

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
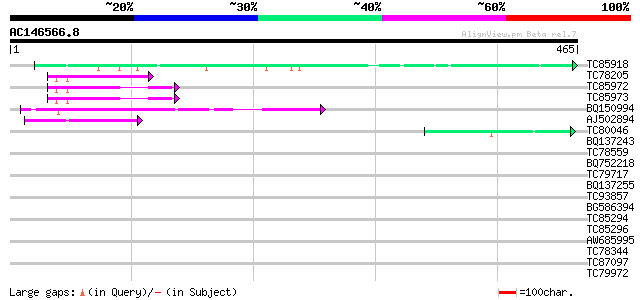
Query= AC146566.8 + phase: 0
(465 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreu... 63 2e-10
TC78205 homologue to PIR|T47559|T47559 60S ribosomal protein-lik... 56 3e-08
TC85972 homologue to SP|O65743|RL24_CICAR 60S ribosomal protein ... 54 2e-07
TC85973 homologue to SP|O65743|RL24_CICAR 60S ribosomal protein ... 54 2e-07
BQ150994 similar to GP|22122173|dbj preproneuropeptide B {Homo s... 50 1e-06
AJ502894 weakly similar to SP|P38665|RL24 60S ribosomal protein ... 46 3e-05
TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor ... 42 7e-04
BQ137243 similar to GP|2624970|gb|A proline-rich protein 9-1 {Mu... 41 0.001
TC78559 weakly similar to GP|5733686|gb|AAD49719.1| maturation p... 40 0.002
BQ752218 similar to GP|19170914|emb hypothetical protein {Enceph... 39 0.003
TC79717 similar to GP|10086499|gb|AAG12559.1 Unknown Protein {Ar... 39 0.004
BQ137255 homologue to PIR|B34768|B347 ORF5 protein - Orf virus (... 39 0.006
TC93857 similar to SP|Q05049|MUC1_XENLA Integumentary mucin C.1 ... 38 0.007
BG586394 similar to GP|7295463|gb| CG4835 gene product {Drosophi... 37 0.012
TC85294 similar to GP|2961298|emb|CAA12357.1 hypothetical protei... 36 0.027
TC85296 weakly similar to GP|2505870|emb|CAA72908.1 hypothetical... 35 0.047
AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - A... 35 0.061
TC78344 similar to GP|13540405|gb|AAK29456.1 histone H1 {Lens cu... 35 0.080
TC87097 similar to GP|14334540|gb|AAK59678.1 putative protein {A... 34 0.10
TC79972 similar to GP|8778288|gb|AAF79297.1| F14D16.30 {Arabidop... 34 0.10
>TC85918 similar to GP|13562014|gb|AAK30610.1 fibroin 1 {Plectreurys tristis},
partial (8%)
Length = 4117
Score = 63.2 bits (152), Expect = 2e-10
Identities = 99/484 (20%), Positives = 187/484 (38%), Gaps = 39/484 (8%)
Frame = -1
Query: 21 KEDDLVEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTESSTVSASEEVIQ---KKRNRE 77
KED++ E K ETV+ PK+ K + TES T+S +E V +
Sbjct: 3019 KEDEVKAEPAVAAEKTDDSVPIETVS-DNPKEEKDLITESETISVTEPVNDTAISHHDET 2843
Query: 78 PEVQDAAREAA------LREDATKREAALQAI-------RSKRAKGERSIQERIREAAEQ 124
P + A+E + L E + + ++ I +S A +E++++A +
Sbjct: 2842 PSTESDAKETSQQIEKELVETNEENQPKIEDIPEAETTEKSSDATETNQFEEKVKQA--E 2669
Query: 125 LRKEEADPSLKKAQKPVSLESPCFRLTPEQEERTRE----------IAANGLAKRRQMTE 174
+ + A+ K +P++ E P PE+E + + I + + + T
Sbjct: 2668 ISETSAEKEEKPELEPLATEEPKVTKEPEKESQEKSEEAEQPKAIAIVESTVEANEEKTA 2489
Query: 175 IYKRNRDEQLKAAGYELDAEKAAVIASLAAEIEEE----TVREGAALLKQALKAKHVSG- 229
+ ++ E E+ V AEI +E T E LK+ L A V
Sbjct: 2488 AAISEETKIIEVEPTETVKEEPMVTEVATAEIVKEEPVVTEVEATETLKEELVATEVEAT 2309
Query: 230 -ATSSEPA------SEAPKSEATRPEAHPS-GIPSVPKASVNIQTPILPSSPSSSSSSSS 281
EPA +E K E E P+ + P A+ T + P ++ +
Sbjct: 2308 ETVKEEPAATEVKVTETVKDEPVVTEVDPTETVKEEPLATEAEATETVKEEPVATEVEPT 2129
Query: 282 TNSDDIPLSQHIKQCLPNFKPSTSTFQDDYEQMQIHFSEQRIKICENLNLSADHFFQPPL 341
+ P++ ++ +T T +D+ ++ +E + E + +
Sbjct: 2128 ETVKEEPVATEVE--------ATETVKDEPAATEVEATE---TVKEEPVATETEATETVK 1982
Query: 342 VEPLNIQHPETTPTPQKASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQPTSTKASKT 401
EP+ + E T T ++ S VA E +E ++ +T+ + + E T +A++T
Sbjct: 1981 EEPVATE-VEATETVKEES-VATEVEATETVKEEPVTTVVEPTEMVKDEPAATEFEATET 1808
Query: 402 VPEKTVLENQPETSTVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHSPTQNQTIPEQQ 461
V ++T++ T TV E++V K V + P TE++ P + P +
Sbjct: 1807 VKDETIVTEVDPTETVKEEAVATEVEITKTVKE-PLVTEVDLTESVKVKPVVTEVDPREM 1631
Query: 462 TTSD 465
+
Sbjct: 1630 VKEE 1619
>TC78205 homologue to PIR|T47559|T47559 60S ribosomal protein-like -
Arabidopsis thaliana, partial (93%)
Length = 858
Score = 56.2 bits (134), Expect = 3e-08
Identities = 39/95 (41%), Positives = 57/95 (59%), Gaps = 8/95 (8%)
Frame = +1
Query: 32 KPSKRS-----RKAEKETVA--APKPKKGKTVKTES-STVSASEEVIQKKRNREPEVQDA 83
KPSK + RK K+ +A A K ++ T K S S V A+ EVIQKKR+ +PEV+DA
Sbjct: 193 KPSKLTWTAMYRKQHKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRSEKPEVRDA 372
Query: 84 AREAALREDATKREAALQAIRSKRAKGERSIQERI 118
AREAALRE + + ++K+A+ + Q+ +
Sbjct: 373 AREAALREIKERVKKTKDEKKAKKAESQAKAQKSV 477
>TC85972 homologue to SP|O65743|RL24_CICAR 60S ribosomal protein L24.
[Chickpea Garbanzo] {Cicer arietinum}, partial (98%)
Length = 764
Score = 53.5 bits (127), Expect = 2e-07
Identities = 45/116 (38%), Positives = 63/116 (53%), Gaps = 8/116 (6%)
Frame = +1
Query: 32 KPSKRS-----RKAEKETVA--APKPKKGKTVKTES-STVSASEEVIQKKRNREPEVQDA 83
KPSK + RK K+ +A A K ++ T K S S V A+ EVIQKKR +PEV+DA
Sbjct: 211 KPSKLTWTAMYRKQHKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDA 390
Query: 84 AREAALREDATKREAALQAIRSKRAKGERSIQERIREAAEQLRKEEADPSLKKAQK 139
AREAALRE I+ERI++ ++ + ++A+ + KAQK
Sbjct: 391 AREAALRE----------------------IKERIKKTKDEKKAKKAEVA-SKAQK 489
>TC85973 homologue to SP|O65743|RL24_CICAR 60S ribosomal protein L24.
[Chickpea Garbanzo] {Cicer arietinum}, partial (98%)
Length = 725
Score = 53.5 bits (127), Expect = 2e-07
Identities = 45/116 (38%), Positives = 63/116 (53%), Gaps = 8/116 (6%)
Frame = +2
Query: 32 KPSKRS-----RKAEKETVA--APKPKKGKTVKTES-STVSASEEVIQKKRNREPEVQDA 83
KPSK + RK K+ +A A K ++ T K S S V A+ EVIQKKR +PEV+DA
Sbjct: 197 KPSKLTWTAMYRKQHKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDA 376
Query: 84 AREAALREDATKREAALQAIRSKRAKGERSIQERIREAAEQLRKEEADPSLKKAQK 139
AREAALRE I+ERI++ ++ + ++A+ + KAQK
Sbjct: 377 AREAALRE----------------------IKERIKKTKDEKKAKKAEVA-SKAQK 475
>BQ150994 similar to GP|22122173|dbj preproneuropeptide B {Homo sapiens},
partial (15%)
Length = 1036
Score = 50.4 bits (119), Expect = 1e-06
Identities = 58/255 (22%), Positives = 104/255 (40%), Gaps = 5/255 (1%)
Frame = +3
Query: 10 KSRKTKSKALTKEDDLVEEATRKPSKRSR---KAEKETVAAPKPKKGKTVKTESSTVSAS 66
+ R+ S+A E + E T+K ++R R K E T P+P++G T +T A
Sbjct: 309 RRRRETSRA---EGEREREGTKKEARRKRARRKKETPTPDGPRPRRGITRPERQATRRAE 479
Query: 67 EEVIQKKRNREPEVQDAAREAAL-REDATKREAALQAIRSKRAKGERSIQERIREAAEQL 125
+KKR EPE + RE + +A +REA +A R R+ A+++
Sbjct: 480 RRRAEKKRRTEPEQKPNHREKRPDKREAARREAERRAKRDATEHAARTRPTTPTHASDEE 659
Query: 126 RKEEADPSLKKAQKPVSLESPCFRLTPEQEERTREIAANGLAKRRQMTEIYKRNRDEQLK 185
R A + ++ Q ++ + T +I+ N ++ TE R R+ +
Sbjct: 660 RGRAARDAHRR-QPATHDNKTRHQVRHHRHRHTSQISQN---RK*TKTEAPARARESK-- 821
Query: 186 AAGYELDAEKAAVIASLAAEIEEETVREGA-ALLKQALKAKHVSGATSSEPASEAPKSEA 244
E + ++GA A + A + K+ +G PA+ + + +
Sbjct: 822 ---------------------ETNSRKQGAVARARDARRNKNNTGIIKEHPANRSSRRDP 938
Query: 245 TRPEAHPSGIPSVPK 259
P+ P G +V K
Sbjct: 939 REPDHQPPGSSAVHK 983
>AJ502894 weakly similar to SP|P38665|RL24 60S ribosomal protein L24 (L30).
