

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146556.9 + phase: 0
(2244 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
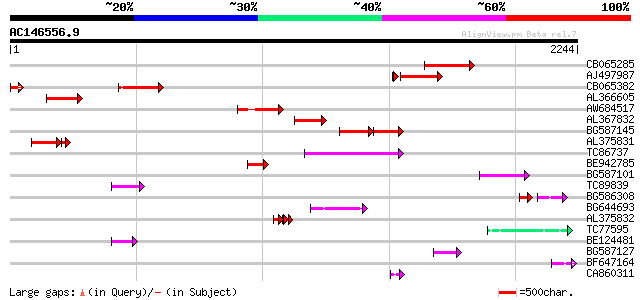
Score E
Sequences producing significant alignments: (bits) Value
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 397 e-110
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 299 7e-81
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 297 4e-80
AL366605 275 1e-73
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 249 7e-66
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 240 5e-63
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 152 3e-59
AL375831 168 2e-56
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 181 2e-45
BE942785 133 7e-31
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 115 3e-25
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 93 1e-18
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 60 7e-18
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 85 3e-16
AL375832 80 9e-15
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 78 3e-14
BE124481 74 5e-13
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 66 1e-10
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 50 1e-05
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 49 2e-05
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 397 bits (1021), Expect = e-110
Identities = 189/197 (95%), Positives = 193/197 (97%)
Frame = -2
Query: 1643 KSKDCEEPLINEGPDPISKWGLVFDGAVNAYGKGIGAVIVSPQGHHIPFTARILFECTNN 1702
KSKDCEEPLI EGPDP SKWGLVFDGAVNAYGKGIGAVIVSPQGH+IPFTARILFECTNN
Sbjct: 591 KSKDCEEPLIGEGPDPNSKWGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNN 412
Query: 1703 MAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYF 1762
MAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHA LIPYRDYARRLLTYF
Sbjct: 411 MAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYF 232
Query: 1763 TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGDVIDQAGE 1822
TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIG+VIDQ GE
Sbjct: 231 TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGNVIDQTGE 52
Query: 1823 NVVDYRPWYYDIKQFLL 1839
NVVDY+PWYYDIKQFL+
Sbjct: 51 NVVDYKPWYYDIKQFLV 1
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 299 bits (766), Expect = 7e-81
Identities = 133/167 (79%), Positives = 155/167 (92%)
Frame = -2
Query: 1547 KTCCALAWAAKRLRHYLVNHTTWLISRMDPIKYIFEKAAVTGKIARWQMLLSEYDIVFKT 1606
KTCCALAWAAKRLRHY++NHTTWL+S+MDPIKYIFEK A+TG+IARWQMLLSEYDI +++
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1607 QKAIKGSILADHLAYQPLDDYQPIEFDFPDEEIMYLKSKDCEEPLINEGPDPISKWGLVF 1666
QKAIKGSILADHLA+QPL+DY+PI+FDFPDEEIMYLK KDC+EPL EGPDP S WGL+F
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 1667 DGAVNAYGKGIGAVIVSPQGHHIPFTARILFECTNNMAEYEACIFGI 1713
DGAVN YG GIGAV+++P+G HIPFTAR+ F+CTNN+AEYEACI GI
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
Score = 48.1 bits (113), Expect = 4e-05
Identities = 20/24 (83%), Positives = 23/24 (95%)
Frame = +2
Query: 1513 CVLGQQDETGKKEHAIYYLSKKFT 1536
CVLGQQDETG+KEHAIYYLS+K +
