

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.14 - phase: 0 /pseudo
(312 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
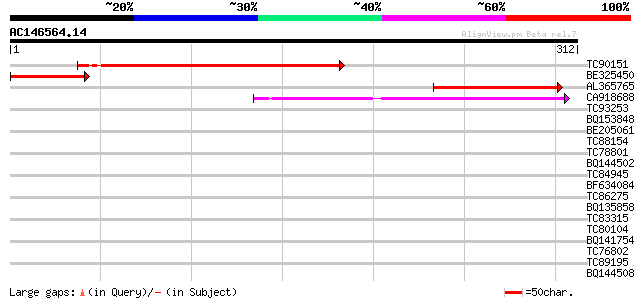
Score E
Sequences producing significant alignments: (bits) Value
TC90151 similar to GP|16323326|gb|AAL15376.1 At2g02370/T16F16.16... 112 2e-25
BE325450 95 3e-20
AL365765 similar to GP|16323326|gb| At2g02370/T16F16.16 {Arabido... 73 2e-13
CA918688 similar to GP|23617115|db contains EST AU173803(R3795)~... 54 6e-08
TC93253 weakly similar to GP|14030731|gb|AAK53040.1 AT5g65380/MN... 33 0.18
BQ153848 similar to GP|22136750|gb| unknown protein {Arabidopsis... 32 0.31
BE205061 30 0.92
TC88154 similar to PIR|T34137|T34137 hypothetical protein C33G8.... 30 1.2
TC78801 weakly similar to GP|13605827|gb|AAK32899.1 AT5g48380/MJ... 30 1.6
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 29 2.0
TC84945 similar to GP|20259551|gb|AAM13895.1 unknown protein {Ar... 29 2.0
BF634084 homologue to GP|21439028|emb unnamed protein product {H... 29 2.0
TC86275 similar to PIR|E96603|E96603 unknown protein F14G9.26 [i... 29 2.7
BQ135858 similar to GP|23491723|db formin homology protein A {Di... 28 3.5
TC83315 weakly similar to GP|19911209|dbj|BAB86931. glucosyltran... 28 3.5
TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR recept... 28 3.5
BQ141754 similar to GP|1903309|emb| orf 1035 {Spodoptera littora... 28 4.5
TC76802 homologue to SP|P35694|BRU1_SOYBN Brassinosteroid-regula... 28 4.5
TC89195 similar to GP|20197375|gb|AAM15048.1 expressed protein {... 28 4.5
BQ144508 weakly similar to GP|13880121|gb PE_PGRS family protein... 28 5.9
>TC90151 similar to GP|16323326|gb|AAL15376.1 At2g02370/T16F16.16
{Arabidopsis thaliana}, partial (28%)
Length = 762
Score = 112 bits (280), Expect = 2e-25
Identities = 55/147 (37%), Positives = 90/147 (60%)
Frame = +2
Query: 38 DYIKLIPGSDECLPLTAVEMVEECSLPSRRSVVWYWVKMVLLFLSLGFLAVAVLKWVGPY 97
+Y++L SDE P A + + SR + +W+K+ + + L++ ++KW P+
Sbjct: 329 EYVRLAI-SDE--PRVAEAEMLQPQAESRITSFKWWMKVSIWGFIIVILSLLLVKWGVPF 499
Query: 98 LIDKEVIPIINWETETFSPPVLTILLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLI 157
+K + P++ WE F PVL ++L AS+A+FP +L+PS PSMW+AG+ GYG GF++I
Sbjct: 500 AFEKVLYPVMEWEATAFGRPVLALVLIASLALFPVLLIPSGPSMWLAGMIFGYGLGFVII 679
Query: 158 ITAAAIGVSLPFIIGSIFHHKIEGWLE 184
+ IG+ LP++IG F +I WLE
Sbjct: 680 MVGTTIGMVLPYLIGLKFRDRIHQWLE 760
>BE325450
Length = 340
Score = 95.1 bits (235), Expect = 3e-20
Identities = 43/44 (97%), Positives = 43/44 (97%)
Frame = +2
Query: 1 MKDNTDDCGGGGGGGGWKKEVVSDVTITIEADDDNKGDYIKLIP 44
MKDNTDDCGGGGGGGGWKKEVVSDVTITIEADDDNKGDYIK IP
Sbjct: 209 MKDNTDDCGGGGGGGGWKKEVVSDVTITIEADDDNKGDYIK*IP 340
