

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146565.8 - phase: 0 /pseudo
(1145 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
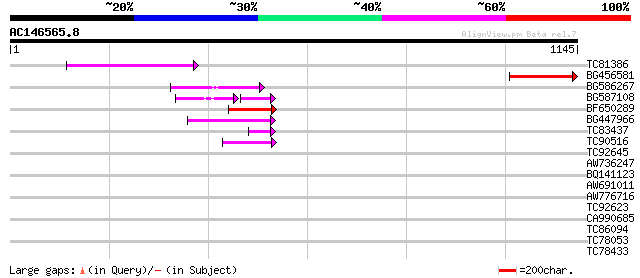
Score E
Sequences producing significant alignments: (bits) Value
TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR re... 223 3e-58
BG456581 217 2e-56
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 92 1e-18
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 59 1e-18
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 79 1e-14
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 78 2e-14
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 48 2e-05
TC90516 47 4e-05
TC92645 35 0.13
AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase... 35 0.13
BQ141123 homologue to GP|14488272|db Na+/H+ exchanger protein {I... 31 2.4
AW691011 similar to PIR|T01804|T018 Na+/H+-exchanging protein 3 ... 31 2.4
AW776716 similar to SP|O22224|Y240 Protein At2g41620. [Mouse-ear... 31 2.4
TC92623 31 3.1
CA990685 weakly similar to GP|22136772|gb putative carbonyl redu... 30 4.1
TC86094 similar to GP|18700188|gb|AAL77705.1 AT4g18710/F28A21_12... 30 4.1
TC78053 similar to GP|5923675|gb|AAD56326.1| putative mRNA cappi... 30 6.9
TC78433 similar to GP|3790100|gb|AAC67586.1| pyrophosphate-depen... 29 9.1
>TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 798
Score = 223 bits (568), Expect = 3e-58
Identities = 116/265 (43%), Positives = 158/265 (58%)
Frame = -1
Query: 116 RRLLWADLTHLQGCFLGPWLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHL 175
RR LW+ ++++Q W IGDFN ++G+HE +G P DF WS+ N L HL
Sbjct: 795 RRQLWSAISNIQTQHKLSWCCIGDFNTILGSHEHQGSHTPARLPMLDFQQWSDVNNLFHL 616
Query: 176 PSSGPLLTWSNGRLGSAYVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADV 235
P+ G TW+NGR G RLDRSI N+ + S CTL + +SDH P+L
Sbjct: 615 PTRGSAFTWTNGRRGRNNTRKRLDRSIVNQLMIDNCESLSACTLTKLRSDHFPILFELQT 436
Query: 236 ASVKNAISFKFYKAWTSHVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVF 295
+++ + SFKF K W++H DC ++ + WA V G M L KL+ LK V K WN++ F
Sbjct: 435 QNIQFSSSFKFMKMWSAHPDCINIIKQCWANQVVGCPMFVLNQKLKNLKEVLKVWNKNTF 256
Query: 296 GDVDRQVRLATDEVSRIQQIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQS 355
G+V QV A E+ IQ IDS+G+SD L Q+ AQL L +ALN+++ FW EKS+
Sbjct: 255 GNVHSQVDNAYKELDDIQVKIDSIGYSDVLMDQEKAAQLNLESALNIEEVFWHEKSKVNW 76
Query: 356 FIHGDRNTAYFHRVARIKASSKNIS 380
GDRNTA+FHRVA+IK +S IS
Sbjct: 75 HCEGDRNTAFFHRVAKIKRTSSLIS 1
>BG456581
Length = 683
Score = 217 bits (553), Expect = 2e-56
Identities = 110/136 (80%), Positives = 117/136 (85%)
Frame = +1
Query: 1010 EVNCQAGLLFSHVAHSLCSMGVNYLEVFHSSISFFYFLEACS**ITHG*QPFFTWVCIGF 1069
+VN QAGLLFSHVA+ LCS+G+NYLE H S SFFY LEA S**ITHG* P FTWVC GF
Sbjct: 10 KVNYQAGLLFSHVAYFLCSVGLNYLEFLHPSFSFFYLLEAGS**ITHG**PLFTWVCFGF 189
Query: 1070 DM*PLFGASRNF*SFVSSL*VCGGALELALFTAAHWAGFLLH*GAAVLSPTPLFFSSGGL 1129
D+* LFGASRNF SFV SL*VCG ALELALF+AA WAGFLL *G AVLSPTPLFFSS GL
