

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.10 + phase: 0
(393 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
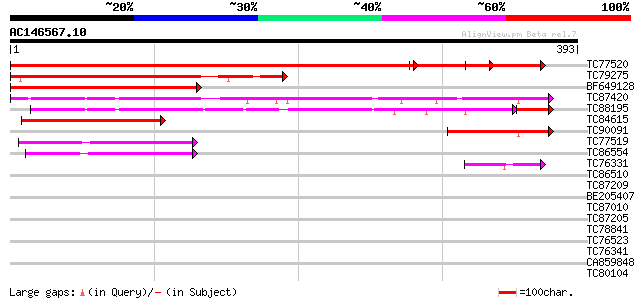
Score E
Sequences producing significant alignments: (bits) Value
TC77520 weakly similar to GP|17063173|gb|AAL32982.1 AT5g60980/MS... 555 0.0
TC79275 similar to GP|17063173|gb|AAL32982.1 AT5g60980/MSL3_100 ... 189 1e-48
BF649128 weakly similar to GP|17979475|gb AT3g25150/MJL12_9 {Ara... 162 3e-40
TC87420 weakly similar to GP|19347889|gb|AAL86001.1 unknown prot... 147 9e-36
TC88195 similar to GP|19347889|gb|AAL86001.1 unknown protein {Ar... 129 7e-33
TC84615 weakly similar to GP|17063173|gb|AAL32982.1 AT5g60980/MS... 124 6e-29
TC90091 weakly similar to GP|17979475|gb|AAL50074.1 AT3g25150/MJ... 71 8e-13
TC77519 similar to PIR|H86398|H86398 protein F17L21.10 [imported... 55 5e-08
TC86554 similar to PIR|H86398|H86398 protein F17L21.10 [imported... 54 8e-08
TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding pro... 41 7e-04
TC86510 similar to GP|20160777|dbj|BAB89718. putative ribonucleo... 40 0.001
TC87209 similar to GP|9663767|emb|CAC01237.1 RNA Binding Protein... 40 0.002
BE205407 weakly similar to GP|10716605|gb putative RNA binding p... 40 0.002
TC87010 weakly similar to PIR|T48456|T48456 rna binding protein-... 35 0.050
TC87205 similar to GP|12324445|gb|AAG52185.1 putative splicing f... 35 0.050
TC78841 similar to GP|5815235|gb|AAD52609.1| splicing factor SR1... 35 0.065
TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 35 0.065
TC76341 similar to GP|7528270|gb|AAF63202.1| poly(A)-binding pro... 35 0.065
CA859848 similar to GP|12842203|dbj data source:MGD source key:... 34 0.11
TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR recept... 33 0.15
>TC77520 weakly similar to GP|17063173|gb|AAL32982.1 AT5g60980/MSL3_100
{Arabidopsis thaliana}, partial (27%)
Length = 1880
Score = 555 bits (1429), Expect(2) = 0.0
Identities = 279/283 (98%), Positives = 280/283 (98%)
Frame = +2
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA
Sbjct: 119 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 298
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 120
EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK
Sbjct: 299 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 478
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPT 180
GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPT
Sbjct: 479 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPT 658
Query: 181 AQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSI 240
AQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSI
Sbjct: 659 AQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSI 838
Query: 241 EKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHP 283
EKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAK + P
Sbjct: 839 EKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKDSCAP 967
Score = 110 bits (276), Expect(2) = 0.0
Identities = 55/55 (100%), Positives = 55/55 (100%)
Frame = +1
Query: 317 FVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAASMQSALE 371
FVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAASMQSALE
Sbjct: 1069 FVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAASMQSALE 1233
Score = 86.7 bits (213), Expect = 1e-17
Identities = 40/59 (67%), Positives = 43/59 (72%)
Frame = +3
Query: 278 KPAVHPPRVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKK 336
K AVHPPRVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIF + F++
Sbjct: 951 KTAVHPPRVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIFCGKFAYECNSRAIGSGFQE 1127
>TC79275 similar to GP|17063173|gb|AAL32982.1 AT5g60980/MSL3_100
{Arabidopsis thaliana}, partial (26%)
Length = 840
Score = 189 bits (481), Expect = 1e-48
Identities = 101/196 (51%), Positives = 130/196 (65%), Gaps = 4/196 (2%)
Frame = +2
Query: 1 MAVPEA--VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTT 58
MAV EA TP+PQ+VGNAFVEQYY ILH+ PD VHRFY DSS++SR + +G M TVTT
Sbjct: 200 MAVQEANPSTTPSPQVVGNAFVEQYYHILHQSPDLVHRFYQDSSLLSRSDSNGVMKTVTT 379
Query: 59 TAEIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQ 118
T I++ I SL Y E+ +ADAQ SY GV+V+VTGCLTG DN++RKF+Q+FFLAPQ
Sbjct: 380 TQAINETIVSLNYEDCTAEIKTADAQESYEKGVIVLVTGCLTGKDNVRRKFSQTFFLAPQ 559
Query: 119 DKGFYVLNDVFRYVDAYKSIDIESVPANDADE--SAPSEAIITPEPEPVHVPEVIPPTQT 176
DKG+YVLNDVFRY++ NDA + SA +I E VH+PE ++
Sbjct: 560 DKGYYVLNDVFRYIE-----------ENDAPQLSSASINTVINENAEAVHIPE----SED 694
Query: 177 VIPTAQTVIPPTQTVI 192
+ P+ V P + +
Sbjct: 695 INPSEHLVEDPATSAV 742
>BF649128 weakly similar to GP|17979475|gb AT3g25150/MJL12_9 {Arabidopsis
thaliana}, partial (25%)
Length = 527
Score = 162 bits (409), Expect = 3e-40
Identities = 76/133 (57%), Positives = 96/133 (72%)
Frame = +2
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
MA E P P +VG+AFVEQYY +LH P+ VHRFY D S + RPE +G + TT A
Sbjct: 53 MATSENQVPPAPDVVGHAFVEQYYYMLHESPEHVHRFYQDVSKLGRPEPNGIIGITTTMA 232
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 120
EIDKKI S+ Y+ E+LS DAQ S+ GV+V+VTG + G DN+K+KF Q FFLAPQ+K
Sbjct: 233 EIDKKILSMGYSELSAEILSVDAQESFGGGVIVLVTGFMIGKDNVKQKFTQCFFLAPQEK 412
Query: 121 GFYVLNDVFRYVD 133
G++VLND+FRYVD
Sbjct: 413 GYFVLNDIFRYVD 451
>TC87420 weakly similar to GP|19347889|gb|AAL86001.1 unknown protein
{Arabidopsis thaliana}, partial (47%)
Length = 1303
Score = 147 bits (370), Expect = 9e-36
Identities = 112/402 (27%), Positives = 188/402 (45%), Gaps = 25/402 (6%)
Frame = +2
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
MA P + Q +G FV QYY +L P+ VH+FY D+S M R + + T T
Sbjct: 104 MATPFPIPLTAAQ-IGTYFVGQYYHVLQNQPELVHQFYSDASTMLRIDGNAR-ETATAML 277
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 120
+I + SL YT +E+ +A + S++ G +V+V+G + DN++RKF Q+FFLAPQ+K
Sbjct: 278 QIHTLVMSLSYTG--IEIKTAHSLESWSGGAIVMVSGSVQIKDNLRRKFMQTFFLAPQEK 451
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPE---PVHVPEVIP-PTQT 176
GF+VLND+F +V+ +D +A++ + ++VP I P
Sbjct: 452 GFFVLNDIFHFVE------------DDLIHHHHHQAVLLAQSNLDSKLNVPSTINMPVSN 595
Query: 177 VIPTAQT---VIPPTQTV----IADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSH 229
+P+ ++ T V +AD + + ++ + N SS
