

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
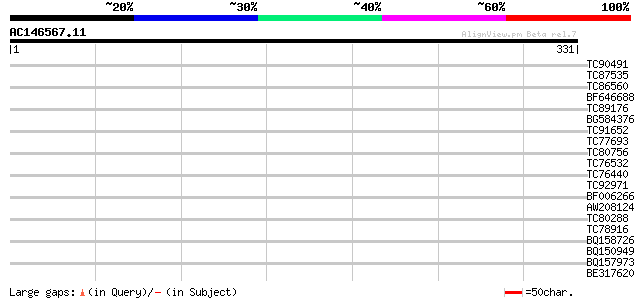
Query= AC146567.11 - phase: 0
(331 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC90491 homologue to PIR|S33706|S33706 DNA-damage repair protein... 35 0.052
TC87535 similar to GP|19310542|gb|AAL85004.1 unknown protein {Ar... 33 0.15
TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-... 32 0.26
BF646688 similar to GP|3851530|gb| nodulin {Glycine max}, partia... 31 0.58
TC89176 homologue to GP|13539605|emb|CAC35733. cyclophilin-RNA i... 31 0.58
BG584376 30 1.7
TC91652 similar to GP|11935197|gb|AAG42014.1 unknown protein {Ar... 29 2.9
TC77693 weakly similar to GP|7108597|gb|AAF36490.1| K+ transport... 29 2.9
TC80756 homologue to PIR|S53498|S53498 dnaK-type molecular chape... 29 2.9
TC76532 weakly similar to SP|Q9BBN6|YCF1_LOTJA Hypothetical 214.... 29 2.9
TC76440 weakly similar to SP|Q9LD55|IF3A_ARATH Eukaryotic transl... 28 3.7
TC92971 similar to GP|9757935|dbj|BAB08478.1 gene_id:MSF19.7~pir... 28 3.7
BF006266 similar to GP|21491|emb|C ST-LS1 protein (AA 1-138) {So... 28 4.9
AW208124 weakly similar to PIR|T49082|T490 hypothetical protein ... 28 6.4
TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein... 28 6.4
TC78916 similar to GP|7267777|emb|CAB81180.1 predicted protein o... 28 6.4
BQ158726 similar to GP|21491|emb|C ST-LS1 protein (AA 1-138) {So... 27 8.3
BQ150949 similar to GP|21491|emb|C ST-LS1 protein (AA 1-138) {So... 27 8.3
BQ157973 similar to GP|21491|emb|C ST-LS1 protein (AA 1-138) {So... 27 8.3
BE317620 similar to GP|21491|emb|CA ST-LS1 protein (AA 1-138) {S... 27 8.3
>TC90491 homologue to PIR|S33706|S33706 DNA-damage repair protein DRT111
precursor - Arabidopsis thaliana, partial (29%)
Length = 630
Score = 34.7 bits (78), Expect = 0.052
Identities = 18/60 (30%), Positives = 36/60 (60%), Gaps = 4/60 (6%)
Frame = +3
Query: 143 QIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIP--ESIRHNFEQME--EERTKVLIAIE 198
++D++++D + +C +Y ++ +T+PN P E++R F Q E EE TK L+ ++
Sbjct: 66 EVDDELEDEVGSECAKYGTVTRVLIFEITEPNFPTDEAVR-IFVQFERSEETTKALVDLD 242
>TC87535 similar to GP|19310542|gb|AAL85004.1 unknown protein {Arabidopsis
thaliana}, partial (86%)
Length = 893
Score = 33.1 bits (74), Expect = 0.15
Identities = 25/84 (29%), Positives = 41/84 (48%), Gaps = 3/84 (3%)
Frame = +1
Query: 181 HNFEQMEEERTK--VLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDS 238
HN + +E++ K + +KQ EKE + KK EKN+ SK K+SE+ S
Sbjct: 1 HNTNRTKEKKRKENTSSSFQKQSKREKERQK*KKKETKNKEKNSKKSKKN*TTKMSEQQS 180
Query: 239 -ARRQEEIENAMYLAREKSLADAD 261
+ Q ++N+ L+ L+ D
Sbjct: 181 NSNNQLVVQNSGSLSFSSHLSKED 252
>TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-like
{Arabidopsis thaliana}, partial (28%)
Length = 2450
Score = 32.3 bits (72), Expect = 0.26
Identities = 24/67 (35%), Positives = 33/67 (48%), Gaps = 2/67 (2%)
Frame = +2
Query: 177 ESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAE--KNANVSKILMEQKLS 234
E R E M+ + ++ E K++ +EAE K A EAE K A S I L+
