

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146566.2 + phase: 2 /partial
(358 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
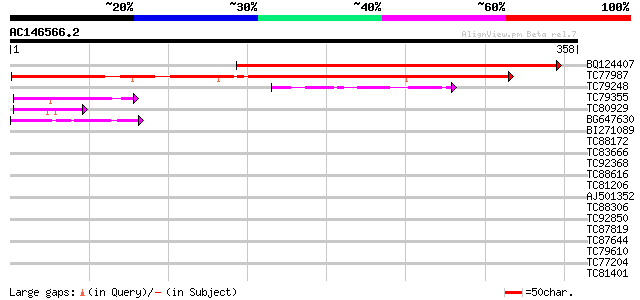
Score E
Sequences producing significant alignments: (bits) Value
BQ124407 similar to GP|17979277|gb unknown protein {Arabidopsis ... 430 e-121
TC77987 weakly similar to GP|17979277|gb|AAL49864.1 unknown prot... 332 1e-91
TC79248 similar to GP|22165059|gb|AAM93676.1 unknown protein {Or... 42 4e-04
TC79355 similar to GP|15081721|gb|AAK82515.1 At1g04990/F13M7_1 {... 42 4e-04
TC80929 similar to GP|10177307|dbj|BAB10568. contains similarity... 41 6e-04
BG647630 similar to GP|20521270|dbj hypothetical protein~similar... 41 8e-04
BI271089 similar to GP|9294325|dbj| gene_id:K24M9.13~unknown pro... 40 0.002
TC88172 similar to GP|12003386|gb|AAG43550.1 Avr9/Cf-9 rapidly e... 39 0.002
TC83666 similar to GP|15042832|gb|AAK82455.1 putative RNA bindin... 39 0.003
TC92368 weakly similar to GP|19347974|gb|AAL86319.1 unknown prot... 39 0.003
TC88616 weakly similar to PIR|T48209|T48209 hypothetical protein... 39 0.004
TC81206 similar to GP|8777433|dbj|BAA97023.1 gene_id:MHM17.4~unk... 39 0.004
AJ501352 similar to PIR|B84584|B845 probable RING zinc finger pr... 38 0.007
TC88306 similar to GP|21593124|gb|AAM65073.1 unknown {Arabidopsi... 37 0.009
TC92850 similar to GP|15529216|gb|AAK97702.1 At1g72220/T9N14_22 ... 37 0.009
TC87819 similar to GP|16209696|gb|AAL14405.1 AT5g20880/F22D1_50 ... 37 0.009
TC87644 similar to GP|10177307|dbj|BAB10568. contains similarity... 37 0.009
TC79610 similar to GP|16323294|gb|AAL15402.1 AT5g19430/F7K24_180... 37 0.009
TC77204 similar to GP|21536625|gb|AAM60957.1 RING-H2 zinc finger... 37 0.009
TC81401 weakly similar to GP|21593762|gb|AAM65729.1 unknown {Ara... 37 0.012
>BQ124407 similar to GP|17979277|gb unknown protein {Arabidopsis thaliana},
partial (62%)
Length = 635
Score = 430 bits (1106), Expect = e-121
Identities = 200/205 (97%), Positives = 201/205 (97%)
Frame = +2
Query: 144 NCPRGEQCPHVHGDLCPSCGRQCLHPFRPEEREEHMMSCRNKQKHLEALKRSQEIECSVC 203
NCPRGEQCPHVHGDLCPSCGRQCLHPFRPEEREEHMMSCRNKQKHLEALKRSQEIECSVC
Sbjct: 2 NCPRGEQCPHVHGDLCPSCGRQCLHPFRPEEREEHMMSCRNKQKHLEALKRSQEIECSVC 181
Query: 204 LERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYF 263
LERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYF
Sbjct: 182 LERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYF 361
Query: 264 VVPSVIWYATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEV 323
VVPSVIWYATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEV
Sbjct: 362 VVPSVIWYATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEV 541
Query: 324 ALRHLGAADGDTIIAKDIRFLSTLS 348
ALRHLGAADGDTIIAKDIR L+
