

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
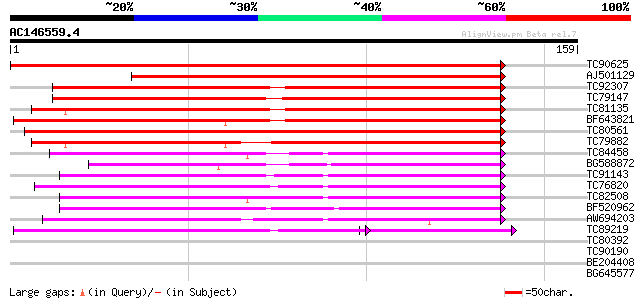
Query= AC146559.4 - phase: 0 /pseudo
(159 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC90625 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 280 2e-76
AJ501129 weakly similar to GP|10177856|dbj phloem-specific lecti... 171 9e-44
TC92307 weakly similar to PIR|E84434|E84434 probable phloem-spec... 168 7e-43
TC79147 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 148 1e-36
TC81135 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 145 5e-36
BF643821 weakly similar to PIR|F84434|F844 lectin-like protein [... 142 4e-35
TC80561 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 138 1e-33
TC79882 weakly similar to PIR|E84434|E84434 probable phloem-spec... 132 5e-32
TC84458 weakly similar to PIR|B96604|B96604 hypothetical protein... 96 8e-21
BG588872 weakly similar to GP|21592660|gb phloem-specific lectin... 85 1e-17
TC91143 similar to PIR|T47557|T47557 hypothetical protein F8J2.1... 72 1e-13
TC76820 weakly similar to GP|18175688|gb|AAL59911.1 unknown prot... 60 3e-10
TC82508 similar to PIR|T50527|T50527 hypothetical protein T27I15... 59 6e-10
BF520962 weakly similar to PIR|C96656|C96 unknown protein 43836... 57 2e-09
AW694203 weakly similar to PIR|C96656|C96 unknown protein 43836... 55 1e-08
TC89219 weakly similar to PIR|C96656|C96656 unknown protein 438... 47 8e-08
TC80392 weakly similar to GP|15088626|gb|AAK84134.1 nictaba {Nic... 38 0.002
TC90190 similar to GP|3426041|gb|AAC32240.1| unknown protein {Ar... 35 0.013
BE204408 similar to GP|10177692|db gene_id:MUL8.13~unknown prote... 34 0.029
BG645577 similar to GP|21535793|em putative carbamoyl phosphate ... 29 0.92
>TC90625 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial (9%)
Length = 892
Score = 280 bits (716), Expect = 2e-76
Identities = 139/139 (100%), Positives = 139/139 (100%)
Frame = +2
Query: 1 MDMKTEEAKSMIEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDL 60
MDMKTEEAKSMIEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDL
Sbjct: 191 MDMKTEEAKSMIEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDL 370
Query: 61 ISIISDSPSASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDIS 120
ISIISDSPSASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDIS
Sbjct: 371 ISIISDSPSASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDIS 550
Query: 121 IALGDTPQFWDWPILPESR 139
IALGDTPQFWDWPILPESR
Sbjct: 551 IALGDTPQFWDWPILPESR 607
>AJ501129 weakly similar to GP|10177856|dbj phloem-specific lectin-like
protein {Arabidopsis thaliana}, partial (33%)
Length = 558
Score = 171 bits (434), Expect = 9e-44
Identities = 84/105 (80%), Positives = 92/105 (87%)
Frame = +2
Query: 35 LSLVCKGFCSAAKSDTVWDRFLPSDLISIISDSPSASSLFSTSPSKKSLYLTLSDHPIVI 94
L L FCSAA+SD VWDRFLPSDLISI+SDS SASSLF+TSPSKKSLYLTLSDHPI+I
Sbjct: 80 LCLSVVSFCSAAESDIVWDRFLPSDLISIVSDSQSASSLFTTSPSKKSLYLTLSDHPILI 259
Query: 95 ENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQFWDWPILPESR 139
NGK SFQLEKQSG+K+YML ARD+SIA GDTP +WDW ILPESR
Sbjct: 260 NNGKTSFQLEKQSGKKVYMLGARDLSIAWGDTPCYWDWIILPESR 394