[Yeast] {Kluyveromyces lactis}, partial (84%)
Length = 501
Score = 46.2 bits (108), Expect = 3e-05
Identities = 31/97 (31%), Positives = 53/97 (53%)
Frame = +2
Query: 13 KTKSKALTKEDDLVEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTESSTVSASEEVIQK 72
K++S L +++ T + +K E VA K + + VK + + V AS E I+
Sbjct: 170 KSESLFLQRKNPRKISWTTVYRRMHKKGITEEVA--KKRTRRAVKHQRAIVGASWEAIKA 343
Query: 73 KRNREPEVQDAAREAALREDATKREAALQAIRSKRAK 109
KRN++PEV++AAR A RE ++A ++++AK
Sbjct: 344 KRNQKPEVREAARAEAARESKENKKAEQSKRKAEKAK 454
>TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor (EC
3.2.1.4) (Endo-1 4-beta-glucanase) (Spore germination
protein 270-6), partial (11%)
Length = 643
Score = 41.6 bits (96), Expect = 7e-04
Identities = 36/128 (28%), Positives = 50/128 (38%), Gaps = 4/128 (3%)
Frame = +3
Query: 341 LVEPLNIQHPETTPTPQKASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQP---TSTK 397
++EP+ + T T V + + P ETST + E + TST+
Sbjct: 174 IIEPVTVTSTATITTTVSPITVTDTSTNTVSPSPTETSTETSTETPTETSTETPTETSTE 353
Query: 398 ASKTVPEKTVLENQPETSTVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHS-PTQNQT 456
S P +T E ETST T E ++ P P+TE TE S T +T
Sbjct: 354 TSTETPTETSTETPTETSTETPTETSTETSTE-TSTETPPPSTETSTETTPPSTETSTET 530
Query: 457 IPEQQTTS 464
P TS
Sbjct: 531 TPPSTETS 554
Score = 34.7 bits (78), Expect = 0.080
Identities = 32/125 (25%), Positives = 46/125 (36%), Gaps = 2/125 (1%)
Frame = +3
Query: 342 VEPLNIQHPETTPTPQKASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQP-TSTKASK 400
V P+ + T +E + E +T E P + T T E TST+
Sbjct: 222 VSPITVTDTSTNTVSPSPTETSTETST-ETPTETSTETPTETSTETSTETPTETSTETPT 398
Query: 401 TVPEKTVLENQPETSTVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHS-PTQNQTIPE 459
+T E ETST P + + ++ P+TE TE S T +T P
Sbjct: 399 ETSTETPTETSTETSTETSTETPPPST--ETSTETTPPSTETSTETTPPSTETSTETTPP 572
Query: 460 QQTTS 464
TS
Sbjct: 573 STETS 587
Score = 33.9 bits (76), Expect = 0.14
Identities = 30/105 (28%), Positives = 42/105 (39%), Gaps = 2/105 (1%)
Frame = +3
Query: 344 PLNIQHPETTPTPQKASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQPTSTKAS-KTV 402
P +T TP + S ++E P + T T E P ST+ S +T
Sbjct: 333 PTETSTETSTETPTETSTETPTETSTETPTETSTET----STETSTETPPPSTETSTETT 500
Query: 403 PEKTVLENQPETSTVPEQSVPEHTVPEKVVS-DLPQPNTEQQTEN 446
P T E ET+ ++ E T P S + P+TE TEN
Sbjct: 501 PPST--ETSTETTPPSTETSTETTPPSTETSTETTPPSTETSTEN 629
Score = 32.0 bits (71), Expect = 0.52
Identities = 21/75 (28%), Positives = 33/75 (44%), Gaps = 3/75 (4%)
Frame = +3
Query: 394 TSTKASKTVPE---KTVLENQPETSTVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHS 450
T+T + TV + TV + ETST P T E + +TE TE +
Sbjct: 213 TTTVSPITVTDTSTNTVSPSPTETSTETSTETPTETSTETPTETSTETSTETPTETSTET 392
Query: 451 PTQNQTIPEQQTTSD 465
PT+ T +T+++
Sbjct: 393 PTETSTETPTETSTE 437
>BQ137243 similar to GP|2624970|gb|A proline-rich protein 9-1 {Mus musculus},