Sbjct: 542 ALRHLGAADGDTIIAKDIRLSDFLA 616
>TC77987 weakly similar to GP|17979277|gb|AAL49864.1 unknown protein
{Arabidopsis thaliana}, partial (53%)
Length = 1338
Score = 332 bits (851), Expect = 1e-91
Identities = 170/324 (52%), Positives = 209/324 (64%), Gaps = 7/324 (2%)
Frame = +3
Query: 2 LCKFFAHGACLKGEHCEFSHDWKAPPNNICTFYQKGVCAYGSRCRYDHVKASRAQSSTPS 61
+CKF+A G CLKG+ C+FSH K ++IC++YQKG CAYGSRCRY HV+AS+A SS
Sbjct: 93 VCKFYARGVCLKGDQCDFSHQRKDTASDICSYYQKGSCAYGSRCRYKHVRASQASSSAS- 269
Query: 62 SSITEHQPLVSESAV--LGNTRVTSNGVATAAEFSLFSTPFVLPSEQAWNQESAQLDFLR 119
+VS+SAV + +V SN V + +PS + Q
Sbjct: 270 --------MVSDSAVPVARSAKVASNWVPKVTK---------VPSPDKRGVKGLQRKHQD 398
Query: 120 EDDVVQSVITS--PSELPICSFAAAGNCPRGEQCPHVHGDLCPSCGRQCLHPFRPEEREE 177
DV +S S P E C FAAA NCP G C +HG+ C C + CLHP +E+E
Sbjct: 399 STDVGESSTGSARPHENLFCKFAAA-NCPGG--CSRIHGNQCLYCRKYCLHPTDKKEKEN 569
Query: 178 HMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWR 237
H+ +C K+K+L ALK S+EIEC+VCLERVLSKP +E KFGLL ECDH FC+SCIRNWR
Sbjct: 570 HLRTCDKKEKYLLALKNSEEIECNVCLERVLSKPKPSECKFGLLPECDHAFCLSCIRNWR 749
Query: 238 SSNPTLGMDVNS---TLRACPICRKLSYFVVPSVIWYATSEEKMEIIDTYKAKLKSIDCK 294
SS PT M++ S T+R CP+CRKLSYFV+PS IW+ T EEK EIID YKA + IDCK
Sbjct: 750 SSAPTSAMEIGSNTNTVRTCPVCRKLSYFVIPSGIWFTTKEEKQEIIDNYKANCRLIDCK 929
Query: 295 HFDFGEGNCPFGTSCFYKHAYRDG 318
HFD G GNCPFG SCFYKH + G
Sbjct: 930 HFDSGNGNCPFGASCFYKHTVKPG 1001
>TC79248 similar to GP|22165059|gb|AAM93676.1 unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (97%)
Length = 1272
Score = 42.0 bits (97), Expect = 4e-04
Identities = 30/117 (25%), Positives = 48/117 (40%)
Frame = +2
Query: 166 CLHPFRPEEREEHMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECD 225
C+ +R E EEH K + +E EC +C+E + SK +L C+
Sbjct: 482 CMERYRRREDEEH--------KQFSDIDFEREEECGICME-MNSKI--------VLPNCN 610
Query: 226 HPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIID 282
H C+ C WR+ + ++CP CR V +W T + +I+D
Sbjct: 611 HVMCLKCYHEWRARS-----------QSCPFCRDSLKRVNSGDLWIFT--DSRDIVD 742
>TC79355 similar to GP|15081721|gb|AAK82515.1 At1g04990/F13M7_1 {Arabidopsis
thaliana}, partial (42%)
Length = 1627
Score = 42.0 bits (97), Expect = 4e-04
Identities = 28/97 (28%), Positives = 48/97 (48%), Gaps = 18/97 (18%)
Frame = +2
Query: 3 CKFF-AHGACLKGEHCEFSHDWK---------------APPNNICTFYQ-KGVCAYGSRC 45
CK+F + G C G C+F H + P N IC++Y+ GVC +G C
Sbjct: 1022 CKYFMSTGTCKYGSDCKFHHPKERIAQTLSINPLGLPMRPGNAICSYYRIYGVCKFGPTC 1201
Query: 46 RYDHVKASRAQS-STPSSSITEHQPLVSESAVLGNTR 81
++DH + +Q+ PS +++ V ++++L N R
Sbjct: 1202 KFDHPVVAISQNYGLPSPTLS-----VFDASLLTNPR 1297
Score = 32.7 bits (73), Expect = 0.22
Identities = 18/60 (30%), Positives = 24/60 (40%), Gaps = 14/60 (23%)
Frame = +1
Query: 9 GACLKGEHCEFSHDWKAPP-------------NNICTFYQK-GVCAYGSRCRYDHVKASR 54