>TC92307 weakly similar to PIR|E84434|E84434 probable phloem-specific lectin
[imported] - Arabidopsis thaliana, partial (19%)
Length = 817
Score = 168 bits (426), Expect = 7e-43
Identities = 79/127 (62%), Positives = 100/127 (78%)
Frame = +1
Query: 13 EVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISIISDSPSASS 72
E LPEGCIA +LS T PVD+CRLS + K F SA+ SD VW+RFLPSD+ SIIS PS S+
Sbjct: 22 EELPEGCIAAVLSSTTPVDACRLSFISKNFLSASDSDAVWNRFLPSDISSIISQFPSLSN 201
Query: 73 LFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQFWDW 132
+ P+KK LY+ LSD PI+I+NGKKSFQLE++SG+K YMLSAR ++I GDT Q+W+W
Sbjct: 202 I----PTKKDLYIALSDRPIIIDNGKKSFQLERKSGKKCYMLSARSLAITWGDTDQYWNW 369
Query: 133 PILPESR 139
++PESR
Sbjct: 370 TVMPESR 390
>TC79147 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial
(32%)
Length = 958
Score = 148 bits (373), Expect = 1e-36
Identities = 74/127 (58%), Positives = 96/127 (75%)
Frame = +2
Query: 13 EVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISIISDSPSASS 72
E L EGCI+ ILS T PVD+ RLS+V K F SAA SD VW+ FLPSD+ SIIS SPS ++
Sbjct: 35 EDLAEGCISSILSRTTPVDAGRLSVVSKTFLSAADSDAVWNHFLPSDVNSIISQSPSLAN 214
Query: 73 LFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQFWDW 132
+PSKK+LYLTLSD PI+I++ KKSFQL+++SG+K YML+AR + I GD ++W W
Sbjct: 215 ----APSKKALYLTLSDRPIIIDDAKKSFQLDRKSGKKCYMLAARSLHIIWGDDDRYWIW 382
Query: 133 PILPESR 139
+P+SR
Sbjct: 383 TAMPDSR 403
>TC81135 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial
(32%)
Length = 1186
Score = 145 bits (367), Expect = 5e-36
Identities = 73/135 (54%), Positives = 100/135 (74%), Gaps = 2/135 (1%)
Frame = +2
Query: 7 EAKSMIEV--LPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISII 64
E K M E+ LPEGCIA I+S T P+D+ RLS+ K F SA+ SD VW++FLPSD+ SII
Sbjct: 86 ERKRMTEIQDLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSII 265
Query: 65 SDSPSASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALG 124
S SPS +++ P+KK LYL LSD P++I++ KKS QLE++SG+K YMLSAR ++I G
Sbjct: 266 SHSPSLTNI----PTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWG 433
Query: 125 DTPQFWDWPILPESR 139
D ++W+W +P+SR
Sbjct: 434 DDRRYWNWISMPDSR 478
>BF643821 weakly similar to PIR|F84434|F844 lectin-like protein [imported] -
Arabidopsis thaliana, partial (14%)
Length = 584
Score = 142 bits (359), Expect = 4e-35
Identities = 74/141 (52%), Positives = 102/141 (71%), Gaps = 3/141 (2%)
Frame = +2
Query: 2 DMKTEEAKSMIEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSD-- 59
++K + + E L EGCI+ I+S T P D+ RLS+V K F SAA SD VW++FLPSD
Sbjct: 53 EVKKSKRMTEFEELSEGCISAIISRTTPADAGRLSVVSKTFLSAADSDAVWNQFLPSDSH 232
Query: 60 -LISIISDSPSASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARD 118
+ SIIS SPS +++ PSKK+LYL LSD PI+I+N +KSFQL+++SG+K YML+AR
Sbjct: 233 FIDSIISHSPSLANV----PSKKALYLALSDLPIIIDNDQKSFQLDRKSGKKCYMLAARS 400
Query: 119 ISIALGDTPQFWDWPILPESR 139
+SIA GD ++W+W +P SR
Sbjct: 401 LSIAWGDDRRYWNWISMPNSR 463
>TC80561 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial
(52%)
Length = 1436
Score = 138 bits (347), Expect = 1e-33
Identities = 73/135 (54%), Positives = 89/135 (65%)
Frame = +3