partial (13%)
Length = 1145
Score = 40.8 bits (94), Expect = 0.001
Identities = 43/201 (21%), Positives = 67/201 (32%), Gaps = 7/201 (3%)
Frame = +3
Query: 3 GGALPVAKSRKTKSKALTKEDDLVEEATRKPSKRSRKAEKETVAAP-------KPKKGKT 55
GG P ++R T+ A E TRKP R EKE A +P++
Sbjct: 108 GGGAPGGRARHTRPTA---------EPTRKPKGRREHREKEHALAENQRQSTGRPQRKVR 260
Query: 56 VKTESSTVSASEEVIQKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQ 115
+ A KK ++ E + R + +D A R +RA + Q
Sbjct: 261 KRVIKRNQGADRRKANKKGKKKEEGERGERTSRRDDDDEPTRTATDTRRQRRASTQTPNQ 440
Query: 116 ERIREAAEQLRKEEADPSLKKAQKPVSLESPCFRLTPEQEERTREIAANGLAKRRQMTEI 175
+ R EQ R ++ + ++ P T Q RE + +
Sbjct: 441 KHARTIEEQHRAKQRGDDRRT*GSRTAVRRPRPATTQRQHTAARETRRSARRTTAAKLDT 620
Query: 176 YKRNRDEQLKAAGYELDAEKA 196
RD + E + EKA
Sbjct: 621 TAGGRDRRTTERRIEREREKA 683
Score = 30.8 bits (68), Expect = 1.2
Identities = 33/123 (26%), Positives = 54/123 (43%), Gaps = 4/123 (3%)
Frame = +1
Query: 12 RKTKSKALTKEDDLVEEATRKP-SKRSRKAEKETVAAPKPKKGKTVKTESSTVSASEEVI 70
R+++ A T+ VE +TR + KA +ET + G + T SS +A
Sbjct: 700 RRSRQTADTRTP--VETSTRDELADEESKAAEETERRIRTYSGSEMNTTSSARNARNTAN 873
Query: 71 QKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKG---ERSIQERIREAAEQLRK 127
+K + AAR + T+RE ALQ R ++ +G + + E+ A Q +
Sbjct: 874 TRKSS-------AARTTEKNQRDTRREDALQRARKRQRRGRPRDPASTEQKATARTQTKH 1032
Query: 128 EEA 130
E A
Sbjct: 1033ESA 1041
>TC78559 weakly similar to GP|5733686|gb|AAD49719.1| maturation protein
pPM32 {Glycine max}, partial (55%)
Length = 1128
Score = 40.0 bits (92), Expect = 0.002
Identities = 53/239 (22%), Positives = 100/239 (41%), Gaps = 27/239 (11%)
Frame = +2
Query: 28 EATRKPSKR------SRKAEKETVAAPKPKKGKTVKTESSTVS-----ASEEVIQKKRNR 76
E+TRKPS S K +++ A + ++ + +S + A+E++ Q R+
Sbjct: 32 ESTRKPSTNWAYDHSSSKTKRDAEIADRARETMSEGVDSEDIKQYSRDANEDIKQFARDT 211
Query: 77 EPEVQDAAREAALREDATKREAALQAIRSKRAKGERS------IQERIREAAEQLRKEEA 130
+ +DAA ++ E A +E +A GE++ +++R EAA + +
Sbjct: 212 NEKTKDAA--GSMWEKA--KEGTDRAAEKAENAGEKARDYAYEVKDRTNEAAGSVWDKAR 379
Query: 131 DPSLKKAQKPVSLESPCFRLTPEQEERTREIAANGLAKRRQMTEIYKRNRDEQLKAAG-- 188
+ + + A+K + + +ERT++ A N R K E ++AG
Sbjct: 380 EGTNRAAEKAENAGEKARDYAYDAKERTKDAAENAGESIRDYAYDAKDRTKEAAESAGEK 559
Query: 189 ---YELDA-----EKAAVIASLAAEIEEETVREGAALLKQALKAKHVSGATSSEPASEA 239
Y DA E A+ +A E E+T + + A +A +G + + A A
Sbjct: 560 ARDYAYDAKERTKEGASYMADKTKEGAEKTAEKTGEVAGSATEALKSAGEMAKKTAQGA 736
>BQ752218 similar to GP|19170914|emb hypothetical protein {Encephalitozoon
cuniculi}, partial (5%)
Length = 637
Score = 39.3 bits (90), Expect = 0.003
Identities = 30/111 (27%), Positives = 58/111 (52%)
Frame = -2
Query: 71 QKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQERIREAAEQLRKEEA 130
+K + + + ARE A ++ REAA + + ++ + E+ +E R+A +++ KEE
Sbjct: 609 EKAKKEAEKKKKKAREEAAKKAKKAREAAYKKAKEEKKEAEKKAKEEKRQAEKKI-KEEK 433
Query: 131 DPSLKKAQKPVSLESPCFRLTPEQEERTREIAANGLAKRRQMTEIYKRNRD 181