G C G +C ++H P C ++ K G C YGS C+Y H K R
Sbjct: 364 GMCGYGSNCRYNHPANISPVTQYGEELPERVGQPDCEYFLKTGTCKYGSTCKYHHPKDRR 543
Score = 31.6 bits (70), Expect = 0.49
Identities = 11/20 (55%), Positives = 15/20 (75%), Gaps = 1/20 (5%)
Frame = +1
Query: 31 CTFYQK-GVCAYGSRCRYDH 49
C +Y + G+C YGS CRY+H
Sbjct: 343 CVYYLRTGMCGYGSNCRYNH 402
Score = 28.9 bits (63), Expect = 3.2
Identities = 30/108 (27%), Positives = 43/108 (39%), Gaps = 7/108 (6%)
Frame = +2
Query: 33 FYQKGVCAYGSRCRYDHVKASRAQSSTPSSSITEHQPLVSESAVLGNTRVTSNGVATAAE 92
F G C YGS C++ H K AQ+ + + P+ +A+ R+ GV
Sbjct: 1031 FMSTGTCKYGSDCKFHHPKERIAQTLSINPL---GLPMRPGNAICSYYRI--YGVCKFGP 1195
Query: 93 FSLFSTPFV-------LPSEQAWNQESAQLDFLREDDVVQSVITSPSE 133
F P V LPS +++ L R VQ TSPS+
Sbjct: 1196 TCKFDHPVVAISQNYGLPSPTLSVFDASLLTNPRRLSTVQPAETSPSK 1339
>TC80929 similar to GP|10177307|dbj|BAB10568. contains similarity to zinc
finger protein~gene_id:MDC12.23 {Arabidopsis thaliana},
partial (28%)
Length = 1274
Score = 41.2 bits (95), Expect = 6e-04
Identities = 23/66 (34%), Positives = 29/66 (43%), Gaps = 19/66 (28%)
Frame = +3
Query: 3 CKFFAH-GACLKGEHCEFSHD----WKAPP-------------NNICTFYQK-GVCAYGS 43
C FF G C HC+F H K PP N+CT Y + G+C +G
Sbjct: 630 CSFFLKTGDCKFKSHCKFHHPKNRITKLPPCNLSDKGLPLRPGQNVCTHYSRYGICKFGP 809
Query: 44 RCRYDH 49
C+YDH
Sbjct: 810 ACKYDH 827
Score = 27.3 bits (59), Expect = 9.3
Identities = 15/68 (22%), Positives = 28/68 (41%), Gaps = 21/68 (30%)
Frame = +2
Query: 3 CKFFAH-GACLKGEHCEFSHDWKAPPNNICTF--------------------YQKGVCAY 41
CK+++ G C G+ C+F H + + I F + G C +
Sbjct: 68 CKYYSRSGGCKFGKDCKFDHTRRKIFSEIEVFRA*LPWPTYSFG*ERMPVITMRTGSCKF 247
Query: 42 GSRCRYDH 49
G+ C+++H
Sbjct: 248 GANCKFNH 271
>BG647630 similar to GP|20521270|dbj hypothetical protein~similar to
Arabidopsis thaliana chromosome 5 At5g56930, partial
(7%)
Length = 699
Score = 40.8 bits (94), Expect = 8e-04
Identities = 24/84 (28%), Positives = 37/84 (43%)
Frame = +2
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNNICTFYQKGVCAYGSRCRYDHVKASRAQSSTP 60
V C +A G+C+KG C + H P F KG C Y RC + H + TP
Sbjct: 152 VPCAHYACGSCMKGNDCPYDHQLSKYP--CSNFVSKGSC-YRGRCMFSHQVQTSQDIPTP 322
Query: 61 SSSITEHQPLVSESAVLGNTRVTS 84
+++ +P + GNT ++
Sbjct: 323 TNAC---KPELKSPLSSGNTNFST 385
Score = 35.8 bits (81), Expect = 0.026
Identities = 16/38 (42%), Positives = 20/38 (52%), Gaps = 1/38 (2%)
Frame = +2
Query: 13 KGEHCEFSHDWKAPPNNI-CTFYQKGVCAYGSRCRYDH 49
KG+ C FSHD ++ C Y G C G+ C YDH
Sbjct: 101 KGDQCNFSHDAIPLTKSVPCAHYACGSCMKGNDCPYDH 214
>BI271089 similar to GP|9294325|dbj| gene_id:K24M9.13~unknown protein
{Arabidopsis thaliana}, partial (2%)
Length = 481
Score = 39.7 bits (91), Expect = 0.002
Identities = 15/23 (65%), Positives = 17/23 (73%)
Frame = +3