Sbjct: 190 DV*LLFGASRNFGSFVPSL*VCGDALELALFSAARWAGFLLL*GVAVLSPTPLFFSSAGL 369
Query: 1130 VCCCGCAYGSFYLLGS 1145
VCCCGCAYGSFYL+GS
Sbjct: 370 VCCCGCAYGSFYLVGS 417
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 92.0 bits (227), Expect = 1e-18
Identities = 65/197 (32%), Positives = 92/197 (45%), Gaps = 8/197 (4%)
Frame = +3
Query: 325 LYRQDLEAQLL---LTNALNVQDQFWKEKSRHQSFIHGDRNTAYFHRVARIKASSKNISL 381
L+ + E LL L + ++ FW +KSR GDRNT +FH V + + + I
Sbjct: 60 LFNKGEELSLLRSELNEEYHNEEIFWMQKSRLNWLRSGDRNTKFFHAVTKNRRAQNRILS 239
Query: 382 LYDGAT----VLSDAGDIAAHILNYFQGIFDVVNNCIPNDLVARSIHALVTEDENNMLIR 437
L D V D G +A DV D SI A+VTE++N L+
Sbjct: 240 LIDDDDKEWFVEEDLGRLADSHFKLLYSSEDV--GITLEDW--NSIPAIVTEEQNAQLMA 407
Query: 438 LPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNIN- 496
R+E++ AVFD+N PGPDG F+Q FWD +G D+ Q+F G + IN
Sbjct: 408 QISREEVREAVFDINPHKCPGPDGMNVFFFQQFWDTMGDDLTSMAQEFLRTGKLEEGINK 587
Query: 497 SNLLVLIPKIPGAQAMG 513
+N+ K G +A G
Sbjct: 588 TNIWFGSKKAGGKEASG 638
Score = 40.0 bits (92), Expect = 0.005
Identities = 20/52 (38%), Positives = 31/52 (59%)
Frame = +1
Query: 488 QGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRLASI 539
QG+ L+PK A+ + +FRPI+L N +KIV+K+L+ RL S+
Sbjct: 559 QGNLKRESTKQTSGLVPKKLEAKRLVEFRPISLCNVAYKIVSKVLSKRLKSV 714
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 59.3 bits (142), Expect(2) = 1e-18
Identities = 37/130 (28%), Positives = 66/130 (50%), Gaps = 4/130 (3%)
Frame = +2
Query: 336 LTNALNVQDQFWKEKSRHQSFIHGDRNTAYFHRVARIKASSKNISLL--YDGATVLSDAG 393
L A ++ +W++KSR+ I GD N ++H + + + + I L YDG + + G
Sbjct: 20 LQEAYKDEEDYWQQKSRNMWHISGDLNKKFYHALTKQRHARNRIVGLYDYDGNWITEEQG 199
Query: 394 DIAAHILNYFQGIFDVVNNCIPN--DLVARSIHALVTEDENNMLIRLPLRDEIKSAVFDL 451
+ ++YF+ +F P D I + +T N L+RL +E++ A+F +
Sbjct: 200 -VEKVAVDYFEDLF---QRTTPTGFDGFLDEITSSITPQMNQRLLRLATEEEVRLALFIM 367
Query: 452 NGDGAPGPDG 461
+ + APGPDG
Sbjct: 368 HPEKAPGPDG 397
Score = 53.1 bits (126), Expect(2) = 1e-18
Identities = 25/71 (35%), Positives = 38/71 (53%)
Frame = +3
Query: 466 FYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQF 525
F+Q W I+ D++ V F G +N+ + LIPK M + RPI+L N +
Sbjct: 411 FFQHSWHIIKMDLLKMVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVGY 590
Query: 526 KIVTKILADRL 536
KI++K+L RL
Sbjct: 591 KIISKVLCQRL 623
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 79.0 bits (193), Expect = 1e-14
Identities = 35/97 (36%), Positives = 59/97 (60%)
Frame = +3
Query: 443 EIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVL 502
E+K+A+F ++ APG DG+ HF++ W+I+G V+ ++ DFF G IN + L
Sbjct: 36 EVKNALFSMDSSKAPGIDGYNVHFFKCSWNIIGDSVIDAILDFFKTGFMPKIINCTYVTL 215
Query: 503 IPKIPGAQAMGDFRPIALANFQFKIVTKILADRLASI 539
+PK ++ +FRPIA + +KI++KIL R+ +
Sbjct: 216 LPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGV 326
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 77.8 bits (190), Expect = 2e-14