Sbjct: 38 CVLGQQDETGRKEHAIYYLSRKLS 109
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 297 bits (760), Expect = 4e-80
Identities = 146/177 (82%), Positives = 153/177 (85%)
Frame = -2
Query: 432 PVQQQQQQQQQQQQYTRPTFPPIPMLYAELLPTLLQRGHCTTRQGKPPPDPLPPRFRSDL 491
PV QQ+Q QQ RPTFP IPMLYAE LPTLL RGHCT RQGKPPPDPLPPRFRSDL
Sbjct: 518 PVHQQRQ*QQ-----ARPTFPLIPMLYAE*LPTLLLRGHCTIRQGKPPPDPLPPRFRSDL 354
Query: 492 KCDFHQGALGHDVEGCYALKYIVKKLIDQGKLTFENNVPHVLDNPLPNHAAVNMIEVCEE 551
KCDFHQGALGHDVEGCYALK+IVKKLI+QGKLTFENNVPHVLDNPLPNHAAVNMIEV EE
Sbjct: 353 KCDFHQGALGHDVEGCYALKHIVKKLINQGKLTFENNVPHVLDNPLPNHAAVNMIEVYEE 174
Query: 552 APRLDVRNVATPLVPLHIKLCKASLFSHDHAKCLGCLRDPLGCHAVQDDIQSLMNDN 608
AP LDVRNV TPLVPLHIKLC+ASLF HDHA C +PLGC VQ+DIQSLMN+N
Sbjct: 173 APGLDVRNVTTPLVPLHIKLCQASLFDHDHANCQE*FYNPLGCCVVQNDIQSLMNNN 3
Score = 57.0 bits (136), Expect = 8e-08
Identities = 32/50 (64%), Positives = 37/50 (74%)
Frame = -1
Query: 3 EENTQLRTKLASLREELAKAHDAMTALLATQEQPVPVVSTAANVTPTVTT 52
EEN QLRT+LAS+REELAKA+D MTALLA QEQ S+ + T TV T
Sbjct: 624 EENAQLRTELASMREELAKANDTMTALLAAQEQ-----SSTGSPTTTVAT 490
>AL366605
Length = 422
Score = 275 bits (704), Expect = 1e-73
Identities = 129/139 (92%), Positives = 135/139 (96%)
Frame = -3
Query: 147 EIKANRGNGDSIKTQDLCLVSKVDVPKKFKVPEFDKYNGLTCPQNHIVKYVRKMGNYKDN 206
EIKANRGN DS KTQDLCLVSKVDVPKKFK+P+FD+YNGLTCPQNHI+KYVRKMGNYKDN
Sbjct: 417 EIKANRGNADSFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDN 238
Query: 207 DSLMIHYFQDSLMEDAAEWYTSLSKDDVHTFDELATAFKSHYGFNTRLKPNREFLRSLSQ 266
DSLMIH FQDSLMEDAAEWYTSLSK+D+HTFDELA AFKSHYGFNTRLKPNREFLRSLSQ
Sbjct: 237 DSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQ 58
Query: 267 KKEESFREYAQRWRGAAAR 285
KKEESFREYAQRWRGAAAR
Sbjct: 57 KKEESFREYAQRWRGAAAR 1
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana chromosome
II BAC F26H6; putative retroelement pol polyprotein,
partial (1%)
Length = 488
Score = 249 bits (637), Expect = 7e-66
Identities = 126/180 (70%), Positives = 140/180 (77%)
Frame = +1
Query: 902 ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKFIRNGKLVTVHGEEAYLVSQLTSF 961
ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKF++NG
Sbjct: 1 ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKFVKNG------------------- 123
Query: 962 SCIEAGSAEGTAFQGLTIEGAEPKKAGAAMASLKDAQKVIQEGQTAGWGKVIQLCENKRK 1021
SAEGTAFQGL++EGAEPKK GAAMASLKDAQK +QEGQ A WGK+IQLCENKRK
Sbjct: 124 ------SAEGTAFQGLSMEGAEPKKVGAAMASLKDAQKAVQEGQAADWGKLIQLCENKRK 285
Query: 1022 EGLGFSPSSRVSSGVFHSAGFVNAISEEATGSGLRPVFVTPGGIARDWNAIDVPSIMHVS 1081
EGL FSP+S VS+G FHSAGFVN ++EE RP+FV PGGIA+DW+A+DVPSIMHVS
Sbjct: 286 EGLRFSPTSGVSTGTFHSAGFVNTLAEEVARFVPRPLFVIPGGIAKDWDAVDVPSIMHVS 465
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 240 bits (612), Expect = 5e-63
Identities = 116/128 (90%), Positives = 122/128 (94%)
Frame = -1
Query: 1125 QEKKAIQPHQEEIELVNIGTEENKREIKIGATLEEGVKQKIIQLLREYPDIFAWSYEDMP 1184
QE+KAIQPHQEEIEL+N+GTEENKREIK+GA LEEGVK+KI QLLREY DIFA SYEDMP
Sbjct: 384 QERKAIQPHQEEIELINLGTEENKREIKVGAALEEGVKRKIFQLLREYLDIFACSYEDMP 205
Query: 1185 GLDPMIVEHQIPTKPECPPVRQKLRRTHPDMALKIKSEVQKQIDAGFLMTVEYPEWVANI 1244