Query: 5 TEEAKSMIEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISII 64
TEE LPEGCI +LS T P D RLSLV F SAA+SD+VWD+FLPSD SII
Sbjct: 141 TEEEGVEFHDLPEGCIENVLSFTTPRDVARLSLVSSTFRSAAESDSVWDKFLPSDYHSII 320
Query: 65 SDSPSASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALG 124
S S + + KK LY+ L+ P++IENGKKSFQL+K G+K YMLSAR + I G
Sbjct: 321 SKSDAEIDNSLSVLPKKDLYVNLTQKPLLIENGKKSFQLDKVDGKKCYMLSARSLFIVWG 500
Query: 125 DTPQFWDWPILPESR 139
DTP++W W P+SR
Sbjct: 501 DTPRYWRWIPDPDSR 545
>TC79882 weakly similar to PIR|E84434|E84434 probable phloem-specific lectin
[imported] - Arabidopsis thaliana, partial (14%)
Length = 636
Score = 132 bits (333), Expect = 5e-32
Identities = 72/138 (52%), Positives = 95/138 (68%), Gaps = 5/138 (3%)
Frame = +1
Query: 7 EAKSMIEV--LPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSD---LI 61
E K M E LPEGCI+ ILS T P D+ RLS+V K SAA SD VW++FLPSD +
Sbjct: 49 ELKKMTEFEELPEGCISAILSRTTPADAGRLSVVSKTILSAADSDAVWNQFLPSDSHFID 228
Query: 62 SIISDSPSASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISI 121
SIIS + P+KKSLYL LSD PI+I+N +KS QL+++SG+K YML+AR +SI
Sbjct: 229 SIIS--------LANVPTKKSLYLALSDSPIIIDNDQKSIQLDRKSGKKCYMLAARSLSI 384
Query: 122 ALGDTPQFWDWPILPESR 139
A G+ ++W+W +P+SR
Sbjct: 385 AWGNDDRYWNWIAMPDSR 438
>TC84458 weakly similar to PIR|B96604|B96604 hypothetical protein F14G9.14
[imported] - Arabidopsis thaliana, partial (24%)
Length = 766
Score = 95.5 bits (236), Expect = 8e-21
Identities = 55/131 (41%), Positives = 75/131 (56%), Gaps = 3/131 (2%)
Frame = +1
Query: 12 IEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISIIS---DSP 68
+E PE C+A ILS T+P+D CR+SL+ S A SD VW++FLP + I+S D P
Sbjct: 16 MEFSPEDCVAHILSFTSPIDVCRVSLISSIVKSMADSDNVWEKFLPHNYQGIVSRLVDPP 195
Query: 69 SASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQ 128
+ S SKK L L P +I++G K F +EK +G+ Y LSAR +SI G+ P
Sbjct: 196 LSCS------SKKELLARLC-KPQLIDDGNKMFSIEKTTGKICYSLSARQLSITFGNNPL 354
Query: 129 FWDWPILPESR 139
W W + SR
Sbjct: 355 HWSWRQVQGSR 387
>BG588872 weakly similar to GP|21592660|gb phloem-specific lectin PP2-like
protein {Arabidopsis thaliana}, partial (55%)
Length = 655
Score = 85.1 bits (209), Expect = 1e-17
Identities = 49/120 (40%), Positives = 71/120 (58%), Gaps = 3/120 (2%)
Frame = +1
Query: 23 ILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLP---SDLISIISDSPSASSLFSTSPS 79
ILS T+P+D CR+SL+ S A SD +W++FLP ++IS + D P S S
Sbjct: 19 ILSFTSPLDVCRVSLLSSVVQSMADSDFLWEKFLPHKYQEIISRLVDPPLPCS------S 180
Query: 80 KKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQFWDWPILPESR 139
KK L+ L P +I++G K F +EK +G+ Y+LSAR +SI G+T +W W + SR
Sbjct: 181 KKELFARLCK-PSLIDDGNKMFSIEKTTGKICYLLSARQLSITFGNTSLYWSWKQVQGSR 357
>TC91143 similar to PIR|T47557|T47557 hypothetical protein F8J2.170 -
Arabidopsis thaliana, partial (60%)
Length = 669
Score = 72.0 bits (175), Expect = 1e-13
Identities = 38/125 (30%), Positives = 66/125 (52%)
Frame = +1
Query: 15 LPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISIISDSPSASSLF 74
+PE C+A++ H P + C L+ + + F AA +D+VW+ LPS+ ++ P
Sbjct: 157 IPENCVARVFLHLTPPEICNLARLNRAFRGAASADSVWESKLPSNYQHLLRLMPEEQR-- 330