Sbjct: 596 YMPSGDIQARIVGRTNEVKENGVADNYGYSEQRIQRGPDSEHIREDNAAEDSNGSLHSSG 775
Query: 230 HVKEPEQPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPF-----HLRASPAKPAVHPP 284
+ + P S E+ A Q+ T +ASI+ K S P H SP++ PP
Sbjct: 776 NAVQDHLPASPEEPAGEPQKHT----YASILRVAKGQSTPVASQPSHKNVSPSEWDYIPP 943
Query: 285 RVHSVPAPEA-------PTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKF 337
+ A P ++P + + +++V NL + + +++ FK F
Sbjct: 944 SSNQQSTASANAFERSEPDAVEELPAAEYEDEI---KSVYVRNLTPTVSPSEIEEEFKNF 1114
Query: 338 GPIKRDGIQVRSNK--GSCFGFVEFESAASMQSALEVWSISI 377
G I+ DG+ +RS K G C+ FVEFE + + +A++ S+ I
Sbjct: 1115GRIRPDGVVIRSRKDVGVCYAFVEFEDMSGVHNAVKAGSVEI 1240
>TC88195 similar to GP|19347889|gb|AAL86001.1 unknown protein {Arabidopsis
thaliana}, partial (55%)
Length = 1716
Score = 129 bits (323), Expect(2) = 7e-33
Identities = 106/354 (29%), Positives = 170/354 (47%), Gaps = 16/354 (4%)
Frame = +1
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
VG+ FV QYY +L + PD VH+FY D S M R + D T T + I + SL +++
Sbjct: 67 VGSYFVGQYYQVLRQQPDHVHQFYSDLSSMIRVDGDYT-ETASDVLHIHNIVTSLNFST- 240
Query: 75 RVEVLSADAQPSYNNGVMVVVTGCLTGTD-NIKRKFAQSFFLAPQDKGFYVLNDVFRYVD 133
+E+ + ++ S++ GV+V+VTG + D N K+KF Q+FFLAPQ+KG++VLND+F++VD
Sbjct: 241 -IEIRTINSLDSWDGGVIVMVTGVVKNKDINRKQKFVQTFFLAPQEKGYFVLNDIFQFVD 417
Query: 134 AYKSIDIESVP-ANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQTVI 192
+ VP A+D +S P + EP P + + + P
Sbjct: 418 E-DVVHPNLVPVASDRIDSQPHVSASFAEP-PASDYGFEEEARDYVNSVHIDDDP----- 576
Query: 193 ADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQEDTP 252
D ++ ++ E+ + V + PV H+V + T V + +E
Sbjct: 577 VDKYSLPEQQQQQLQEDFETEVVVDETPVQEASPPVHNVAHTIRETPAAPVEESFEEPA- 753
Query: 253 KKSFASIVNALKD---NSAPFHLRASPAKPAVHPPRVH---SVPAPEAPTPNMDIPLEKN 306
KK++ASI+ A ++AP H V P V + PA + + E
Sbjct: 754 KKTYASILRAKGQSALSAAPQHAPPPSEYNHVTQPAVQQSVAQPAFQQSSSASAYVSESG 933
Query: 307 NENAGRAH--------AIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKG 352
E A + +++V NLP T ++D+ FK FG IK DGI ++ + G
Sbjct: 934 PEAAEEGYRFEEEEVTSVYVRNLPADITEAEIDQEFKNFGRIKPDGIFIKGSTG 1095
Score = 29.6 bits (65), Expect(2) = 7e-33
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +3
Query: 352 GSCFGFVEFESAASMQSALEVWSISI 377
GSC+ FVEFE Q+AL+ I +
Sbjct: 1101 GSCYAFVEFEDVVGTQNALQASPIQL 1178
>TC84615 weakly similar to GP|17063173|gb|AAL32982.1 AT5g60980/MSL3_100
{Arabidopsis thaliana}, partial (20%)
Length = 507
Score = 124 bits (311), Expect = 6e-29
Identities = 57/100 (57%), Positives = 77/100 (77%)
Frame = +3
Query: 9 TPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQS 68
TP+P+MVGNAFVEQYY ILH P+ V+RFY +SSV+SRP+ +G MT+VTT I++KI S
Sbjct: 186 TPSPEMVGNAFVEQYYHILHHSPELVYRFYQESSVISRPDSNGVMTSVTTMKGINEKILS 365
Query: 69 LEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRK 108
L + F+ E+ +ADAQ S+ GV V+VTGCLTG +++K
Sbjct: 366 LNFKEFKAEIKTADAQKSHKEGVTVLVTGCLTGXGXLEKK 485
>TC90091 weakly similar to GP|17979475|gb|AAL50074.1 AT3g25150/MJL12_9
{Arabidopsis thaliana}, partial (9%)
Length = 886
Score = 70.9 bits (172), Expect = 8e-13
Identities = 38/77 (49%), Positives = 51/77 (65%), Gaps = 3/77 (3%)