>AL365765 similar to GP|16323326|gb| At2g02370/T16F16.16 {Arabidopsis
thaliana}, partial (24%)
Length = 461
Score = 72.8 bits (177), Expect = 2e-13
Identities = 35/71 (49%), Positives = 48/71 (67%)
Frame = +2
Query: 234 PYIIGSLVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIIT 293
PY GS+ GMVPE F+ IY+G LIK+LADA R ++ +I+ N + F + V+T V T
Sbjct: 2 PYFCGSVAGMVPEAFIYIYSGRLIKSLADAQYGRHHMTTVEIVYNIISFIIAVITIVGFT 181
Query: 294 VYAKRRLKELQ 304
VYAKR L +L+
Sbjct: 182 VYAKRTLNKLK 214
>CA918688 similar to GP|23617115|db contains EST AU173803(R3795)~integral
membrane protein-like {Oryza sativa (japonica
cultivar-group)}, partial (59%)
Length = 888
Score = 54.3 bits (129), Expect = 6e-08
Identities = 43/176 (24%), Positives = 77/176 (43%), Gaps = 2/176 (1%)
Frame = -3
Query: 135 LPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHK-IEGWLEKYPKKASIL 193
+P++ +WV V+ G G L + + L F + F + YPK S+
Sbjct: 853 VPASVLLWV-WVSFGLPVGLLPTLLVQLLVQELHFFLVEQFEGPFVVSR*RDYPKFKSVA 677
Query: 194 KSAGAGNWFHQFRAVALIRISPF-PYMVFNYCAVATNVKYGPYIIGSLVGMVPEIFVAIY 252
+ F+ V L+R+ P P+ + NY T V Y++ S +GM+P +Y
Sbjct: 676 IAIRRSG----FKIVLLLRLVPLLPFNMLNYLLSVTPVSLVEYMLASWLGMMPITLALVY 509
Query: 253 TGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLKELQEEDK 308
G +K L+D ++ S + +G ++VV + +T AK L + E++
Sbjct: 508 VGTTLKDLSDVTHGWNEFSKSRWAFIIIGLIVSVVLMICVTKVAKSALDKALAENE 341
>TC93253 weakly similar to GP|14030731|gb|AAK53040.1 AT5g65380/MNA5_11
{Arabidopsis thaliana}, partial (25%)
Length = 633
Score = 32.7 bits (73), Expect = 0.18
Identities = 24/73 (32%), Positives = 40/73 (53%), Gaps = 4/73 (5%)
Frame = +2
Query: 113 TFSPPVLTILLFASVAIFPTILLPSTPSMWVAGVTLGYGF----GFLLIITAAAIGVSLP 168
T SPPVL + S+ + TILL S + ++GV +G G+ ++ I IG+ L
Sbjct: 35 TTSPPVLEAVNDMSILLAVTILLNSVQPI-LSGVAVGSGWQVFVAYVNIGCYYLIGLPLG 211
Query: 169 FIIGSIFHHKIEG 181
++G +F+ +EG
Sbjct: 212 ILMGWVFNTGVEG 250
>BQ153848 similar to GP|22136750|gb| unknown protein {Arabidopsis thaliana},
partial (36%)
Length = 591
Score = 32.0 bits (71), Expect = 0.31
Identities = 20/67 (29%), Positives = 34/67 (49%), Gaps = 2/67 (2%)
Frame = +2
Query: 205 FRAVALIRISPFPYMVFNYCAVATNVKYG-PYIIGSLVGMVPEIFVAIYTGILI-KTLAD 262
++ V L R SP P + NY AT V++ +++ ++ G +P I G L +A
Sbjct: 11 WKFVLLARFSPVPSYIINYTLAATEVRFFLDFLLPTIXGCIPMILQNTSIGSLAGAAVAT 190
Query: 263 ASNERQS 269
AS ++S
Sbjct: 191 ASGSKKS 211
>BE205061
Length = 468
Score = 30.4 bits (67), Expect = 0.92
Identities = 15/58 (25%), Positives = 26/58 (43%)
Frame = +2
Query: 167 LPFIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMVFNYC 224
+P ++G+ F+ I+ KY + GAG FH + +P P +F +C
Sbjct: 92 IPIVLGNSFYSSIDHPQNKYSSPPIFRCAGGAGTLFHD------VEYAPSPLGIFFFC 247