+ KKA+K +++E+ R+ A + L+ RQ TE K ++D
Sbjct: 432 KEAEKKAKK-------------DKKEQDRKDAESALS--RQDTEDSKESKD 325
>TC79717 similar to GP|10086499|gb|AAG12559.1 Unknown Protein {Arabidopsis
thaliana}, partial (5%)
Length = 1579
Score = 38.9 bits (89), Expect = 0.004
Identities = 44/188 (23%), Positives = 70/188 (36%), Gaps = 26/188 (13%)
Frame = +2
Query: 302 PSTSTFQDDYEQMQIHFSEQRIKICENLNLSADHFFQPPLVEPLNIQHPETTPTPQKASE 361
PS Q +E++ + + R + ++S + + I +TP P + ++
Sbjct: 353 PSFIAAQSKFEELSSNSNSGRPSGLLDQDVSVESQADSAYISKEFISSENSTPYPSRNAD 532
Query: 362 -----VALEAATSEDPQQHETSTLHNFEKHLGGEMQPTSTKASKTVPEKTVLENQP---- 412
V ++T + P + ET + + K L + K V T N P
Sbjct: 533 PESGTVLSISSTLDSPDRSETLEIEHDAKDLVEGIVNPENKTDHGVEANTPTSNLPISDS 712
Query: 413 ---ET----------STVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNH----SPTQNQ 455
ET S +PE S PEK SDL + TE ++ + SP
Sbjct: 713 DQLETVNGSRGNVVDSVMPENSKEHAVEPEKTASDLLREQTETVLQDFTYSQQASPGSYM 892
Query: 456 TIPEQQTT 463
TIPE Q T
Sbjct: 893 TIPESQGT 916
>BQ137255 homologue to PIR|B34768|B347 ORF5 protein - Orf virus (strain NZ2),
partial (10%)
Length = 1161
Score = 38.5 bits (88), Expect = 0.006
Identities = 58/253 (22%), Positives = 90/253 (34%), Gaps = 30/253 (11%)
Frame = +3
Query: 12 RKTKSKALTKEDDLVEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTES-------STVS 64
R ++ TK D A +++S +E+ T A + + V+ + T S
Sbjct: 408 RLRRAAGTTKPSDRQSSARGASARQSGASERNTERAERRARASRVRAHAMRGRRAHETTS 587
Query: 65 ASEEVIQKKRNREPEVQDAAREAALREDATKREAALQAIR-------------------- 104
A EE + + DAAR+ R REA Q R
Sbjct: 588 AREEE*RGAEDGRRASDDAARQRRRRRAPRAREAHRQRARRARAARDRRRPAPRRKRARS 767
Query: 105 --SKRAKGERSIQERIREAAEQLRKEEADPSLKKAQKPVSLESPCFRLTPEQEERTREIA 162
++RA+ SIQE +A + E AD +A++ S R T + +T+E
Sbjct: 768 RAARRARARHSIQEDADDARDDEIDESADTRAARARRRDS-RRRASRATAARARQTKE-- 938
Query: 163 ANGLAKRRQMTEIYKRNRDE-QLKAAGYELDAEKAAVIASLAAEIEEETVREGAALLKQA 221
RR+ + R DE + A G + A +A ET + A +A
Sbjct: 939 ------RRERQATHARVLDETRAHADGRDTRARATMTRDVASAARHGETRADACAARARA 1100
Query: 222 LKAKHVSGATSSE 234
A A SE
Sbjct: 1101RAAARTGRARGSE 1139
>TC93857 similar to SP|Q05049|MUC1_XENLA Integumentary mucin C.1 (FIM-C.1)
(Fragment). [African clawed frog] {Xenopus laevis},
partial (8%)
Length = 763
Score = 38.1 bits (87), Expect = 0.007
Identities = 23/92 (25%), Positives = 40/92 (43%)
Frame = +2
Query: 372 PQQHETSTLHNFEKHLGGEMQPTSTKASKTVPEKTVLENQPETSTVPEQSVPEHTVPEKV 431
P+ + T+T N + + + + T + V +K L + T+ V ++ P HT+
Sbjct: 35 PKPNHTTTTTNVTQ----KPKLSHTTTTTNVTQKPKLSHTTTTTNVTQKPKPSHTITTTN 202
Query: 432 VSDLPQPNTEQQTENLNHSPTQNQTIPEQQTT 463
V+ P+PN T N+ P N T T
Sbjct: 203 VTQKPKPNPTTTTTNVTPKPKPNPTTTTTNVT 298
Score = 36.6 bits (83), Expect = 0.021
Identities = 25/104 (24%), Positives = 38/104 (36%)
Frame = +2
Query: 353 TPTPQKASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQPTSTKASKTVPEKTVLENQP 412