Identities = 48/179 (26%), Positives = 80/179 (43%), Gaps = 1/179 (0%)
Frame = +2
Query: 359 GDRNTAYFHRVARIKASSKNISLLYDGA-TVLSDAGDIAAHILNYFQGIFDVVNNCIPND 417
GD+NT +FH A + I L D ++ ++ YF +F N +
Sbjct: 17 GDKNTKFFHSKASQRRKVNEIKKLKDETGNWCKGEENVERLLITYFNNLFTSSNPTAIEE 196
Query: 418 LVARSIHALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSD 477
+ ++ + + +E+ A+ ++ APGPDG F+Q +W IVG +
Sbjct: 197 -TCEVVKGKLSHEHIVWCEKEFTEEEVLEAINQMHPVKAPGPDGLPALFFQKYWHIVGKE 373
Query: 478 VVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRL 536
V V + +N +VLIPK D+RPI+L N KI+TK++A+R+
Sbjct: 374 VQQMVLQVLNNSMETEELNKTFIVLIPKGKNPNTPKDYRPISLCNVVMKIITKVIANRV 550
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 47.8 bits (112), Expect = 2e-05
Identities = 23/55 (41%), Positives = 32/55 (57%)
Frame = +2
Query: 482 VQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRL 536
V +F INS + LIPK+ Q + DFRPI+L +KI+ K+LA+RL
Sbjct: 11 VSEFHRNRKLFKGINSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRL 175
>TC90516
Length = 983
Score = 47.0 bits (110), Expect = 4e-05
Identities = 29/109 (26%), Positives = 56/109 (50%)
Frame = -2
Query: 431 ENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGS 490
+N +L L +++ V L+G+ +P PDGF +F+ +++ +D+ + + F+ +
Sbjct: 544 DNVLLSAQFLVSKMELVVSSLDGNESPRPDGFNLNFFIRLRNMLKADIEIMFEQFYTPAN 365
Query: 491 FADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRLASI 539
+ L LI K+ +GDF ++ K++ K+LA RLA I
Sbjct: 364 LLKIFS*YFLTLISKVEYPILLGDFSLMSFLGSL*KLMAKVLALRLAHI 218
>TC92645
Length = 561
Score = 35.4 bits (80), Expect = 0.13
Identities = 19/49 (38%), Positives = 28/49 (56%), Gaps = 1/49 (2%)
Frame = +3
Query: 430 DENNMLIRLPLR-DEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSD 477
D+NN ++ P DEI +A+F ++ D A G DG Y F ++VG D
Sbjct: 288 DQNNCMLTAPFSVDEIWTAIFQMHHDKAFGLDGLNLAVYYRF*NLVGGD 434
>AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (4%)
Length = 305
Score = 35.4 bits (80), Expect = 0.13
Identities = 15/50 (30%), Positives = 28/50 (56%), Gaps = 2/50 (4%)
Frame = +1
Query: 442 DEIKSAVFDLNGDGA--PGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQG 489
+E++ AV+D + + PGPDG F + +W+++ D + V +F G
Sbjct: 130 EEVRKAVWDCDSSHS*NPGPDGVNFTFVKEYWELIKVDFLRVVMEFHTNG 279
>BQ141123 homologue to GP|14488272|db Na+/H+ exchanger protein {Ipomoea nil},
partial (18%)
Length = 655
Score = 31.2 bits (69), Expect = 2.4
Identities = 14/40 (35%), Positives = 21/40 (52%)
Frame = +2
Query: 603 FFFCFLQLDFGNFTLGQTFYFGEWQGAGVLLLCPWCSSRG 642
+F+CF ++ +F ++ G V L CPW SSRG
Sbjct: 326 YFYCFKTINVIHFRSCFCCFYELVCGTSVCLYCPWSSSRG 445
>AW691011 similar to PIR|T01804|T018 Na+/H+-exchanging protein 3 homolog
A_TM021B04.4 - Arabidopsis thaliana, partial (21%)
Length = 654
Score = 31.2 bits (69), Expect = 2.4
Identities = 14/40 (35%), Positives = 21/40 (52%)
Frame = +2
Query: 603 FFFCFLQLDFGNFTLGQTFYFGEWQGAGVLLLCPWCSSRG 642
+F+CF ++ +F ++ G V L CPW SSRG
Sbjct: 320 YFYCFKTINVIHFRSCFCCFYELVCGTSVCLYCPWSSSRG 439