Query: 3 CKFFAHGACLKGEHCEFSHDWKA 25
CKFFA G C G+HC FSHD +A
Sbjct: 171 CKFFASGNCRNGKHCRFSHDTQA 239
Score = 28.5 bits (62), Expect = 4.2
Identities = 12/35 (34%), Positives = 15/35 (42%), Gaps = 4/35 (11%)
Frame = +3
Query: 19 FSHDWKAPPNNI----CTFYQKGVCAYGSRCRYDH 49
F H + PN C F+ G C G CR+ H
Sbjct: 123 FEHGGRHVPNRTIDAPCKFFASGNCRNGKHCRFSH 227
>TC88172 similar to GP|12003386|gb|AAG43550.1 Avr9/Cf-9 rapidly elicited
protein 132 {Nicotiana tabacum}, partial (26%)
Length = 1445
Score = 39.3 bits (90), Expect = 0.002
Identities = 21/62 (33%), Positives = 30/62 (47%)
Frame = +3
Query: 197 EIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPI 256
++ECSVCL V+ K +L +C+H F + CI W S+ T CP+
Sbjct: 483 DLECSVCLSEVVEG-----EKVRVLPKCNHMFHIDCIDMWFHSHST-----------CPL 614
Query: 257 CR 258
CR
Sbjct: 615 CR 620
>TC83666 similar to GP|15042832|gb|AAK82455.1 putative RNA binding protein
{Oryza sativa}, partial (52%)
Length = 896
Score = 38.9 bits (89), Expect = 0.003
Identities = 24/78 (30%), Positives = 33/78 (41%)
Frame = +1
Query: 2 LCKFFAHGACLKGEHCEFSHDWKAPPNNICTFYQKGVCAYGSRCRYDHVKASRAQSSTPS 61
+C+ F G C +G C+FSHD + N T + A G +YD K R +
Sbjct: 565 VCRAFQRGECTRGAGCKFSHDEQRAAN---TGWGDNGIAKGDNDKYDGPKKERRYGNNQP 735
Query: 62 SSITEHQPLVSESAVLGN 79
I E + S S GN
Sbjct: 736 DRIPETRDRDSRSRAHGN 789
Score = 28.1 bits (61), Expect = 5.5
Identities = 10/43 (23%), Positives = 20/43 (46%)
Frame = +1
Query: 24 KAPPNNICTFYQKGVCAYGSRCRYDHVKASRAQSSTPSSSITE 66
K +C +Q+G C G+ C++ H + A + + I +
Sbjct: 547 KREARGVCRAFQRGECTRGAGCKFSHDEQRAANTGWGDNGIAK 675
>TC92368 weakly similar to GP|19347974|gb|AAL86319.1 unknown protein
{Arabidopsis thaliana}, partial (63%)
Length = 999
Score = 38.9 bits (89), Expect = 0.003
Identities = 22/53 (41%), Positives = 27/53 (50%), Gaps = 5/53 (9%)
Frame = +3
Query: 2 LC-KFFAHGACLKGEHCEFSHDWKAPPN---NIC-TFYQKGVCAYGSRCRYDH 49
LC KF + G+C GE C F HD A + +C F KG C G CR+ H
Sbjct: 90 LCFKFVSSGSCQWGETCNFRHDTDAREHCLRGVCFDFLNKGKCERGPDCRFRH 248
>TC88616 weakly similar to PIR|T48209|T48209 hypothetical protein T20L15.150
- Arabidopsis thaliana, partial (27%)
Length = 1148
Score = 38.5 bits (88), Expect = 0.004
Identities = 27/76 (35%), Positives = 32/76 (41%), Gaps = 2/76 (2%)
Frame = +2
Query: 185 KQKHLEALKRSQE--IECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPT 242
KQK E K + +EC+VCL V E LL C H F V CI W +S+ T
Sbjct: 452 KQKQQEENKNVSKNIVECAVCLSVVED-----EEMMRLLPNCKHSFHVGCIDKWLASHST 616
Query: 243 LGMDVNSTLRACPICR 258
CP CR
Sbjct: 617 -----------CPNCR 631
>TC81206 similar to GP|8777433|dbj|BAA97023.1 gene_id:MHM17.4~unknown
protein {Arabidopsis thaliana}, partial (5%)
Length = 842
Score = 38.5 bits (88), Expect = 0.004
Identities = 20/52 (38%), Positives = 26/52 (49%), Gaps = 5/52 (9%)
Frame = +3