Sbjct: 1499 EEARREVEDMKNKTDELKKEAEATKLALEEAEKKLKEATEEAEAAKAAEASAIEQITVLT 1678
Query: 235 EKDSARR 241
E+ SARR
Sbjct: 1679 ERTSARR 1699
>BF646688 similar to GP|3851530|gb| nodulin {Glycine max}, partial (46%)
Length = 669
Score = 31.2 bits (69), Expect = 0.58
Identities = 27/96 (28%), Positives = 50/96 (51%), Gaps = 9/96 (9%)
Frame = +3
Query: 185 QMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSK--ILMEQKLSEK--DSAR 240
Q+ E K +A+ + ++ + E E M + +E K +SK + E K+ E + +
Sbjct: 189 QVAEVEAKKAVALREAEL-QGEVEKMNALTTTEKLKADLLSKASVQYETKVQEANWELYK 365
Query: 241 RQEEIENAMYLAR-----EKSLADADFYRVIKEAEA 271
+Q+E E ++ + +K+LAD+ FY +EAEA
Sbjct: 366 KQKEAEAILFEKKAEAEAQKALADSTFYARKQEAEA 473
>TC89176 homologue to GP|13539605|emb|CAC35733. cyclophilin-RNA interacting
protein {Paramecium tetraurelia}, partial (3%)
Length = 783
Score = 31.2 bits (69), Expect = 0.58
Identities = 19/60 (31%), Positives = 32/60 (52%)
Frame = +2
Query: 184 EQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQE 243
E+ EER K + +Q+VSE EA + E E+ +++ E++L E+ + RR E
Sbjct: 77 EKEREEREKEREKVRRQRVSEWEAIDKEN---EEKERKRREEELIRERELEERMNCRRLE 247
>BG584376
Length = 741
Score = 29.6 bits (65), Expect = 1.7
Identities = 30/111 (27%), Positives = 48/111 (43%), Gaps = 5/111 (4%)
Frame = -3
Query: 143 QIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHNFEQMEEERTKVLIAIEKQKV 202
QIDEK ++ QV ++ + TK E + E++E + IEK
Sbjct: 739 QIDEKPQNGQQVLESKPT------SIERTKAQTNEKAQKGEEEVEPK------TIEKLVT 596
Query: 203 SEKEAETMKKMAISEAEKNANVSKILMEQK-----LSEKDSARRQEEIENA 248
E E + M+ +AEK N K +E+K S K+ ++E+E A
Sbjct: 595 KENLEEKIVGMSAEDAEKERNSKKEEVEEKNEKTYESSKNQNLEEKEVEKA 443
>TC91652 similar to GP|11935197|gb|AAG42014.1 unknown protein {Arabidopsis
thaliana}, partial (78%)
Length = 887
Score = 28.9 bits (63), Expect = 2.9
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = -3
Query: 35 WRGGALLKTITEPGFHMKMPFLTQFEPVQVTLQTDEVT 72
+ GG T EP H+ + FLTQ PVQ + T +T
Sbjct: 603 YNGGF*GNTFREPSLHLPV-FLTQGTPVQAAVNTTRIT 493
>TC77693 weakly similar to GP|7108597|gb|AAF36490.1| K+ transporter HAK5
{Arabidopsis thaliana}, partial (67%)
Length = 2635
Score = 28.9 bits (63), Expect = 2.9
Identities = 13/53 (24%), Positives = 28/53 (52%)
Frame = +3
Query: 59 FEPVQVTLQTDEVTDIPCGTKGGVMIVFGKIEVVNRLHKESVYETLLNYGVQY 111
+EP+ + +E+ I T+ GV+ + G+ EVV + + + ++NY +
Sbjct: 2229 YEPIIIQGPEEEIKFIDKATENGVVYMIGEAEVVAHPNSSILNKIVVNYAYSF 2387
>TC80756 homologue to PIR|S53498|S53498 dnaK-type molecular chaperone
HSP71.2 - garden pea, partial (53%)
Length = 1189
Score = 28.9 bits (63), Expect = 2.9
Identities = 39/187 (20%), Positives = 80/187 (41%), Gaps = 13/187 (6%)
Frame = +3
Query: 145 DEKMKDALQVDCTRYAPGIEIIGVRVT-----KPNIPESIRHNFEQMEEERTKVLIAI-E 198
DEK++D L +D T + G+E G +T IP F + + VLI + E
Sbjct: 267 DEKVQDLLLLDVTPLSLGLETAGGVMTTLIPRNTTIPTKKEQIFSTYSDNQPGVLIQVFE 446
Query: 199 KQKVSEKEAETMKKMAIS------EAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLA 252
++ K+ + K ++ NV + + + + ++N + +
Sbjct: 447 GERARTKDNNLLGKFELTGIPPAPRGVPQINVCFDIDANGILNVSAEDKTAGVKNKITIT 626