GLDP IVEH+IPTKPECPPVR KLRRTHPDMALKIKSEVQKQIDAGFLMTVEYPEWVANI
Sbjct: 204 GLDPKIVEHRIPTKPECPPVR*KLRRTHPDMALKIKSEVQKQIDAGFLMTVEYPEWVANI 25
Query: 1245 VPVPKKDG 1252
VPVPKKDG
Sbjct: 24 VPVPKKDG 1
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 152 bits (383), Expect(2) = 3e-59
Identities = 70/136 (51%), Positives = 101/136 (73%)
Frame = +2
Query: 1304 MSPEDREKTSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVK 1363
M P+D EKT+FIT GT+CYKVMPFGL NAG+TYQR + +F D + +EVY+DDM+VK
Sbjct: 2 MHPDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVK 181
Query: 1364 STDEEQHVEYLTKMFERLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIR 1423
S H+ +L + F+ L +Y ++LNP KCTFGV SG+ LG+IV+Q+GIEV+P ++ AI
Sbjct: 182 SLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAIL 361
Query: 1424 EMPAPQTEKQVRGFLG 1439
++P+P+ ++V+ G
Sbjct: 362 DLPSPKNSREVQRLTG 409
Score = 97.4 bits (241), Expect(2) = 3e-59
Identities = 49/120 (40%), Positives = 75/120 (61%)
Frame = +3
Query: 1439 GRLNYISRFISHMTATCGPIFKLLRKNQPIVWNDECQEAFDSIKNYLLEPPILVPPVEGR 1498
GR+ ++RFIS T C P +KLL N+ VW+++C+EAF+ +K YL PP+L P G
Sbjct: 408 GRIAALNRFISRSTDKCLPFYKLLCGNKRFVWDEKCEEAFEQLKQYLTTPPVLSKPEAGD 587
Query: 1499 PLIMYLAVFDESMGCVLGQQDETGKKEHAIYYLSKKFTDCETRYTMLEKTCCALAWAAKR 1558
L +Y+A+ ++ VL ++D +K I+Y SK+ TD ETRY LEK A+ +A++
Sbjct: 588 TLSLYIAISSTAVSSVLIREDRGEQK--PIFYTSKRMTDPETRYPTLEKMAFAVITSARK 761
>AL375831
Length = 467
Score = 168 bits (426), Expect(2) = 2e-56
Identities = 78/119 (65%), Positives = 91/119 (75%)
Frame = +2
Query: 87 IPGANLTPVSTALTQAATTVTEPIVNAVPLFVHANAHHGSTATTGTIEERMAELAKELRR 146
IP N + L Q VTEP+V+ +P ++ N H S T T+EE M ELAKELR
Sbjct: 8 IPATNAASIIATLPQTTAAVTEPLVHTLPQGININTQHRSIPVTKTMEEMMEELAKELRH 187
Query: 147 EIKANRGNGDSIKTQDLCLVSKVDVPKKFKVPEFDKYNGLTCPQNHIVKYVRKMGNYKD 205
EI+ANRGN DS+KTQDLCLVSKVDVPKKFK+P+FD+YNGLTCPQNHI+KYVRKMGNYKD
Sbjct: 188 EIQANRGNADSVKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 364
Score = 71.2 bits (173), Expect(2) = 2e-56
Identities = 32/36 (88%), Positives = 34/36 (93%)
Frame = +3
Query: 204 KDNDSLMIHYFQDSLMEDAAEWYTSLSKDDVHTFDE 239
K NDSLMIH FQDSLMEDAAEWYTSLSK+D+HTFDE
Sbjct: 360 KTNDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDE 467
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 181 bits (460), Expect = 2e-45
Identities = 123/398 (30%), Positives = 200/398 (49%), Gaps = 7/398 (1%)
Frame = +1
Query: 1168 LLREYPDIFAWSYEDMPGLDPMIVEHQIPTKPEC----PPVRQ-KLRRTHPDMALKIKSE 1222
+L E+PD+F +++H IP P+ PP+ L L +K
Sbjct: 343 VLEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKDGNDPPLPWGPLYGMSRQELLVLKKT 522
Query: 1223 VQKQIDAGFLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVD 1282
++ +D GF+ A ++ V K G +R CVD+R LN + KD +PLP I +
Sbjct: 523 LEDLLDKGFIKASGSAAG-APVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLR 699
Query: 1283 NTAQSKVFSFMDGFSGYNQIKMSPEDREKTSFITPWGTFCYKVMPFGLINAGATYQRGMT 1342
A ++ F+ +D + ++++++ ED+EKT+F T +G F + V PFGL A AT+QR +
Sbjct: 700 RVAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEWIVCPFGLTGAPATFQRYIN 879
Query: 1343 TLFHDMIHKEVEVYVDDMIVKST-DEEQHVEYLTKMFERLRKYKLRLNPNKCTFGVRSGK 1401
