP|8096410|dbj|BAA95880.1 hypothetical protei... 28 0.84
BG453631 homologue to GP|21106839|gb| disulfide oxidoreductase {... 25 5.4
TC76706 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulat... 25 5.4
AW693766 similar to GP|16611746|gb| putative glycosyl transferas... 24 9.3
TC92278 similar to SP|O43093|KINH_SYNRA Kinesin heavy chain (Syn... 24 9.3
>TC80538 homologue to PIR|D96797|D96797 Sm-like protein [imported] -
Arabidopsis thaliana, partial (95%)
Length = 554
Score = 142 bits (359), Expect = 2e-35
Identities = 72/74 (97%), Positives = 73/74 (98%)
Frame = +2
Query: 1 MAGTEEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV 60
MAGTEEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV
Sbjct: 41 MAGTEEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV 220
Query: 61 EIDDETYEEIVRVS 74
EIDDETYEEIVR +
Sbjct: 221 EIDDETYEEIVRTT 262
>TC78297 homologue to GP|9758886|dbj|BAB09440.1 U6 snRNA-associated Sm-like
protein-like {Arabidopsis thaliana}, partial (95%)
Length = 693
Score = 40.4 bits (93), Expect = 1e-04
Identities = 19/49 (38%), Positives = 30/49 (60%)
Frame = +1
Query: 13 PLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVE 61
P +LI + +I+V ++ D+EL G L +D ++NM+L DV E T E
Sbjct: 193 PSELIDRCIGSKIWVIMKGDKELVGTLRGFDVYVNMVLEDVTEYEITAE 339
>TC77017 homologue to PIR|T00420|T00420 probable small nuclear
ribonucleoprotein D2 [imported] - Arabidopsis thaliana,
complete
Length = 1186
Score = 40.4 bits (93), Expect = 1e-04
Identities = 19/58 (32%), Positives = 38/58 (64%), Gaps = 2/58 (3%)
Frame = +3
Query: 5 EEESAVKEPLDLIRLSL--DERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV 60
EEE PL ++ +S+ + ++ + R++++L G++ A+D+H NM+L +V E+ T V
Sbjct: 624 EEEEFNTGPLSVLMMSVKNNTQVLINCRNNKKLLGRVRAFDRHCNMVLENVREMWTEV 797
>CA921199 similar to PIR|T51950|T51 DNA-directed RNA polymerase (EC 2.7.7.6)
23K chain [imported] - Arabidopsis thaliana, partial
(49%)
Length = 756
Score = 40.4 bits (93), Expect = 1e-04
Identities = 19/49 (38%), Positives = 30/49 (60%)
Frame = +2
Query: 13 PLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVE 61
P +LI + +I+V ++ D+EL G L +D ++NM+L DV E T E
Sbjct: 605 PSELIDRCIGSKIWVIMKGDKELVGTLRGFDVYVNMVLEDVTEYEITAE 751
>TC88753 similar to PIR|T10586|T10586 small nuclear
ribonucleoprotein-associated protein homolog F9F13.90 -
Arabidopsis thaliana, partial (77%)
Length = 1273
Score = 38.1 bits (87), Expect = 6e-04
Identities = 15/35 (42%), Positives = 26/35 (73%)
Frame = +2
Query: 21 LDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEE 55
++ R+ V ++ R+L GK A+D+H+N++LGD EE
Sbjct: 158 INYRMRVTIQDGRQLVGKFMAFDRHMNLVLGDCEE 262
>BF644261 homologue to GP|20196966|gb| putative small nuclear
ribonucleoprotein D2 {Arabidopsis thaliana}, partial
(86%)
Length = 668
Score = 37.4 bits (85), Expect = 0.001
Identities = 18/58 (31%), Positives = 38/58 (65%), Gaps = 2/58 (3%)
Frame = +1
Query: 5 EEESAVKEPLDLIRLSL--DERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV 60
+EE PL ++ +S+ + ++ + R++++L G++ A+D+H NM+L +V E+ T V
Sbjct: 145 QEEEFNIGPLSVLYMSVRNNTQVLINCRNNKKLLGRVMAFDRHCNMVLENVREMWTEV 318
>TC82136 similar to GP|22202729|dbj|BAC07386. putative snRNP protein
{Oryza sativa (japonica cultivar-group)}, partial (51%)
Length = 629
Score = 37.4 bits (85), Expect = 0.001
Identities = 15/35 (42%), Positives = 26/35 (73%)
Frame = +2
Query: 21 LDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEE 55
++ R+ V ++ R+L GK A+D+H+N++LGD EE
Sbjct: 59 INYRMRVTIQDGRQLVGKFMAFDRHMNLVLGDWEE 163
>TC79964 homologue to PIR|B84453|B84453 probable snRNP splicing factor
