

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.3 - phase: 0 /pseudo
(281 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
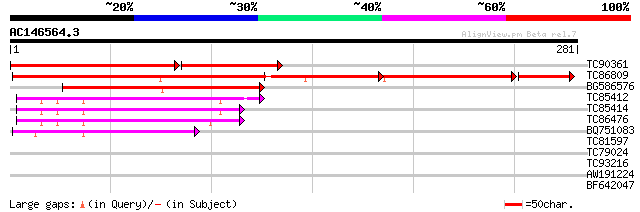
Score E
Sequences producing significant alignments: (bits) Value
TC90361 similar to PIR|S35701|S35701 translation elongation fact... 166 4e-65
TC86809 homologue to PIR|S35701|S35701 translation elongation fa... 215 2e-56
BG586576 similar to GP|10129885|db mitochondrial elongation fact... 82 3e-16
TC85412 homologue to GP|15450763|gb|AAK96653.1 elongation factor... 53 1e-07
TC85414 homologue to GP|15450763|gb|AAK96653.1 elongation factor... 52 3e-07
TC86476 similar to GP|15450763|gb|AAK96653.1 elongation factor E... 46 1e-05
BQ751083 homologue to GP|13925370|gb elongation factor 2 {Neuros... 42 3e-04
TC81597 similar to PIR|D96510|D96510 probable mitochondrial elon... 34 0.072
TC79024 similar to PIR|A84716|A84716 probable GTP-binding protei... 32 0.27
TC93216 similar to PIR|T51606|T51606 probable 26S proteasome non... 31 0.61
AW191224 similar to GP|14190379|gb| AT5g60590/muf9_240 {Arabidop... 27 8.8
BF642047 similar to GP|14190379|gb| AT5g60590/muf9_240 {Arabidop... 27 8.8
>TC90361 similar to PIR|S35701|S35701 translation elongation factor EF-G
chloroplast - soybean, partial (15%)
Length = 790
Score = 166 bits (420), Expect(2) = 4e-65
Identities = 82/84 (97%), Positives = 83/84 (98%)
Frame = +2
Query: 1 MDIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLA 60
MDIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLA
Sbjct: 359 MDIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLA 538
Query: 61 EEDPSFRVSQDKEDRTIIQGTGEL 84
EEDPSFRVSQDKEDRTIIQGTG +
Sbjct: 539 EEDPSFRVSQDKEDRTIIQGTGRI 610
Score = 99.8 bits (247), Expect(2) = 4e-65
Identities = 47/50 (94%), Positives = 49/50 (98%)
Frame = +1
Query: 86 LETIVDRLKREFKVEANVGAPQANYRESISEVTEVRYVFNKPYILNPWTQ 135
L+ +VDRLKREFKVEANVGAPQANYRESISEVTEVRYVFNKPYILNPWTQ
Sbjct: 616 LKPLVDRLKREFKVEANVGAPQANYRESISEVTEVRYVFNKPYILNPWTQ 765
>TC86809 homologue to PIR|S35701|S35701 translation elongation factor EF-G
chloroplast - soybean, partial (91%)
Length = 2674
Score = 215 bits (547), Expect = 2e-56
Identities = 116/198 (58%), Positives = 139/198 (69%), Gaps = 14/198 (7%)
Frame = +2
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLKDTI GETLCDP+ P VL++MDFP VIKIAIEPKTKAD++KM AGL+KLA+
Sbjct: 1409 DIVALAGLKDTITGETLCDPESPVVLERMDFPDPVIKIAIEPKTKADIDKMAAGLVKLAQ 1588
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKRE+KVEANVGAPQ NYRESIS++ E
Sbjct: 1589 EDPSFHFSRDEEINQTVIEGMGELHLEIIVDRLKREYKVEANVGAPQVNYRESISKIHEA 1768
Query: 121 RYVFNKPYILNPWTQVVDTNLRIKS-------------KKKHFQKNSLYWGF*RIGKLHE 167
RYV K Q D +R + K K + G *RIG+++E
Sbjct: 1769 RYVHKKQ--SGGQGQFADITVRFEPMEPGSGYEFKSEIKGGAVPKEYIPGGC*RIGRVYE 1942
Query: 168 QWCSSRLSGCWCTRSTCG 185
QWCS R+S C CT STCG
Sbjct: 1943 QWCSGRISSCRCTCSTCG 1996
Score = 119 bits (297), Expect(2) = 1e-32
Identities = 70/128 (54%), Positives = 83/128 (64%), Gaps = 3/128 (2%)
Frame = +3
Query: 127 PYILNPWTQVVDTNLRIKSKKKHFQKNSLYWGF*RIGKLHEQWCSSRLSGCWCTRSTC-- 184
PY N W+QVVD N R+KSK++ FQKN+ + G + L E + L+G
Sbjct: 1821 PYGSNQWSQVVDMNSRVKSKEEQFQKNT-FLGV--VKGLEECMSNGVLAGFPVVDVRAVL 1991
Query: 185 -G*YLP*CGFKCEGIPVGSKPSFYGRNEKSQTINA*TYNET*SCYS*RTL*RCTW*SQQK 243