TP P+ T + H T+T + K +P+ T + V +K
Sbjct: 251 TPKPKPNPTTTTTNVTPKPKPNHTTTTTNVTPKP-----KPSHTTTTTNVTQKPKPNPTT 415
Query: 413 ETSTVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHSPTQNQT 456
T+ V + P HT V+ P+PN T N+ P T
Sbjct: 416 TTTNVAPKPKPNHTTTTTNVTQKPKPNHTTTTTNVTQKPNHTTT 547
Score = 36.2 bits (82), Expect = 0.027
Identities = 28/120 (23%), Positives = 43/120 (35%), Gaps = 3/120 (2%)
Frame = +2
Query: 347 IQHPETTPT---PQKASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQPTSTKASKTVP 403
+ H TT K S ++ P+ + T+T N + +P T + V
Sbjct: 131 LSHTTTTTNVTQKPKPSHTITTTNVTQKPKPNPTTTTTNVTP----KPKPNPTTTTTNVT 298
Query: 404 EKTVLENQPETSTVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHSPTQNQTIPEQQTT 463
K + T+ V + P HT V+ P+PN T N+ P N T T
Sbjct: 299 PKPKPNHTTTTTNVTPKPKPSHTTTTTNVTQKPKPNPTTTTTNVAPKPKPNHTTTTTNVT 478
>BG586394 similar to GP|7295463|gb| CG4835 gene product {Drosophila
melanogaster}, partial (2%)
Length = 734
Score = 37.4 bits (85), Expect = 0.012
Identities = 50/251 (19%), Positives = 92/251 (35%), Gaps = 13/251 (5%)
Frame = +1
Query: 227 VSGATSS-EPASEAPKSEATRPEAHPSGIPSVP------KASVNIQTPILPSSPSSSSSS 279
V+G T+ E + +SE P + +P K + T +P+ S ++
Sbjct: 4 VTGTTTQPEKETTTTESETNTPTKSETDVPETTTTIEKSKNDLGTTTESETETPTKSETN 183
Query: 280 SSTNSDDIPLSQHIKQCLPNFKP----STSTFQDDYEQMQIHFSEQRIKICENLNLSADH 335
+ P+ Q + P KP +T T + + E + K ++ +
Sbjct: 184 VPETTTTEPIVQTDSEETPTEKPEKETTTKTTKPEAETTTEPETNTPTKSETDVPETTTT 363
Query: 336 FFQPPLVEPLNIQHPETTPTPQKASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQPTS 395
+ V + TPT + E +E P+ T EK+ +PT
Sbjct: 364 LEKNQNVPGSTTESETETPTKSETETTTTEETPTEKPETDIPKTTTASEKNKTTTTEPTV 543
Query: 396 TKASKTVPEKTVLENQPETSTVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHSPTQNQ 455
S+ P + + + ET+T PE + T+ +T+N +P +N+
Sbjct: 544 PTVSEETPTE---KPEKETTTTPE----------------TEATTQPETDNTPTNPEKNK 666
Query: 456 --TIPEQQTTS 464
T PE +TT+
Sbjct: 667 ATTTPETETTT 699
Score = 33.9 bits (76), Expect = 0.14
Identities = 54/267 (20%), Positives = 93/267 (34%), Gaps = 3/267 (1%)
Frame = +1
Query: 21 KEDDLVEEATRKPSKRSRKAEKETVAAPKPKK--GKTVKTESSTVSASEEVIQKKRNREP 78
KE E T P+K + T K K G T ++E+ T + SE + + EP
Sbjct: 31 KETTTTESETNTPTKSETDVPETTTTIEKSKNDLGTTTESETETPTKSETNVPETTTTEP 210
Query: 79 EVQDAAREAALREDATKREAALQAIRSKRAKGERSIQ-ERIREAAEQLRKEEADPSLKKA 137
VQ + E T++ ++ + + E + + E + E +L+K
Sbjct: 211 IVQTDSEET-----PTEKPEKETTTKTTKPEAETTTEPETNTPTKSETDVPETTTTLEKN 375
Query: 138 QKPVSLESPCFRLTPEQEERTREIAANGLAKRRQMTEIYKRNRDEQLKAAGYELDAEKAA 197
Q + TP + E T + T+I K + K
Sbjct: 376 QNVPGSTTESETETPTKSE-TETTTTEETPTEKPETDIPKTTTAS---------EKNKTT 525
Query: 198 VIASLAAEIEEETVREGAALLKQALKAKHVSGATSSEPASEAPKSEATRPEAHPSGIPSV 257
+ EET E T++ P +EA T+PE P+
Sbjct: 526 TTEPTVPTVSEETPTE------------KPEKETTTTPETEA----TTQPET--DNTPTN 651
Query: 258 PKASVNIQTPILPSSPSSSSSSSSTNS 284
P+ + TP ++ +S ++ S TN+