Query: 3 CKFFAHGACLKGEHCEFSHD----WKAPPNNICTFY-QKGVCAYGSRCRYDH 49
C+ + +G C KG+ C FSHD K+ P C Y C G+ C YDH
Sbjct: 117 CRHYMNGRCHKGDECNFSHDAIPLTKSKP---CVHYAHHRSCMKGNDCPYDH 263
Score = 28.5 bits (62), Expect = 4.2
Identities = 23/89 (25%), Positives = 37/89 (40%), Gaps = 9/89 (10%)
Frame = +3
Query: 5 FFAHGACLKGEHCEFSHDWKAPPN-----NICTFYQKGVCAYG----SRCRYDHVKASRA 55
F + G+C +G+ C FSH + + N C K + G S +H S
Sbjct: 291 FVSKGSCYRGDACLFSHQVQTSQDIPTATNACKPELKSSLSSGNTNFSTLLNNHGTGSVQ 470
Query: 56 QSSTPSSSITEHQPLVSESAVLGNTRVTS 84
QS++ S I H + + + T+ TS
Sbjct: 471 QSNSNSKGIYSHTNVEHKVTDVSKTKPTS 557
>AJ501352 similar to PIR|B84584|B845 probable RING zinc finger protein
[imported] - Arabidopsis thaliana, partial (17%)
Length = 607
Score = 37.7 bits (86), Expect = 0.007
Identities = 21/63 (33%), Positives = 29/63 (45%)
Frame = +3
Query: 196 QEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACP 255
Q +EC+VCL + E LL +C H F ++CI W S+ T CP
Sbjct: 399 QGLECTVCLSKFED-----EEILRLLPKCKHAFHMNCIDKWLESHST-----------CP 530
Query: 256 ICR 258
+CR
Sbjct: 531 LCR 539
>TC88306 similar to GP|21593124|gb|AAM65073.1 unknown {Arabidopsis
thaliana}, partial (80%)
Length = 1304
Score = 37.4 bits (85), Expect = 0.009
Identities = 22/87 (25%), Positives = 34/87 (38%)
Frame = +2
Query: 196 QEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACP 255
+E EC +CLE +L C H C+ C R W N+ +CP
Sbjct: 740 REDECGICLEPCTKM---------VLPNCCHAMCIKCYRKW-----------NTKSESCP 859
Query: 256 ICRKLSYFVVPSVIWYATSEEKMEIID 282
CR V +W T ++ +++D
Sbjct: 860 FCRGSIRRVNSEDLWVLTCDD--DVVD 934
>TC92850 similar to GP|15529216|gb|AAK97702.1 At1g72220/T9N14_22
{Arabidopsis thaliana}, partial (19%)
Length = 687
Score = 37.4 bits (85), Expect = 0.009
Identities = 25/74 (33%), Positives = 31/74 (41%), Gaps = 2/74 (2%)
Frame = +3
Query: 199 ECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
ECSVCL T LL +C H F +SCI W S+ CP+CR
Sbjct: 480 ECSVCLNEFHEDETLR-----LLPKCSHAFHISCIDTWLRSHTN-----------CPLCR 611
Query: 259 K--LSYFVVPSVIW 270
+S V P V +
Sbjct: 612 AGIVSNNVTPEVTY 653
>TC87819 similar to GP|16209696|gb|AAL14405.1 AT5g20880/F22D1_50
{Arabidopsis thaliana}, partial (28%)
Length = 1121
Score = 37.4 bits (85), Expect = 0.009
Identities = 31/116 (26%), Positives = 45/116 (38%), Gaps = 10/116 (8%)
Frame = +1
Query: 171 RPEEREEHMMSCRN----------KQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGL 220
+P E ++ SC K++H E E C+VCL ++ + E
Sbjct: 433 KPVEDQQQYSSCCQMLPLTSFGEIKERHPET-----EETCAVCLNKLKMEDEVRE----- 582
Query: 221 LSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEE 276
L CDH F CI W G D + + CP+CR P + Y+ S E
Sbjct: 583 LMNCDHVFHKECIDKWLEH----GHDNENHNQTCPLCR------APLINSYSLSSE 720
>TC87644 similar to GP|10177307|dbj|BAB10568. contains similarity to zinc