+LP*C FKC G PVGSK SF GRN+KS+T NA*TYNE+*SCYS*RT RC W*SQ K
Sbjct: 1992 VDGFLP*CRFKCVGFPVGSKRSFSGRNKKSRTENA*TYNES*SCYS*RTFRRCNW*SQLK 2171
Query: 244 KRPNHKCW 251
KRPN W
Sbjct: 2172 KRPNQ*FW 2195
Score = 38.5 bits (88), Expect(2) = 1e-32
Identities = 17/28 (60%), Positives = 22/28 (77%)
Frame = +1
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASYSMQ FD V ++IQN+L T
Sbjct: 2278 MTKGRASYSMQLAMFDVVPQHIQNQLAT 2361
>BG586576 similar to GP|10129885|db mitochondrial elongation factor G {Oryza
sativa (japonica cultivar-group)}, partial (30%)
Length = 745
Score = 81.6 bits (200), Expect = 3e-16
Identities = 39/101 (38%), Positives = 63/101 (61%), Gaps = 1/101 (0%)
Frame = +1
Query: 27 LDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAEEDPSFRVSQDKED-RTIIQGTGELH 85
+ M P V+ +A++P +K + L + EDP+FRV D E +TII G GELH
Sbjct: 112 MTSMSVPEPVMSLAVQPVSKDSGGQFSKALNRFQREDPTFRVGLDPESGQTIISGMGELH 291
Query: 86 LETIVDRLKREFKVEANVGAPQANYRESISEVTEVRYVFNK 126
L+ V+R++RE+KV+A VG P+ N+RE++++ + Y+ K
Sbjct: 292 LDIYVERIRREYKVDATVGKPRVNFRETVTQRADFDYLHKK 414
>TC85412 homologue to GP|15450763|gb|AAK96653.1 elongation factor EF-2
{Arabidopsis thaliana}, complete
Length = 3229
Score = 52.8 bits (125), Expect = 1e-07
Identities = 41/129 (31%), Positives = 67/129 (51%), Gaps = 6/129 (4%)
Frame = +3
Query: 4 VTLVGLKDTIA-GETLCDPK--GPFVLDQMDFPVH-VIKIAIEPKTKADMEKMEAGLIKL 59
V LVGL I TL + K + M F V V+++A++ K +D+ K+ GL +L
Sbjct: 1521 VALVGLDQFITKNATLTNEKEVDAHPIRAMKFSVSPVVRVAVQCKVASDLPKLVEGLKRL 1700
Query: 60 AEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFKVEANV--GAPQANYRESISEV 117
A+ DP + ++ I+ G GELHLE + L+ +F A + P ++RE++ +
Sbjct: 1701 AKSDPMVVCTIEESGEHIVAGAGELHLEICLKDLQDDFMGGAEIIKSDPVVSFRETVLD- 1877
Query: 118 TEVRYVFNK 126
VR V +K
Sbjct: 1878 RSVRTVMSK 1904
>TC85414 homologue to GP|15450763|gb|AAK96653.1 elongation factor EF-2
{Arabidopsis thaliana}, complete
Length = 2892
Score = 51.6 bits (122), Expect = 3e-07
Identities = 37/119 (31%), Positives = 62/119 (52%), Gaps = 6/119 (5%)
Frame = +3
Query: 4 VTLVGLKDTIA-GETLCDPK--GPFVLDQMDFPVH-VIKIAIEPKTKADMEKMEAGLIKL 59
V +VGL I TL + K + M F V V+++A++ K +D+ K+ GL +L
Sbjct: 1479 VAMVGLDQFITKNATLTNEKEVDAHPIRAMKFSVSPVVRVAVQCKVASDLPKLVEGLKRL 1658
Query: 60 AEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFKVEANV--GAPQANYRESISE 116
A+ DP + ++ I+ G GELHLE + L+ +F A + P ++RE++ E
Sbjct: 1659 AKSDPMVVCTIEESGEHIVAGAGELHLEICLKDLQDDFMGGAEIIKSDPVVSFRETVLE 1835
>TC86476 similar to GP|15450763|gb|AAK96653.1 elongation factor EF-2
{Arabidopsis thaliana}, complete
Length = 2974
Score = 46.2 bits (108), Expect = 1e-05
Identities = 37/119 (31%), Positives = 57/119 (47%), Gaps = 6/119 (5%)
Frame = +2
Query: 4 VTLVGLKDTIA-GETLCDPK--GPFVLDQMDFPVH-VIKIAIEPKTKADMEKMEAGLIKL 59
V +VGL I TL + K + M F V V+ +A+ K +D+ K+ GL +L
Sbjct: 1598 VAMVGLDQFITKNATLTNEKEVDAHPIRAMKFSVSPVVSVAVTCKVASDLPKLVEGLKRL 1777
Query: 60 AEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFK--VEANVGAPQANYRESISE 116
A+ DP + + II GELHLE + L+ +F E P ++RE++ E
Sbjct: 1778 AKSDPMVVCTISETGEHIIAAAGELHLEICLKDLQDDFMNGAEITKSDPIVSFRETVLE 1954
>BQ751083 homologue to GP|13925370|gb elongation factor 2 {Neurospora
crassa}, partial (28%)
Length = 721
Score = 42.0 bits (97), Expect = 3e-04