Frame = +3
Query: 304 EKNNENAG-RAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK--GSCFGFVEF 360
E N+ N H+I++ NLP++ TV QL+ F+KFGPIK IQVR+NK G CFGFVE+
Sbjct: 15 ESNDSNEEVEGHSIYIRNLPLNVTVAQLETEFQKFGPIKPGCIQVRNNKQQGYCFGFVEY 194
Query: 361 ESAASMQSALEVWSISI 377
S SM +A++ I I
Sbjct: 195 LSVNSMNNAIQASPIPI 245
>TC77519 similar to PIR|H86398|H86398 protein F17L21.10 [imported] -
Arabidopsis thaliana, partial (98%)
Length = 1037
Score = 55.1 bits (131), Expect = 5e-08
Identities = 38/127 (29%), Positives = 61/127 (47%), Gaps = 3/127 (2%)
Frame = +1
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKI 66
V+ P + AFVE YY+ + + Y D S+++ + + + I K+
Sbjct: 64 VRAMDPNALSKAFVEHYYTTFDTNRPNLAALYQDGSMLTFEGQQ-----IMGSQNIVTKL 228
Query: 67 QSLEYTSFRVEVLSADAQPS-YNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKG-FY 123
SL + + + D QPS N G++V V+G L + KF+Q F L P +G +Y
Sbjct: 229 TSLPFQQCHHSITTVDCQPSGANGGMLVFVSGNLQLAGEQHALKFSQMFHLIPTPQGSYY 408
Query: 124 VLNDVFR 130
V ND+FR
Sbjct: 409 VWNDIFR 429
>TC86554 similar to PIR|H86398|H86398 protein F17L21.10 [imported] -
Arabidopsis thaliana, partial (97%)
Length = 781
Score = 54.3 bits (129), Expect = 8e-08
Identities = 36/122 (29%), Positives = 61/122 (49%), Gaps = 3/122 (2%)
Frame = +3
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P ++ AFVE YY+ + + Y + S+++ + + + I K+ SL +
Sbjct: 153 PDVLAKAFVEHYYTTFDNNRGGLATLYQEGSMLTFEGQ-----KIQGSPNIVAKLTSLPF 317
Query: 72 TSFRVEVLSADAQPS-YNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKG-FYVLNDV 128
+ + D QPS N G++V V+G L + KF+Q F L P +G +YV+ND+
Sbjct: 318 QQCHHSITTVDCQPSGANGGMLVFVSGNLQLAGEQHALKFSQMFHLMPTPQGSYYVMNDI 497
Query: 129 FR 130
FR
Sbjct: 498 FR 503
>TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding protein
{Nicotiana tabacum}, partial (97%)
Length = 2581
Score = 41.2 bits (95), Expect = 7e-04
Identities = 25/61 (40%), Positives = 31/61 (49%), Gaps = 5/61 (8%)
Frame = +3
Query: 316 IFVANLPMSATVEQLDRAFKKFGPIK-----RDGIQVRSNKGSCFGFVEFESAASMQSAL 370
+FV NL S T ++L + F +FG I RDG K CFGFV FES A+
Sbjct: 987 VFVKNLSESTTDDELKKTFGEFGTITSAVVMRDG----DGKSKCFGFVNFESTDDAARAV 1154
Query: 371 E 371
E
Sbjct: 1155 E 1157
>TC86510 similar to GP|20160777|dbj|BAB89718. putative ribonucleoprotein
{Oryza sativa (japonica cultivar-group)}, partial (67%)
Length = 1587
Score = 40.4 bits (93), Expect = 0.001
Identities = 27/90 (30%), Positives = 43/90 (47%), Gaps = 7/90 (7%)
Frame = +3
Query: 307 NENAGRAHA-----IFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKG--SCFGFVE 359
N G+A +F+ ++P ++L AF+ FG + + V G CFGFV
Sbjct: 1158 NSGGGQAEGPPGANLFIYHIPQEFGDQELANAFQPFGRVLSAKVFVDKATGVSKCFGFVS 1337
Query: 360 FESAASMQSALEVWSISINSCYQQGCKLNV 389
++S + QSA+ + +N C G KL V
Sbjct: 1338 YDSPEAAQSAISM----MNGCQLGGKKLKV 1415
>TC87209 similar to GP|9663767|emb|CAC01237.1 RNA Binding Protein 45
{Nicotiana plumbaginifolia}, partial (69%)
Length = 1504
Score = 40.0 bits (92), Expect = 0.002
Identities = 26/85 (30%), Positives = 39/85 (45%)
Frame = +3
Query: 307 NENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAASM 366