H+ + V Y+DD+++ +T ++ H + ++ RL L L+P KC F V + K
Sbjct: 880 KTLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEFSVTTVK 1059
Query: 1402 LLGFIVSQ-KGIEVDPDKVRAIREMPAPQTEKQVRGFLGRLNYISRFISHMTATCGPIFK 1460
+GFI++ KG+ DP K+ AIR+ P + K R FLG NY FI + P+ +
Sbjct: 1060 YVGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYYKDFIPGYSEITEPLTR 1239
Query: 1461 LLRKNQPIVWNDECQEAFDSIKNYLLEPPILVPPVEGRPLIMYLAVFDESMGCVLGQQDE 1520
L RK+ P W E + AF +K E P+L + ++G VL Q+D
Sbjct: 1240 LTRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPEAVTTVETDCSGFALGGVLTQEDG 1419
Query: 1521 TGKKEHAIYYLSKKFTDCETRYTMLEKTCCALAWAAKR 1558
TG H + + S++ + E Y + +K A+ WA R
Sbjct: 1420 TG-AAHPVAFHSQRLSPAEYNYPIHDKELLAV-WACLR 1527
>BE942785
Length = 460
Score = 133 bits (335), Expect = 7e-31
Identities = 62/82 (75%), Positives = 73/82 (88%)
Frame = -2
Query: 941 NGKLVTVHGEEAYLVSQLTSFSCIEAGSAEGTAFQGLTIEGAEPKKAGAAMASLKDAQKV 1000
+G LVT+HGEEAYL+SQL+SFSCIEAGSAEGTAFQGLT+EG EPK+ G AMASLKDAQ+
Sbjct: 378 SGXLVTIHGEEAYLISQLSSFSCIEAGSAEGTAFQGLTVEGTEPKRDGTAMASLKDAQRA 199
Query: 1001 IQEGQTAGWGKVIQLCENKRKE 1022
+QE Q AGWG++IQL ENK K+
Sbjct: 198 VQESQAAGWGRLIQLRENKHKD 133
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 115 bits (287), Expect = 3e-25
Identities = 63/196 (32%), Positives = 98/196 (49%)
Frame = +2
Query: 1861 RFLLDGDILYKRNYDMVLLRCVDEHEAEQLMHDVHDGTFGTHATGHTMSRKLLRAGYYWM 1920
R+L D LYK+ D + +RCV E E ++ H + H K+ +AG++W
Sbjct: 17 RYLWDEPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGFWWP 196
Query: 1921 TMEHDCYQYARKCHKCQIYADKIHVPPHALNVMSSPWPFSMWGIDMIGRIEPKASNGHRF 1980
TM D + + KC CQ + N + F +WGID +G +N ++
Sbjct: 197 TMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPSSYNN--KY 370
Query: 1981 ILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALC 2040
ILVA+DY +KWVEA + VV K K+ I R+GVP +I+D G++ N V + L
Sbjct: 371 ILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLL 550
Query: 2041 EEFKIEHHNSSPYRPQ 2056
++ + H ++ Y PQ
Sbjct: 551 KKNGVRHKVATAYHPQ 598
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 93.2 bits (230), Expect = 1e-18
Identities = 54/132 (40%), Positives = 70/132 (52%), Gaps = 1/132 (0%)
Frame = +3
Query: 403 PPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQQQQQQQQYTRPT-FPPIPMLYAEL 461
PP Q P P QP P Q +Q +P Q Q+ Q Q + + PIP+ YA+L
Sbjct: 228 PPLVQQRPQSPYQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVKYADL 407
Query: 462 LPTLLQRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIVKKLIDQG 521
LP LL++ T P+ LPP +R DL C FHQGA GHD E CY LK V+KLI+
Sbjct: 408 LPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHDTEQCYPLKEEVQKLIENN 587
Query: 522 KLTFENNVPHVL 533
+F++ VL
Sbjct: 588 VWSFDDQDIKVL 623
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 60.5 bits (145), Expect(2) = 7e-18
Identities = 39/122 (31%), Positives = 68/122 (54%), Gaps = 4/122 (3%)
Frame = -3
Query: 2090 LHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSL---RVIMEAKLSEAEWCQSRYDQL 2146
L +RT R +T +TPFS+ + +EA+ P EV + SL R+ +L+ ++ L
Sbjct: 462 LWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNIELNN----DRLFNAL 295