Query: 75 STSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQFWDWPI 134
+ SKK ++ S P+ ++G K L++ +GR +SA+ +SI D ++W W
Sbjct: 331 CRNLSKKDIFAVFS-KPLPFDDGNKQLWLDRVTGRVCMSISAKGLSITGIDDRRYWTWVP 507
Query: 135 LPESR 139
ESR
Sbjct: 508 TEESR 522
>TC76820 weakly similar to GP|18175688|gb|AAL59911.1 unknown protein
{Arabidopsis thaliana}, partial (67%)
Length = 1388
Score = 60.5 bits (145), Expect = 3e-10
Identities = 37/132 (28%), Positives = 64/132 (48%)
Frame = +2
Query: 8 AKSMIEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISIISDS 67
+K+ ++ +PE CI+ I+ P + C+L+ V K F A+ D VW+ LP ++
Sbjct: 176 SKTCLDDIPESCISSIMMRLDPQEICKLARVNKTFHRASSDDYVWESKLPPSYKFLVKKI 355
Query: 68 PSASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALGDTP 127
L S +KK +Y L P + G K L++ SG+ +S++ I D
Sbjct: 356 LGEEKL--CSMTKKEIYAKLC-QPNFFDGGTKEILLDRCSGQVCLFISSKSFKITGIDDR 526
Query: 128 QFWDWPILPESR 139
++W++ ESR
Sbjct: 527 RYWNYISTEESR 562
>TC82508 similar to PIR|T50527|T50527 hypothetical protein T27I15_150 -
Arabidopsis thaliana, partial (71%)
Length = 1761
Score = 59.3 bits (142), Expect = 6e-10
Identities = 36/128 (28%), Positives = 65/128 (50%), Gaps = 3/128 (2%)
Frame = +2
Query: 15 LPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISIIS---DSPSAS 71
+PE C+A +L + P D C+L+ + + F A+ +D VW+ LP + I+ + + S
Sbjct: 719 IPESCVALVLMYLDPPDICKLARLNRAFRDASFADFVWESKLPLNYEFIMGKALEDDTTS 898
Query: 72 SLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQFWD 131
S K+ +Y L P + +NG K L+K++G +S++ + I D ++W+
Sbjct: 899 SSSGAELGKRDIYARLC-KPNLFDNGTKEIWLDKRTGGVCLAISSKALRITGIDDRRYWN 1075
Query: 132 WPILPESR 139
ESR
Sbjct: 1076HISTEESR 1099
>BF520962 weakly similar to PIR|C96656|C96 unknown protein 43836-42510
[imported] - Arabidopsis thaliana, partial (58%)
Length = 681
Score = 57.4 bits (137), Expect = 2e-09
Identities = 38/125 (30%), Positives = 62/125 (49%)
Frame = +3
Query: 15 LPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISIISDSPSASSLF 74
LP+ CIA I+ + P C+L+ + + F A+ +D VW+ LPS+ II SS F
Sbjct: 180 LPQSCIAGIMEYMDPPQICKLASLNRAFHGASLADFVWESKLPSNYHVIIQKIFDGSS-F 356
Query: 75 STSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQFWDWPI 134
+ K+ +Y L + G K L++ G+ +SA+ +SI D ++W+
Sbjct: 357 PSHLGKRGIYSRLCSLN-TFDEGTKKVWLDRSMGKVCLSVSAKGLSITGIDDRRYWNHVP 533
Query: 135 LPESR 139
ESR
Sbjct: 534 TEESR 548
>AW694203 weakly similar to PIR|C96656|C96 unknown protein 43836-42510
[imported] - Arabidopsis thaliana, partial (52%)
Length = 621
Score = 55.1 bits (131), Expect = 1e-08
Identities = 34/131 (25%), Positives = 69/131 (51%), Gaps = 1/131 (0%)
Frame = +1
Query: 10 SMIEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISIISDSPS 69
S + LPE C+A I+ + P C+L+ + + F +A+ +D VW+ LP + +++
Sbjct: 157 STLGELPESCVASIIGYMDPPQICQLATLNRAFRAASSADFVWESKLPPNYHLLLA---K 327
Query: 70 ASSLFSTSPSKKSLYLTLSDHPIVIENGKKSFQLEKQSGRKIYMLSA-RDISIALGDTPQ 128
F + K+ +Y +L I++G K +++ +G+ +SA + +S+ D +
Sbjct: 328 IFHHFPINLGKRDIYASLC-RLNTIDDGTKKVWIDRATGKLCLAISANKGLSVLGVDDRR 504
Query: 129 FWDWPILPESR 139
+W++ I ESR
Sbjct: 505 YWNYIITEESR 537
>TC89219 weakly similar to PIR|C96656|C96656 unknown protein 43836-42510
[imported] - Arabidopsis thaliana, partial (46%)