Identities = 28/95 (29%), Positives = 48/95 (50%), Gaps = 2/95 (2%)
Frame = +3
Query: 2 DIVTLVGLKD-TIAGETLCDPKGPFVLDQMDFPVH-VIKIAIEPKTKADMEKMEAGLIKL 59
+IV LVG+ + TL L M F V V++ +++ K D+ K+ GL +L
Sbjct: 432 NIVGLVGIDQFLLKSGTLTTSDTAHNLKVMKFSVSPVVQRSVQVKNAQDLPKLVEGLKRL 611
Query: 60 AEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLK 94
++ DP ++ ++ G GELHLE ++ L+
Sbjct: 612 SKSDPCVLTMTNESGEHVVAGAGELHLEICLNDLE 716
>TC81597 similar to PIR|D96510|D96510 probable mitochondrial elongation
factor [imported] - Arabidopsis thaliana, partial (31%)
Length = 1065
Score = 33.9 bits (76), Expect = 0.072
Identities = 11/34 (32%), Positives = 24/34 (70%)
Frame = +2
Query: 93 LKREFKVEANVGAPQANYRESISEVTEVRYVFNK 126
++RE++ +A VG P+ N+RE++++ + Y+ K
Sbjct: 2 IRREYRADATVGKPRVNFRETVTQRADFDYLHKK 103
>TC79024 similar to PIR|A84716|A84716 probable GTP-binding protein [imported]
- Arabidopsis thaliana, partial (67%)
Length = 1513
Score = 32.0 bits (71), Expect = 0.27
Identities = 33/133 (24%), Positives = 55/133 (40%), Gaps = 16/133 (12%)
Frame = +2
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLD---------QMDFPVHVIKIAIEPKTKADMEKM 52
DI+++ GL G T+ + L M F V+ +A T K+
Sbjct: 1025 DIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGKI 1204
Query: 53 EAGLIKLAEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYR- 111
L+ +E + + V + +QG GEL L +++ ++RE E +V P+ Y+
Sbjct: 1205 GDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSPPKVMYKT 1381
Query: 112 ------ESISEVT 118
E I EVT
Sbjct: 1382 DKGQKLEPIEEVT 1420
>TC93216 similar to PIR|T51606|T51606 probable 26S proteasome non-ATPase
chain S5a [imported] - rice, partial (45%)
Length = 661
Score = 30.8 bits (68), Expect = 0.61
Identities = 15/38 (39%), Positives = 19/38 (49%), Gaps = 3/38 (7%)
Frame = +1
Query: 150 FQKNSLYWGF*---RIGKLHEQWCSSRLSGCWCTRSTC 184
FQK S+ + F RI +QWCS R W T + C
Sbjct: 22 FQKKSVLYSFKSRFRISSTFQQWCSRRHLSLWITLNIC 135
>AW191224 similar to GP|14190379|gb| AT5g60590/muf9_240 {Arabidopsis
thaliana}, partial (28%)
Length = 444
Score = 26.9 bits (58), Expect = 8.8
Identities = 14/40 (35%), Positives = 20/40 (50%)
Frame = +1
Query: 144 KSKKKHFQKNSLYWGF*RIGKLHEQWCSSRLSGCWCTRST 183
K+ K FQK+ L +G +G WC S + C C R +
Sbjct: 220 KN*KGFFQKHGLEYGKM*LGS*C*DWCGSSSNRCLCCRGS 339
>BF642047 similar to GP|14190379|gb| AT5g60590/muf9_240 {Arabidopsis
thaliana}, partial (29%)
Length = 489
Score = 26.9 bits (58), Expect = 8.8
Identities = 14/40 (35%), Positives = 20/40 (50%)
Frame = +1
Query: 144 KSKKKHFQKNSLYWGF*RIGKLHEQWCSSRLSGCWCTRST 183
K+ K FQK+ L +G +G WC S + C C R +
Sbjct: 211 KN*KGFFQKHGLEYGKM*LGS*C*DWCGSSSNRCLCCRGS 330
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.334 0.145 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,260,785
Number of Sequences: 36976
Number of extensions: 120821
Number of successful extensions: 1009
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 986
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1003
length of query: 281
length of database: 9,014,727
effective HSP length: 95
effective length of query: 186
effective length of database: 5,502,007
effective search space: 1023373302
effective search space used: 1023373302
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146564.3