finger protein~gene_id:MDC12.23 {Arabidopsis thaliana},
partial (36%)
Length = 1541
Score = 37.4 bits (85), Expect = 0.009
Identities = 21/77 (27%), Positives = 36/77 (46%), Gaps = 17/77 (22%)
Frame = +1
Query: 3 CKFFAH-GACLKGEHCEFSHD--WKAPPNNI-------------CTFYQK-GVCAYGSRC 45
CK++ G C G+ C+++H + AP + + C +Y + G C +GS C
Sbjct: 424 CKYYQRSGGCKFGKACKYNHSRGFTAPISELNFLGLPIRLGERECPYYMRTGSCKFGSNC 603
Query: 46 RYDHVKASRAQSSTPSS 62
R++H + S P S
Sbjct: 604 RFNHPDPTTVGGSDPQS 654
Score = 35.4 bits (80), Expect = 0.034
Identities = 21/70 (30%), Positives = 32/70 (45%), Gaps = 19/70 (27%)
Frame = +1
Query: 3 CKFFAH-GACLKGEHCEFSHDW----KAPPNN-------------ICTFYQK-GVCAYGS 43
C FF G C +C+F H K PP N +C+ Y + G+C +G
Sbjct: 958 CSFFIKTGDCKFKSNCKFHHPKNRVAKLPPCNLSDKGLPLRPDQSVCSHYSRYGICKFGP 1137
Query: 44 RCRYDHVKAS 53
CR+DH +++
Sbjct: 1138 ACRFDHPESA 1167
>TC79610 similar to GP|16323294|gb|AAL15402.1 AT5g19430/F7K24_180
{Arabidopsis thaliana}, partial (54%)
Length = 789
Score = 37.4 bits (85), Expect = 0.009
Identities = 16/59 (27%), Positives = 31/59 (52%)
Frame = +1
Query: 200 CSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
C++CL+ ++ + TA L+ C+H +CV+CI +W + + + CP C+
Sbjct: 253 CAICLDNIVLQETA------LVKGCEHAYCVTCILHWATYSQKV---------TCPQCK 384
>TC77204 similar to GP|21536625|gb|AAM60957.1 RING-H2 zinc finger
protein-like {Arabidopsis thaliana}, partial (30%)
Length = 1218
Score = 37.4 bits (85), Expect = 0.009
Identities = 21/61 (34%), Positives = 27/61 (43%)
Frame = +3
Query: 198 IECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPIC 257
+EC+VCL T L+ +CDH F CI W SS+ T CP+C
Sbjct: 396 LECAVCLMEFEDTETLR-----LIPKCDHVFHPECIDEWLSSHTT-----------CPVC 527
Query: 258 R 258
R
Sbjct: 528 R 530
>TC81401 weakly similar to GP|21593762|gb|AAM65729.1 unknown {Arabidopsis
thaliana}, partial (30%)
Length = 768
Score = 37.0 bits (84), Expect = 0.012
Identities = 19/60 (31%), Positives = 28/60 (46%)
Frame = +2
Query: 199 ECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
EC+VCL+ ++ + A LL C+H F + C W S +P CP+CR
Sbjct: 290 ECAVCLDDIVDEQPAR-----LLPGCNHAFHLQCADTWLSKHP-----------MCPLCR 421
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,075,384
Number of Sequences: 36976
Number of extensions: 267904
Number of successful extensions: 1443
Number of sequences better than 10.0: 130
Number of HSP's better than 10.0 without gapping: 1350
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1426
length of query: 358
length of database: 9,014,727
effective HSP length: 97
effective length of query: 261
effective length of database: 5,428,055
effective search space: 1416722355
effective search space used: 1416722355
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146566.2