[imported] - Arabidopsis thaliana, partial (93%)
Length = 607
Score = 33.9 bits (76), Expect = 0.012
Identities = 17/44 (38%), Positives = 27/44 (60%)
Frame = +1
Query: 14 LDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIV 57
LDL + +D+ + VKL R++ G L YDQ LN++L + E +
Sbjct: 88 LDLAKF-VDKGVQVKLTGGRQVTGTLKGYDQLLNLVLDEAVEFL 216
>TC84710 homologue to PIR|B84453|B84453 probable snRNP splicing factor
[imported] - Arabidopsis thaliana, partial (93%)
Length = 543
Score = 32.7 bits (73), Expect = 0.026
Identities = 17/44 (38%), Positives = 26/44 (58%)
Frame = +3
Query: 14 LDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIV 57
LDL + +D+ + VKL R++ G L YDQ LN+ L + E +
Sbjct: 42 LDLAKF-VDKGVQVKLTGGRQVTGTLKGYDQLLNLFLDEAVEFL 170
>TC82092 similar to PIR|H86164|H86164 hypothetical protein AAC72111.1
[imported] - Arabidopsis thaliana, partial (80%)
Length = 311
Score = 31.2 bits (69), Expect = 0.076
Identities = 15/49 (30%), Positives = 29/49 (58%)
Frame = +1
Query: 25 IYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEEIVRV 73
+ V+L++D +RG LH+ DQ+LN+ ++ T +D + Y ++ V
Sbjct: 70 VTVELKNDLAIRGTLHSVDQYLNI------KLENTRVVDQDRYPHMLSV 198
>TC91947 similar to GP|8096410|dbj|BAA95880.1 hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 872
Score = 27.7 bits (60), Expect = 0.84
Identities = 9/18 (50%), Positives = 16/18 (88%)
Frame = +2
Query: 32 DRELRGKLHAYDQHLNMI 49
+R+LRG +HA++ HLN++
Sbjct: 515 ERDLRGSVHAHNTHLNVL 568
>BG453631 homologue to GP|21106839|gb| disulfide oxidoreductase {Xanthomonas
axonopodis pv. citri str. 306}, partial (5%)
Length = 661
Score = 25.0 bits (53), Expect = 5.4
Identities = 16/51 (31%), Positives = 27/51 (52%), Gaps = 6/51 (11%)
Frame = +1
Query: 4 TEEESAVKEPL----DLIRLSLDERIYVK--LRSDRELRGKLHAYDQHLNM 48
++EE +++ L D + SL + + VK L R G++ AYD H N+
Sbjct: 301 SDEEDVMEDNLSDEQDFMEDSLADNLKVKIFLYDGRSYEGQVRAYDFHFNI 453
>TC76706 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulated protein
CORA. [Alfalfa] {Medicago sativa}, partial (94%)
Length = 1034
Score = 25.0 bits (53), Expect = 5.4
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +2
Query: 53 VEEIVTTVEIDDETYEEIVRVSHNPN 78
VE + TTVE+D E E+V V P+
Sbjct: 563 VEVVTTTVEVDTEDTVEVVTVVTVPS 640
>AW693766 similar to GP|16611746|gb| putative glycosyl transferase
{Shigella boydii}, partial (5%)
Length = 656
Score = 24.3 bits (51), Expect = 9.3
Identities = 11/30 (36%), Positives = 18/30 (59%)
Frame = +1
Query: 43 DQHLNMILGDVEEIVTTVEIDDETYEEIVR 72
++H + ILG VEE+ + E YEE ++
Sbjct: 34 NEHEHGILGFVEEVKSVFHGKREEYEEFIK 123
>TC92278 similar to SP|O43093|KINH_SYNRA Kinesin heavy chain (Synkin).
{Syncephalastrum racemosum}, partial (10%)
Length = 709
Score = 24.3 bits (51), Expect = 9.3
Identities = 16/60 (26%), Positives = 33/60 (54%)
Frame = +3
Query: 12 EPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEEIV 71
E L +R L E + + ++ + +LH ++HL ++E +TT+E++ YEE++
Sbjct: 135 EELSTLRRELSESKSLVTQHEQTIN-ELHYENEHLTRKRDELEIRLTTLELE---YEELL 302
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.134 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,923,802
Number of Sequences: 36976
Number of extensions: 17933
Number of successful extensions: 70
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 70
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 70
length of query: 80
length of database: 9,014,727
effective HSP length: 56
effective length of query: 24
effective length of database: 6,944,071
effective search space: 166657704
effective search space used: 166657704
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC146567.9