Sbjct: 652 PEKNKATTTPETETTTASETNVSETNT 732
>TC85294 similar to GP|2961298|emb|CAA12357.1 hypothetical protein {Cicer
arietinum}, partial (61%)
Length = 1353
Score = 36.2 bits (82), Expect = 0.027
Identities = 23/75 (30%), Positives = 35/75 (46%), Gaps = 10/75 (13%)
Frame = +2
Query: 392 QPTSTKASKTVPEKTVLENQPET----STVPEQSVPEHTVPEKVVSDLP------QPNTE 441
Q S + ++ P TV E PE T+PEQ+ E PE+ +++P + TE
Sbjct: 263 QMASVEVAQVPPPTTVPETTPEVIKTEETIPEQTAIEVPAPEQPEAEVPATEVPTEETTE 442
Query: 442 QQTENLNHSPTQNQT 456
Q TE +P +T
Sbjct: 443 QPTETTEETPAAVET 487
>TC85296 weakly similar to GP|2505870|emb|CAA72908.1 hypothetical protein
{Arabidopsis thaliana}, partial (3%)
Length = 774
Score = 35.4 bits (80), Expect = 0.047
Identities = 22/66 (33%), Positives = 32/66 (48%), Gaps = 4/66 (6%)
Frame = +1
Query: 392 QPTSTKASKTVPEKTVLENQPET----STVPEQSVPEHTVPEKVVSDLPQPNTEQQTENL 447
Q S + ++ P TV E PE T+PEQ+ E PE+ +++P TE TE
Sbjct: 556 QMASVEVAQVPPPTTVPETTPEVIKTEETIPEQTAIEVPAPEQPEAEVPA--TEVPTEET 729
Query: 448 NHSPTQ 453
PT+
Sbjct: 730 TEQPTE 747
>AW685995 similar to PIR|I51618|I516 nucleolar phosphoprotein - African
clawed frog, partial (5%)
Length = 516
Score = 35.0 bits (79), Expect = 0.061
Identities = 32/127 (25%), Positives = 55/127 (43%), Gaps = 5/127 (3%)
Frame = +3
Query: 9 AKSRKTKSKALTKEDDL-----VEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTESSTV 63
AK+ + K K K+++ VEE + + S ++ P K K ES
Sbjct: 108 AKAEEEKKKQQKKKEEKPAPMEVEEDSSSSEESSSSEDEAPAKKAAPAKKAAAKKES--- 278
Query: 64 SASEEVIQKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQERIREAAE 123
S+ EE + E E + A + AA +E +++ E++ + E S +E EA
Sbjct: 279 SSEEESSSSESESEEEEKPAKKAAAKKESSSEEESS--------SSEEESDEEEKNEARR 434
Query: 124 QLRKEEA 130
+ +EEA
Sbjct: 435 R*EQEEA 455
>TC78344 similar to GP|13540405|gb|AAK29456.1 histone H1 {Lens culinaris},
partial (86%)
Length = 1215
Score = 34.7 bits (78), Expect = 0.080
Identities = 56/238 (23%), Positives = 92/238 (38%), Gaps = 1/238 (0%)
Frame = +2
Query: 26 VEEATRKPSKRSRKAEKETVAAPKPK-KGKTVKTESSTVSASEEVIQKKRNREPEVQDAA 84
V+ + + P+K + A+K A PKPK K K ++ + + K + + + A
Sbjct: 443 VKGSYKLPAKSAAPAKKPAAAKPKPKPKAKAPVAKAPAAKSKAKAAPAKAKAKAKAKAAP 622
Query: 85 REAALREDATKREAALQAIRSKRAKGERSIQERIREAAEQLRKEEADPSLKKAQKPVSLE 144
+A + A K + A +A + +AK + + +A + K A P KA KP +
Sbjct: 623 AKA---KPAAKAKPAAKAKPAAKAK---PVAKAKPKAKAPVAKANAVPVAAKA-KPAAKA 781
Query: 145 SPCFRLTPEQEERTREIAANGLAKRRQMTEIYKRNRDEQLKAAGYELDAEKAAVIASLAA 204
P AK + K +R + G A K A
Sbjct: 782 KPA-------------------AKAKPAARPAKASRTSTRTSPGKRAAAAKPA------- 883
Query: 205 EIEEETVREGAALLKQALKAKHVSGATSSEPASEAPKSEATRPEAHPSGIPSVPKASV 262
V++ A +K+A+ AK AT + A+ A K+ A R G + KASV
Sbjct: 884 ------VKKAATPVKKAVPAK--KAATPVKKAAPARKAPAKRGGRK*KGGGVIRKASV 1033
Score = 33.9 bits (76), Expect = 0.14
Identities = 42/148 (28%), Positives = 63/148 (42%), Gaps = 8/148 (5%)
Frame = +2