Length = 764
Score = 47.4 bits (111), Expect(2) = 8e-08
Identities = 30/100 (30%), Positives = 50/100 (50%)
Frame = +3
Query: 2 DMKTEEAKSMIEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLI 61
+ + +K+ + +PE CI+ IL H P D C+L+ V + F A+ +D VW+ LPS
Sbjct: 243 ESNSSSSKTGLGDIPESCISSILMHLDPHDICKLARVNRAFHRASSADFVWESKLPSCYK 422
Query: 62 SIISDSPSASSLFSTSPSKKSLYLTLSDHPIVIENGKKSF 101
+ + +L ++ SKK +Y L K+SF
Sbjct: 423 FLANKFLGEENL--STMSKKDVYTKLCQPNRFDGGTKRSF 536
Score = 24.6 bits (52), Expect(2) = 8e-08
Identities = 13/44 (29%), Positives = 22/44 (49%)
Frame = +1
Query: 99 KSFQLEKQSGRKIYMLSARDISIALGDTPQFWDWPILPESRCVN 142
K L+K SG+ +S++ + I D ++W + ESR N
Sbjct: 526 KEVLLDKYSGQVCLFMSSKSLKITGIDDRRYWIYIPTEESRIKN 657
>TC80392 weakly similar to GP|15088626|gb|AAK84134.1 nictaba {Nicotiana
tabacum}, partial (29%)
Length = 1161
Score = 37.7 bits (86), Expect = 0.002
Identities = 19/45 (42%), Positives = 29/45 (64%)
Frame = +2
Query: 95 ENGKKSFQLEKQSGRKIYMLSARDISIALGDTPQFWDWPILPESR 139
E+ K+ +K+S MLSAR +SIA GD ++W+W +P+SR
Sbjct: 53 EHQKRKRDKKKES----CMLSARSLSIAWGDDKRYWNWINMPDSR 175
>TC90190 similar to GP|3426041|gb|AAC32240.1| unknown protein {Arabidopsis
thaliana}, partial (60%)
Length = 1626
Score = 35.0 bits (79), Expect = 0.013
Identities = 20/64 (31%), Positives = 34/64 (52%), Gaps = 2/64 (3%)
Frame = +2
Query: 6 EEAKSMIEV--LPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFLPSDLISI 63
EE S++++ LP CI + LS P + C ++ VCK +SD +W++ + I
Sbjct: 527 EEKASLLDLHDLPLDCILEKLS---PSELCNVAQVCKSLREKCRSDYLWEKHMKMKWGKI 697
Query: 64 ISDS 67
+ DS
Sbjct: 698 LGDS 709
>BE204408 similar to GP|10177692|db gene_id:MUL8.13~unknown protein
{Arabidopsis thaliana}, partial (23%)
Length = 658
Score = 33.9 bits (76), Expect = 0.029
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = +2
Query: 10 SMIEVLPEGCIAKILSHTAPVDSCRLSLVCKGFCSAAKSDTVW 52
+M+ LP+ A I +P D C LSL CK S S+ +W
Sbjct: 122 NMLLYLPDDVFAIISKSFSPRDICNLSLCCKSLNSLVASEKIW 250
>BG645577 similar to GP|21535793|em putative carbamoyl phosphate synthase
small subunit {Nicotiana tabacum}, partial (38%)
Length = 721
Score = 28.9 bits (63), Expect = 0.92
Identities = 22/81 (27%), Positives = 35/81 (43%), Gaps = 6/81 (7%)
Frame = +1
Query: 29 PVDSCRLS------LVCKGFCSAAKSDTVWDRFLPSDLISIISDSPSASSLFSTSPSKKS 82
P+ SC +S +C G S K + PS +S+ + P+ +L P
Sbjct: 7 PLSSCHIS*SLLQTSLCIGVTSKKKQR*WRQKLWPSPYLSMTALVPNRHTLLQKFP---- 174
Query: 83 LYLTLSDHPIVIENGKKSFQL 103
+LSD P+V+E G Q+
Sbjct: 175 --FSLSDVPLVMEKGHGKIQM 231
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.133 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,317,859
Number of Sequences: 36976
Number of extensions: 74201
Number of successful extensions: 524
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 512
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 513
length of query: 159
length of database: 9,014,727
effective HSP length: 88
effective length of query: 71
effective length of database: 5,760,839
effective search space: 409019569
effective search space used: 409019569
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146559.4