Query: 2147 NLIEEKRMDAMARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQPD-PRGKWTPNY 2205
IEE+R A+ R Q+YQ ++++ ++K V + K+G++VL + + GK N+
Sbjct: 294 ETIEERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNW 115
Query: 2206 EG 2207
EG
Sbjct: 114 EG 109
Score = 50.4 bits (119), Expect(2) = 7e-18
Identities = 22/52 (42%), Positives = 35/52 (67%)
Frame = -2
Query: 2017 YGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEAANKNI 2068
+G+P +I+TDNG++ +N + CE ++I + +SP PQ NG EA+NK I
Sbjct: 685 HGLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKII 530
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 85.1 bits (209), Expect = 3e-16
Identities = 65/230 (28%), Positives = 109/230 (47%), Gaps = 5/230 (2%)
Frame = +2
Query: 1191 VEHQIPTKPECPPVRQKLRRTHPDMALKIKSEVQKQIDAGFLMTVEYPEWVANIVPVPKK 1250
++ I P P+ R +P +K +++ ++ GF+ YP V ++ + KK
Sbjct: 35 IDFGIDLLPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKK 211
Query: 1251 DGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDRE 1310
DG +RM +D+ LN + K +PLP ID L DN SK F +D G +Q ++ ED
Sbjct: 212 DGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQHRVIGEDVP 391
Query: 1311 KTSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSTDEEQH 1370
KT+F +G + VM FG N + M +F D + V V+ +D+++ S +E +H
Sbjct: 392 KTAFRIRYGHYEILVMSFG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDILIYSKNENEH 571
Query: 1371 VEYLTKMFERLRKYKLRLNPNKCTFG-----VRSGKLLGFIVSQKGIEVD 1415
+L + L+ L C V G ++S +G++VD
Sbjct: 572 ENHLRLALKVLKDIGL------CQISYV*ILVEVGFFSLHVISGEGLKVD 703
>AL375832
Length = 535
Score = 80.1 bits (196), Expect = 9e-15
Identities = 37/41 (90%), Positives = 39/41 (94%)
Frame = +3
Query: 1042 FVNAISEEATGSGLRPVFVTPGGIARDWNAIDVPSIMHVSE 1082
FVNAISEEATGSGLRP FVTPGGIA DW+AID+PSIMHVSE
Sbjct: 3 FVNAISEEATGSGLRPAFVTPGGIASDWDAIDIPSIMHVSE 125
Score = 34.7 bits (78), Expect(2) = 5e-05
Identities = 14/22 (63%), Positives = 16/22 (72%)
Frame = +1
Query: 1080 VSELNHNKPVEHSNPTVPPSFD 1101
+ LNHNKPVEHSN P+FD
Sbjct: 418 ICRLNHNKPVEHSNSMPLPNFD 483
Score = 32.3 bits (72), Expect(2) = 5e-05
Identities = 13/17 (76%), Positives = 14/17 (81%)
Frame = +3
Query: 1102 FPVYEAEDEEGDDIPYE 1118
FPVYE + EGDDIPYE
Sbjct: 483 FPVYEPKMREGDDIPYE 533
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 78.2 bits (191), Expect = 3e-14
Identities = 75/353 (21%), Positives = 141/353 (39%), Gaps = 15/353 (4%)
Frame = +2
Query: 1889 QLMHDVHDGTFGTHATGHTMSRKLLRAGYYWMTMEHDCYQYARKCHKCQIYADKIHVPPH 1948
+L+ + HD T H G + +++ ++W ++ R C C IH+
Sbjct: 176 KLVQESHDSTAAGHP-GRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVC----GGIHIWRQ 340
Query: 1949 ALNVMSSPWPF-----SMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYTNVTKQ 2003
A P P S +D I + P G +++ V +D +K V + +
Sbjct: 341 AKRGFLKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAE 520
Query: 2004 VVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEA 2063
A+ + +G+P I++D G+N + C + S+ Y PQ +G E
Sbjct: 521 ACAQRFLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTER 700
Query: 2064 ANKNIKRIVQKMVTTYKD-WHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEI 2122