>TC88154 similar to PIR|T34137|T34137 hypothetical protein C33G8.2 -
Caenorhabditis elegans, partial (9%)
Length = 640
Score = 30.0 bits (66), Expect = 1.2
Identities = 12/41 (29%), Positives = 24/41 (58%)
Frame = -2
Query: 118 VLTILLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLII 158
V+ +++F + + P I++ S+W+ + GYG G L+I
Sbjct: 321 VVILIVFLLLLVNPRIIVRIVVSVWIVTMGFGYGCGVSLVI 199
>TC78801 weakly similar to GP|13605827|gb|AAK32899.1 AT5g48380/MJE7_1
{Arabidopsis thaliana}, partial (47%)
Length = 2001
Score = 29.6 bits (65), Expect = 1.6
Identities = 32/143 (22%), Positives = 54/143 (37%), Gaps = 16/143 (11%)
Frame = +3
Query: 158 ITAAAIGVSLPFIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPF- 216
+ A +GV L F + S+ H K E + W + I++S F
Sbjct: 1410 LAALGVGVGLLFFVRSVSHRKKE-------------EDPEGNKWARILKGTKKIKVSMFE 1550
Query: 217 ---PYMVFNYCAVATNVKYGPYIIGSLVGMVPEIFVAIY---TGILIKTLADASNERQSL 270
M + ATN +IG+ G ++ A+ T +++K L ++ + Q
Sbjct: 1551 KSISKMNLSDLMKATNNFSKSNVIGT--GRSGTVYKAVLDDGTSLMVKRLLESQHSEQEF 1724
Query: 271 SAQQIILNA---------LGFCL 284
+A+ L LGFCL
Sbjct: 1725 TAEMATLGTVRHRNLVPLLGFCL 1793
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 29.3 bits (64), Expect = 2.0
Identities = 9/14 (64%), Positives = 13/14 (92%)
Frame = +2
Query: 6 DDCGGGGGGGGWKK 19
++ GGGGGGGGW++
Sbjct: 665 EEGGGGGGGGGWRR 706
>TC84945 similar to GP|20259551|gb|AAM13895.1 unknown protein {Arabidopsis
thaliana}, partial (18%)
Length = 785
Score = 29.3 bits (64), Expect = 2.0
Identities = 10/16 (62%), Positives = 13/16 (80%)
Frame = -1
Query: 1 MKDNTDDCGGGGGGGG 16
++ + DCGGGGGGGG
Sbjct: 158 VRRSESDCGGGGGGGG 111
>BF634084 homologue to GP|21439028|emb unnamed protein product {Homo
sapiens}, partial (2%)
Length = 163
Score = 29.3 bits (64), Expect = 2.0
Identities = 10/13 (76%), Positives = 11/13 (83%)
Frame = -1
Query: 3 DNTDDCGGGGGGG 15
+ DDCGGGGGGG
Sbjct: 115 EEEDDCGGGGGGG 77
>TC86275 similar to PIR|E96603|E96603 unknown protein F14G9.26 [imported] -
Arabidopsis thaliana, partial (51%)
Length = 961
Score = 28.9 bits (63), Expect = 2.7
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = +2
Query: 2 KDNTDDCGGGGGGGGW 17
+D+ D GGGGGGG W
Sbjct: 536 RDDDDGGGGGGGGGNW 583
>BQ135858 similar to GP|23491723|db formin homology protein A {Dictyostelium
discoideum}, partial (4%)
Length = 1033
Score = 28.5 bits (62), Expect = 3.5
Identities = 10/11 (90%), Positives = 10/11 (90%)
Frame = -1
Query: 6 DDCGGGGGGGG 16
D CGGGGGGGG
Sbjct: 715 DGCGGGGGGGG 683
>TC83315 weakly similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13
{Vigna angularis}, partial (42%)
Length = 1048
Score = 28.5 bits (62), Expect = 3.5