NEN IFV NL + T E L + F ++G + + V+ G GFV+F +S
Sbjct: 837 NENDPNNTTIFVGNLDPNVTDEHLKQVFTQYGEL----VHVKIPSGKRCGFVQFADRSSA 1004
Query: 367 QSALEVWSISINSCYQQGCKLNVGW 391
+ AL V +N G + + W
Sbjct: 1005EEALRV----LNGTLLGGQNVRLSW 1067
>BE205407 weakly similar to GP|10716605|gb putative RNA binding protein
{Oryza sativa}, partial (23%)
Length = 608
Score = 39.7 bits (91), Expect = 0.002
Identities = 26/69 (37%), Positives = 38/69 (54%), Gaps = 2/69 (2%)
Frame = +1
Query: 304 EKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRS--NKGSCFGFVEFE 361
E NE+ R IFV LP + +VE+ R F++FG I + S +K FGF+ F+
Sbjct: 379 ECGNEHI-RTKKIFVGGLPANMSVEEFKRYFERFGRITDVVVMQDSLTHKPRGFGFITFD 555
Query: 362 SAASMQSAL 370
S S+QS +
Sbjct: 556 SEDSVQSVM 582
>TC87010 weakly similar to PIR|T48456|T48456 rna binding protein-like -
Arabidopsis thaliana, partial (77%)
Length = 1097
Score = 35.0 bits (79), Expect = 0.050
Identities = 21/68 (30%), Positives = 36/68 (52%), Gaps = 2/68 (2%)
Frame = +1
Query: 297 PNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQ--VRSNKGSC 354
P +P K EN+ + +++ +P ++++ F +FG IKR I ++ K
Sbjct: 211 PGRKLPELKQLENS--SPVLYIGRIPHGFYEKEMEAYFGQFGTIKRLRISRNKKTGKSRH 384
Query: 355 FGFVEFES 362
FGF+EFES
Sbjct: 385 FGFIEFES 408
>TC87205 similar to GP|12324445|gb|AAG52185.1 putative splicing factor;
53460-55514 {Arabidopsis thaliana}, partial (81%)
Length = 1197
Score = 35.0 bits (79), Expect = 0.050
Identities = 22/85 (25%), Positives = 42/85 (48%)
Frame = +2
Query: 305 KNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAA 364
++N ++ + I+V NLP +++ F K+G I ++V + C+ FVEF++A
Sbjct: 92 QSNMSSRFSRTIYVGNLPADIRESEIEDLFYKYGRIMEIELKV-PPRPPCYCFVEFDNAR 268
Query: 365 SMQSALEVWSISINSCYQQGCKLNV 389
+ A+ + GC+L V
Sbjct: 269 DAEDAIR----GRDGYNFDGCRLRV 331
>TC78841 similar to GP|5815235|gb|AAD52609.1| splicing factor SR1
{Arabidopsis thaliana}, partial (74%)
Length = 1380
Score = 34.7 bits (78), Expect = 0.065
Identities = 18/58 (31%), Positives = 31/58 (53%)
Frame = +3
Query: 313 AHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAASMQSAL 370
+ ++V NLP + +++ F K+GPI +++ K + FVEFE A Q A+
Sbjct: 57 SRTLYVGNLPGDIRLREVEDLFYKYGPIVDIDLKI-PPKPPGYAFVEFEDARDAQDAI 227
>TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (69%)
Length = 1615
Score = 34.7 bits (78), Expect = 0.065
Identities = 48/183 (26%), Positives = 80/183 (43%), Gaps = 10/183 (5%)
Frame = +3
Query: 200 SKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQEDTPKKSFASI 259
+K VS P K S ++ + +SS E ++P + +K S ++ D+
Sbjct: 210 AKPVSKPAAVAKKSKKDSSDSDDEDDDSSSD--EDKKPVAAKKEVSESESDSSDDE--DK 377
Query: 260 VNALKDNS-APFHLRASPAKPAVHPPR----VHSVPAPEAPTPNMDIPLEKNNENAGRAH 314
+N KD S + S +P+ P + V V A ++ + P EN+G +
Sbjct: 378 MNVDKDGSDSDESEEESEDEPSKTPQKKIKDVEMVDAGKSGKKAPNTPATPI-ENSG-SK 551
Query: 315 AIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVR-----SNKGSCFGFVEFESAASMQSA 369
+FV NL S +++ F+ G + + VR + FG VEF SA + QSA
Sbjct: 552 TLFVGNLSFSVQRSDIEKFFQDCGEV----VDVRFSSDEEGRFKGFGHVEFASAEAAQSA 719
Query: 370 LEV 372
LE+
Sbjct: 720 LEM 728
>TC76341 similar to GP|7528270|gb|AAF63202.1| poly(A)-binding protein