N+ I+ +++ V +D W ++LP R SS GATPF + +G V P+
Sbjct: 701 WNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYH-VDPIPTVE 877
Query: 2123 PSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAMARGQSYQARMKTAFDKKVHPREFKV 2182
+ V+ E + + + D I+ + + A R ++ + + D+ ++V
Sbjct: 878 DTGGVVSEGEAAAQLLVKRMKDVTGFIQAEIVAAQQRSEASANKRRCPADR------YQV 1039
Query: 2183 GELV---LKRRISQQPDPRGKW------TPNYEGPYVVKKAFSGGALILTHMD 2226
G+ V + S +P + W + P+VV+ G H+D
Sbjct: 1040 GDKVWLNVSNYKSPRPSKKLDWLHHKYEVTRFVTPHVVELNVPGTVYPKFHVD 1198
>BE124481
Length = 538
Score = 74.3 bits (181), Expect = 5e-13
Identities = 42/102 (41%), Positives = 53/102 (51%), Gaps = 1/102 (0%)
Frame = +3
Query: 403 PPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQQQQQQQQYTRPT-FPPIPMLYAEL 461
PP Q P P QP P Q +Q +P Q Q+ Q Q + + PIP+ YA+L
Sbjct: 231 PPLVQQRPQSPYQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVKYADL 410
Query: 462 LPTLLQRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHD 503
LP LL++ T P+ LPP +R DL C FHQGA GHD
Sbjct: 411 LPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHD 536
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 66.2 bits (160), Expect = 1e-10
Identities = 38/109 (34%), Positives = 55/109 (49%)
Frame = +3
Query: 1678 GAVIVSPQGHHIPFTARILFECTNNMAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQI 1737
G + SP + + R+ F +NN YEA I G+ A ++I+++ Y DS LV +Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1738 KGEWETHHAKLIPYRDYARRLLTYFTKVELHHIPRDENQMADALATLSS 1786
GE+E + Y ++L L IPR EN ADALA L+S
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALAS 329
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 49.7 bits (117), Expect = 1e-05
Identities = 35/101 (34%), Positives = 52/101 (50%), Gaps = 3/101 (2%)
Frame = +1
Query: 2145 QLNLIEEKRMDAMARGQSYQARMKTAFDKKVHPREFKVGELVL--KRRISQQPDPRGKWT 2202
+LN +EE R+DA + Y+ R K D+++ REF+ GELVL R+ P GK
Sbjct: 103 ELNELEELRLDAYENAKIYKERTKKWHDRRIIRREFREGELVLLFNSRLKLFP---GKLR 273
Query: 2203 PNYEGPYVVKKAFSGGAL-ILTHMDGVELPNPVNADIVKKY 2242
++ GP+ VK GA+ + + G P VN +K Y
Sbjct: 274 SHWSGPFQVKNVMPSGAVEVWSESTG---PFTVNGQRLKHY 387
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 48.9 bits (115), Expect = 2e-05
Identities = 24/56 (42%), Positives = 31/56 (54%)
Frame = +1
Query: 1508 DESMGCVLGQQDETGKKEHAIYYLSKKFTDCETRYTMLEKTCCALAWAAKRLRHYL 1563
D +G L Q+DE EH I Y S+ T E YT++E+ C A WA + RHYL
Sbjct: 7 DWGIGAGLSQKDEENH-EHPIAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYL 171
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,686,613
Number of Sequences: 36976
Number of extensions: 991994
Number of successful extensions: 7178
Number of sequences better than 10.0: 104
Number of HSP's better than 10.0 without gapping: 5506
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6482
length of query: 2244
length of database: 9,014,727
effective HSP length: 111
effective length of query: 2133
effective length of database: 4,910,391
effective search space: 10473864003
effective search space used: 10473864003
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146556.9