Query: 253 REKS-LADADFYRVIKEAEANRLKLTPEFLELKFIESIANNTKIFFGDKIPNMILDQRLL 311
+K L+ + +++K+AE + K E ++ K +E A N+ + + N I D+++
Sbjct: 627 NDKGRLSKEEIEKMVKDAE--KYKAEDEEVKKK-VE--AKNSIENYAYNMRNTIKDEKIG 791
Query: 312 GNFLVEE 318
G E+
Sbjct: 792 GKLSHED 812
>TC76532 weakly similar to SP|Q9BBN6|YCF1_LOTJA Hypothetical 214.8 kDa protein
ycf1. {Lotus japonicus}, partial (21%)
Length = 4509
Score = 28.9 bits (63), Expect = 2.9
Identities = 31/98 (31%), Positives = 40/98 (40%), Gaps = 3/98 (3%)
Frame = +1
Query: 141 FDQIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHNFEQMEEERTKVLIAIEKQ 200
F IDE + ++ IE+I V P ESI E+R K L K
Sbjct: 1930 FKIIDELSESKKTSTISKNNSKIEVIEVIEESPVKMESINWTNSSFTEKRIKDLNV--KT 2103
Query: 201 KVSEKEAETM---KKMAISEAEKNANVSKILMEQKLSE 235
K K+ ETM KK I +E N N +K + K E
Sbjct: 2104 KTIIKQIETMTEEKKEGILTSEINLNSNKTTYDAKRLE 2217
>TC76440 weakly similar to SP|Q9LD55|IF3A_ARATH Eukaryotic translation
initiation factor 3 subunit 10 (eIF-3 theta), partial
(41%)
Length = 1953
Score = 28.5 bits (62), Expect = 3.7
Identities = 48/233 (20%), Positives = 91/233 (38%)
Frame = +1
Query: 43 TITEPGFHMKMPFLTQFEPVQVTLQTDEVTDIPCGTKGGVMIVFGKIEVVNRLHKESVYE 102
++ E F +P L + +++ Q V MI F VV ++ ++V +
Sbjct: 16 SVPEVQFSQYVPALEKLATLRLLQQVSNVYQSMKIENLAGMIPFFDFSVVEKISVDAVKQ 195
Query: 103 TLLNYGVQYDKTWIYDKIHHEINQFCSSHSLQQVYIDVFDQIDEKMKDALQVDCTRYAPG 162
L+ V + K + FC + D E++ A Q+ C P
Sbjct: 196 KFLSMKVDHMKNVVI---------FCKTSLEADGLRDHLASFAEQLNKARQMICP---PD 339
Query: 163 IEIIGVRVTKPNIPESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNA 222
+ + P + E + +++ ++ IEK+K E+E + ++K E E+ +
Sbjct: 340 RKQSKLGALLPTLSEVVAKEHKRLLARKS----IIEKRK-EEQERQLLEK----EREEES 492
Query: 223 NVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLK 275
++L + +E+ + E L REK D + EAEA RL+
Sbjct: 493 KRLRLLKIDEEAEQRRLATEIEQRKIQRLQREKEERDRE------EAEALRLE 633
>TC92971 similar to GP|9757935|dbj|BAB08478.1
gene_id:MSF19.7~pir||T01615~similar to unknown protein
{Arabidopsis thaliana}, partial (67%)
Length = 696
Score = 28.5 bits (62), Expect = 3.7
Identities = 27/119 (22%), Positives = 52/119 (43%), Gaps = 14/119 (11%)
Frame = +2
Query: 128 CSSHSLQQVYIDV-FDQIDEKMKDALQVDCTRYAPGIE------------IIGVRVTKPN 174
C LQQ DV F + + + L D +RY +E I + T+
Sbjct: 272 CIYFLLQQRQRDVEFRESANEQRQRLLSDISRYEAKVERLDGQLQAKDREIATITRTEAK 451
Query: 175 IPESIRHNFEQMEEERTKVL-IAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQK 232
+++ E++++ER + + I Q+V ++ MKK + ++++LME+K
Sbjct: 452 NTAALKAQIEKLQKERDEFQRMVIGNQQVRTQQIHEMKKKEKEYIKLQERLNQVLMEKK 628
>BF006266 similar to GP|21491|emb|C ST-LS1 protein (AA 1-138) {Solanum
tuberosum}, partial (75%)
Length = 472
Score = 28.1 bits (61), Expect = 4.9
Identities = 18/60 (30%), Positives = 28/60 (46%)
Frame = +2
Query: 53 MPFLTQFEPVQVTLQTDEVTDIPCGTKGGVMIVFGKIEVVNRLHKESVYETLLNYGVQYD 112
+P L++ V + T TD P GT GG+ + GK + + VY+ + YG D
Sbjct: 53 LPALSRRFKVVASGVTKIKTDTPYGTGGGMALPDGKDASGRKQKGKGVYQFVDKYGANVD 232
>AW208124 weakly similar to PIR|T49082|T490 hypothetical protein F4F15.140 -
Arabidopsis thaliana, partial (19%)
Length = 451