{Cucumis sativus}, partial (90%)
Length = 2501
Score = 34.7 bits (78), Expect = 0.065
Identities = 21/58 (36%), Positives = 29/58 (49%), Gaps = 2/58 (3%)
Frame = +3
Query: 316 IFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKG--SCFGFVEFESAASMQSALE 371
++V NL S T + L F +G I + +R G CFGFV FE+A A+E
Sbjct: 837 VYVKNLSESFTEDDLKNEFGAYGTIT-SAVLMRDADGRSKCFGFVNFENAEDAAKAVE 1007
Score = 32.0 bits (71), Expect = 0.42
Identities = 21/69 (30%), Positives = 36/69 (51%), Gaps = 1/69 (1%)
Frame = +3
Query: 304 EKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVR-SNKGSCFGFVEFES 362
+ ++ +G A+ IF+ NL + + L F FG I I S + +GFV+FE+
Sbjct: 531 DPSSRKSGTAN-IFIKNLDKTIDHKALHDTFSSFGQIMSCKIATDGSGQSKGYGFVQFEA 707
Query: 363 AASMQSALE 371
S Q+A++
Sbjct: 708 EDSAQNAID 734
>CA859848 similar to GP|12842203|dbj data source:MGD source key:MGI:1917118
evidence:ISS~peptidylprolyl isomerase E (cyclophilin
E)~, partial (30%)
Length = 387
Score = 33.9 bits (76), Expect = 0.11
Identities = 24/77 (31%), Positives = 38/77 (49%), Gaps = 3/77 (3%)
Frame = +2
Query: 316 IFVANLPMSATVEQLDRAFKKFGPIKRDGIQ---VRSNKGSCFGFVEFESAASMQSALEV 372
I+V L + L AF FG I I N+ FGF+EFE AA Q A++
Sbjct: 47 IYVGGLDDQVNEKILHAAFVPFGDIVEIQIPPDPASHNQHRGFGFIEFEEAADAQEAID- 223
Query: 373 WSISINSCYQQGCKLNV 389
+++++ Y + K+N+
Sbjct: 224 -NMNLSELYGKVIKVNL 271
>TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR receptor protein
kinase {Arabidopsis thaliana}, partial (28%)
Length = 1549
Score = 33.5 bits (75), Expect = 0.15
Identities = 33/129 (25%), Positives = 51/129 (38%), Gaps = 6/129 (4%)
Frame = +1
Query: 175 QTVIPTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTEN-VIPVNHVKESSHHVKE 233
QT V+PP+QT E + +++VS P + + + +P K+ +
Sbjct: 613 QTPSSLRAIVLPPSQT-----EKVPARDVSRPNDVRQEEPRKVWAVPNAQDKQEKDVQRM 777
Query: 234 PEQPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHP-----PRVHS 288
P + + N QE+ + I NA A+ AKP H P V+S
Sbjct: 778 TAIPRDVLRPNDNRQEEPRRVWAVPIPNAHDKQEKDVQRMATIAKPVDHEIDISTPEVYS 957
Query: 289 VPAPEAPTP 297
VP P P P
Sbjct: 958 VPPPPPPPP 984
Score = 27.7 bits (60), Expect = 8.0
Identities = 22/73 (30%), Positives = 32/73 (43%)
Frame = +1
Query: 127 DVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIP 186
DV R K +D ++ D S P E P P P P PP+ IPT + ++
Sbjct: 883 DVQRMATIAKPVD------HEIDISTP-EVYSVPPPPPPPPPPPPPPS---IPTKKGIVE 1032
Query: 187 PTQTVIADTETII 199
PT + T+ +I
Sbjct: 1033PTTSHSLPTKRVI 1071
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.129 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,157,112
Number of Sequences: 36976
Number of extensions: 161249
Number of successful extensions: 1364
Number of sequences better than 10.0: 110
Number of HSP's better than 10.0 without gapping: 1153
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1328
length of query: 393
length of database: 9,014,727
effective HSP length: 98
effective length of query: 295
effective length of database: 5,391,079
effective search space: 1590368305
effective search space used: 1590368305
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146567.10