>AW776716 similar to SP|O22224|Y240 Protein At2g41620. [Mouse-ear cress]
{Arabidopsis thaliana}, partial (23%)
Length = 611
Score = 31.2 bits (69), Expect = 2.4
Identities = 13/36 (36%), Positives = 22/36 (61%), Gaps = 1/36 (2%)
Frame = -3
Query: 27 LSNKPTFIFLAEPMISFSYVP-SWYWHSIGVKKCCL 61
+ +PT +++ I SY+P S +WH+ +K CCL
Sbjct: 351 IEKEPTLFWIS---IDLSYIPDS*HWHAASIKNCCL 253
>TC92623
Length = 549
Score = 30.8 bits (68), Expect = 3.1
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = +3
Query: 1118 SPTPLFFSSGGLVCCCGCAYGSFYLLG 1144
S + +S VCCC C+YGS L+G
Sbjct: 18 SKVSIHYSHFFCVCCCSCSYGSHNLVG 98
>CA990685 weakly similar to GP|22136772|gb putative carbonyl reductase
{Arabidopsis thaliana}, partial (36%)
Length = 554
Score = 30.4 bits (67), Expect = 4.1
Identities = 13/27 (48%), Positives = 15/27 (55%)
Frame = +2
Query: 283 LKTVFKQWNRSVFGDVDRQVRLATDEV 309
LK + +W R VFGDVD DEV
Sbjct: 263 LKRIKNEWTREVFGDVDNLTEEKVDEV 343
>TC86094 similar to GP|18700188|gb|AAL77705.1 AT4g18710/F28A21_120
{Arabidopsis thaliana}, partial (49%)
Length = 1160
Score = 30.4 bits (67), Expect = 4.1
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = +3
Query: 1013 CQAGLLFSHVAHSLCSMGVNYLEVFHSSISFFYFLEAC 1050
CQ L + + +CS + YL VFH I F+Y L C
Sbjct: 1047 CQTSYLPCQM*YDICSF*ICYLLVFH--ICFYYLLNVC 1154
>TC78053 similar to GP|5923675|gb|AAD56326.1| putative mRNA capping enzyme
RNA guanylyltransferase {Arabidopsis thaliana}, partial
(71%)
Length = 2017
Score = 29.6 bits (65), Expect = 6.9
Identities = 21/78 (26%), Positives = 36/78 (45%), Gaps = 11/78 (14%)
Frame = +1
Query: 422 SIHALVTEDENNMLIRLPLRDEIKSAVFDLNG--------DGAPGPDGFGGHFYQSFW-- 471
+++ E + M++ P + +S+ DLNG DG PGPD H S
Sbjct: 988 ALYTFYHEKKPEMVVCPPTPEWKRSSELDLNGEAVPDEDDDGVPGPDLQENHETSSQMTN 1167
Query: 472 -DIVGSDVVLSVQDFFLQ 488
D++G ++ + Q+ F Q
Sbjct: 1168 DDVLGDEIPVDQQNAFRQ 1221
>TC78433 similar to GP|3790100|gb|AAC67586.1| pyrophosphate-dependent
phosphofructokinase beta subunit {Citrus x paradisi},
partial (29%)
Length = 658
Score = 29.3 bits (64), Expect = 9.1
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = +1
Query: 729 WFYDGGA*EDAWRFVGFWRWLNSFY 753
WF DG A* +A + GF RW +S++
Sbjct: 325 WFGDGAA*SEAEDWCGFVRWASSWW 399
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.346 0.153 0.561
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,647,820
Number of Sequences: 36976
Number of extensions: 830283
Number of successful extensions: 8671
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 3575
Number of HSP's successfully gapped in prelim test: 409
Number of HSP's that attempted gapping in prelim test: 4830
Number of HSP's gapped (non-prelim): 4499
length of query: 1145
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1038
effective length of database: 5,058,295
effective search space: 5250510210
effective search space used: 5250510210
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146565.8