Score = 27.7 bits (60), Expect = 6.4
Identities = 12/37 (32%), Positives = 21/37 (56%)
Frame = +1
Query: 96 HKESVYETLLNYGVQYDKTWIYDKIHHEINQFCSSHS 132
HK+ V++TLL+ V+ W+Y + HE + H+
Sbjct: 325 HKDLVFKTLLSAFVRRISCWVYILLGHEDTKTVVEHT 435
>TC80288 weakly similar to PIR|T06029|T06029 hypothetical protein T28I19.100
- Arabidopsis thaliana, partial (14%)
Length = 1460
Score = 27.7 bits (60), Expect = 6.4
Identities = 17/43 (39%), Positives = 24/43 (55%)
Frame = +1
Query: 198 EKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSAR 240
EK++ S K + +K M + + EKN KI MEQK K A+
Sbjct: 478 EKKETSTKLKKKVKTMCM-KGEKNRMKKKISMEQKCRRKMKAK 603
>TC78916 similar to GP|7267777|emb|CAB81180.1 predicted protein of unknown
function {Arabidopsis thaliana}, partial (79%)
Length = 1941
Score = 27.7 bits (60), Expect = 6.4
Identities = 15/51 (29%), Positives = 28/51 (54%)
Frame = +1
Query: 184 EQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLS 234
++ EEE A+E ++ ++E E ++ A AEK A ++K+ E+ S
Sbjct: 1270 QRREEEERLAREAVEAERKLKEEEEARERAAQEAAEKQAALAKLREEKAQS 1422
>BQ158726 similar to GP|21491|emb|C ST-LS1 protein (AA 1-138) {Solanum
tuberosum}, partial (77%)
Length = 689
Score = 27.3 bits (59), Expect = 8.3
Identities = 14/41 (34%), Positives = 20/41 (48%)
Frame = +1
Query: 72 TDIPCGTKGGVMIVFGKIEVVNRLHKESVYETLLNYGVQYD 112
TD P GT GG+ + GK + + VY+ + YG D
Sbjct: 142 TDTPYGTGGGMALPDGKDASGRKQKGKGVYQFVDKYGANVD 264
>BQ150949 similar to GP|21491|emb|C ST-LS1 protein (AA 1-138) {Solanum
tuberosum}, partial (77%)
Length = 689
Score = 27.3 bits (59), Expect = 8.3
Identities = 14/41 (34%), Positives = 20/41 (48%)
Frame = +3
Query: 72 TDIPCGTKGGVMIVFGKIEVVNRLHKESVYETLLNYGVQYD 112
TD P GT GG+ + GK + + VY+ + YG D
Sbjct: 111 TDTPYGTGGGMALPDGKDASGRKQKGKGVYQFVDKYGANVD 233
>BQ157973 similar to GP|21491|emb|C ST-LS1 protein (AA 1-138) {Solanum
tuberosum}, partial (77%)
Length = 704
Score = 27.3 bits (59), Expect = 8.3
Identities = 14/41 (34%), Positives = 20/41 (48%)
Frame = +3
Query: 72 TDIPCGTKGGVMIVFGKIEVVNRLHKESVYETLLNYGVQYD 112
TD P GT GG+ + GK + + VY+ + YG D
Sbjct: 111 TDTPYGTGGGMALPDGKDASGRKQKGKGVYQFVDKYGANVD 233
>BE317620 similar to GP|21491|emb|CA ST-LS1 protein (AA 1-138) {Solanum
tuberosum}, partial (51%)
Length = 283
Score = 27.3 bits (59), Expect = 8.3
Identities = 14/41 (34%), Positives = 20/41 (48%)
Frame = +3
Query: 72 TDIPCGTKGGVMIVFGKIEVVNRLHKESVYETLLNYGVQYD 112
TD P GT GG+ + GK + + VY+ + YG D
Sbjct: 114 TDTPYGTGGGMALPDGKDASGRKQKGKGVYQFVDKYGANVD 236
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,224,151
Number of Sequences: 36976
Number of extensions: 97351
Number of successful extensions: 512
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 511
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 512
length of query: 331
length of database: 9,014,727
effective HSP length: 97
effective length of query: 234
effective length of database: 5,428,055
effective search space: 1270164870
effective search space used: 1270164870
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146567.11