Query: 7 PVAKSR--KTKSKALTKEDDLVEEATRKPSKRSRKAEKETVAAPKP-KKGKTVKTESSTV 63
PVAK+ K+K+KA + +A P+K A+ + A KP K K V
Sbjct: 536 PVAKAPAAKSKAKAAPAKAKAKAKAKAAPAKAKPAAKAKPAAKAKPAAKAKPVAKAKPKA 715
Query: 64 SASEEVIQKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQERIREAAE 123
A K N P V A+ AA + A K + A + ++ R S +R A
Sbjct: 716 KAP----VAKANAVP-VAAKAKPAAKAKPAAKAKPAARPAKASRTSTRTSPGKRAAAAKP 880
Query: 124 QLRK-----EEADPSLKKAQKPVSLESP 146
++K ++A P+ KKA PV +P
Sbjct: 881 AVKKAATPVKKAVPA-KKAATPVKKAAP 961
Score = 28.1 bits (61), Expect = 7.5
Identities = 25/97 (25%), Positives = 46/97 (46%), Gaps = 3/97 (3%)
Frame = +2
Query: 5 ALPVAKSRKTKSKALTKEDDLVEEATRKPSKRSRKAEKETVAAPKPKKGKT-VKTESSTV 63
A PVAK++ + K + + A KP+ +++ A K AA K +T +T
Sbjct: 683 AKPVAKAKPKAKAPVAKANAVPVAAKAKPAAKAKPAAKAKPAARPAKASRTSTRTSPGKR 862
Query: 64 SASEEVIQKKRNREPEVQDAAREAA--LREDATKREA 98
+A+ + KK + A++AA +++ A R+A
Sbjct: 863 AAAAKPAVKKAATPVKKAVPAKKAATPVKKAAPARKA 973
>TC87097 similar to GP|14334540|gb|AAK59678.1 putative protein {Arabidopsis
thaliana}, partial (62%)
Length = 1234
Score = 34.3 bits (77), Expect = 0.10
Identities = 35/168 (20%), Positives = 75/168 (43%), Gaps = 14/168 (8%)
Frame = +1
Query: 11 SRKTKSKALTKEDDLVEEATRKPSKRS---RKAEKETVAAPKPKKGKTVKTESSTVSASE 67
S+ + S + T+E + R+P+ R+ R+ + TVA+ T+ + S+
Sbjct: 136 SQSSSSSSSTEEAH--QAPVREPAARATATRRMRRRTVASGASSSSAPPATQQESGDESD 309
Query: 68 EVIQKKRNREPEVQDAAREAALREDATKREAALQAIRSK--RAKGERSIQERIREAAEQL 125
+ E E + + ++ R++ R A +A +SK R R +++ REA E+
Sbjct: 310 N---EAAGDEQEAKASKKKELKRQEREARRQAEEARQSKQDRYSEMRRLKDEEREAQERK 480
Query: 126 RKEEADPSLKKAQKPVSLESPCFR---------LTPEQEERTREIAAN 164
+EEA + ++ +LE ++ E++++T ++ N
Sbjct: 481 LEEEAKAQKAREEEAAALEFDKWKGEFSVDDEGTLEEEQDKTEDLLTN 624
>TC79972 similar to GP|8778288|gb|AAF79297.1| F14D16.30 {Arabidopsis
thaliana}, partial (46%)
Length = 772
Score = 34.3 bits (77), Expect = 0.10
Identities = 18/46 (39%), Positives = 29/46 (62%), Gaps = 1/46 (2%)
Frame = +1
Query: 242 SEATRPEAHPSGI-PSVPKASVNIQTPILPSSPSSSSSSSSTNSDD 286
+E +P ++P I ++P ++ I+ P L S SSS+SSSST +D
Sbjct: 139 TEINQPFSNPHSIHDAIPISTTEIENPTLYDSTSSSTSSSSTTDED 276
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.303 0.119 0.314
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,105,404
Number of Sequences: 36976
Number of extensions: 165652
Number of successful extensions: 2092
Number of sequences better than 10.0: 170
Number of HSP's better than 10.0 without gapping: 1465
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1818
length of query: 465
length of database: 9,014,727
effective HSP length: 100
effective length of query: 365
effective length of database: 5,317,127
effective search space: 1940751355
effective search space used: 1940751355
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 17 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146566.8