Identities = 18/57 (31%), Positives = 27/57 (46%), Gaps = 1/57 (1%)
Frame = +2
Query: 149 GYG-FGFLLIITAAAIGVSLPFIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQ 204
GYG GF + T IG S ++G H+ WL+ P+ + + S G+G Q
Sbjct: 656 GYGKIGFFPVGTITQIGSSNNDVVGD--EHECLKWLKNQPQNSVLYVSFGSGGTLSQ 820
>TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR receptor protein
kinase {Arabidopsis thaliana}, partial (28%)
Length = 1549
Score = 28.5 bits (62), Expect = 3.5
Identities = 13/21 (61%), Positives = 14/21 (65%)
Frame = -3
Query: 9 GGGGGGGGWKKEVVSDVTITI 29
GGGGGGGG E S V I+I
Sbjct: 989 GGGGGGGGGGTEYTSGVLISI 927
>BQ141754 similar to GP|1903309|emb| orf 1035 {Spodoptera littoralis
nucleopolyhedrovirus}, partial (4%)
Length = 1308
Score = 28.1 bits (61), Expect = 4.5
Identities = 10/13 (76%), Positives = 10/13 (76%)
Frame = -3
Query: 4 NTDDCGGGGGGGG 16
N CGGGGGGGG
Sbjct: 40 NRGSCGGGGGGGG 2
>TC76802 homologue to SP|P35694|BRU1_SOYBN Brassinosteroid-regulated protein
BRU1 precursor. [Soybean] {Glycine max}, partial (71%)
Length = 890
Score = 28.1 bits (61), Expect = 4.5
Identities = 16/47 (34%), Positives = 27/47 (57%)
Frame = +3
Query: 249 VAIYTGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVY 295
V Y +LIK LA ASN R+S+ ++I N ++ +++T+Y
Sbjct: 168 VNFYIFLLIKLLALASNPRKSIYLVELICNLNLLLEILLELLLLTMY 308
>TC89195 similar to GP|20197375|gb|AAM15048.1 expressed protein {Arabidopsis
thaliana}, partial (55%)
Length = 896
Score = 28.1 bits (61), Expect = 4.5
Identities = 16/51 (31%), Positives = 25/51 (48%), Gaps = 8/51 (15%)
Frame = -1
Query: 133 ILLPSTPSMWVAGVTLGYGFGFLL--------IITAAAIGVSLPFIIGSIF 175
I LP TPS+WVA +G FG L +I++ ++ P ++ F
Sbjct: 821 ITLPVTPSVWVAAPEVGTPFGTWLFDAEGLNRVISSLSVRWKTPVVLAENF 669
>BQ144508 weakly similar to GP|13880121|gb PE_PGRS family protein
{Mycobacterium tuberculosis CDC1551}, partial (2%)
Length = 967
Score = 27.7 bits (60), Expect = 5.9
Identities = 11/25 (44%), Positives = 15/25 (60%)
Frame = -1
Query: 4 NTDDCGGGGGGGGWKKEVVSDVTIT 28
+T GG GGGGW ++V + IT
Sbjct: 97 STTAWSGGNGGGGWNTQLVRFIFIT 23
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.142 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,523,593
Number of Sequences: 36976
Number of extensions: 161781
Number of successful extensions: 3020
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 1484
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2624
length of query: 312
length of database: 9,014,727
effective HSP length: 96
effective length of query: 216
effective length of database: 5,465,031
effective search space: 1180446696
effective search space used: 1180446696
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146564.14


