

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146553.6 + phase: 0 /pseudo
(799 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
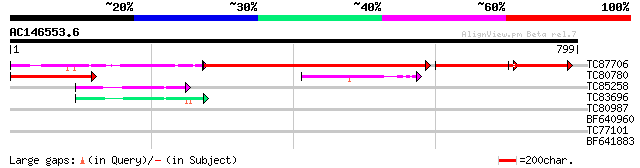
TC87706 weakly similar to GP|22137232|gb|AAM91461.1 AT5g18860/F1... 551 0.0
TC80780 similar to GP|22137232|gb|AAM91461.1 AT5g18860/F17K4_110... 219 2e-57
TC85258 similar to GP|21593497|gb|AAM65464.1 unknown {Arabidopsi... 58 2e-08
TC83696 similar to GP|21593497|gb|AAM65464.1 unknown {Arabidopsi... 51 2e-06
TC80987 similar to GP|15809830|gb|AAL06843.1 At2g36310/F2H17.8 {... 39 0.008
BF640960 similar to GP|22137232|gb| AT5g18860/F17K4_110 {Arabido... 35 0.15
TC77101 similar to GP|15148920|gb|AAK84887.1 homeodomain leucine... 29 6.2
BF641883 similar to GP|15983384|gb AT5g09860/MYH9_7 {Arabidopsis... 29 8.1
>TC87706 weakly similar to GP|22137232|gb|AAM91461.1 AT5g18860/F17K4_110
{Arabidopsis thaliana}, partial (53%)
Length = 1902
Score = 551 bits (1419), Expect(2) = 0.0
Identities = 266/322 (82%), Positives = 295/322 (91%), Gaps = 2/322 (0%)
Frame = +2
Query: 274 EYINITVITSNKPYGISDGSNPLFDGLKVPKFNLKKGGVHSGHIQQGLTDPFCFVKNGKG 333
EY+NITVITSNKPYGISDGSNPLF+GLKVPKFNL+KGGVHSGHIQQGL DP CFV+NGKG
Sbjct: 2 EYMNITVITSNKPYGISDGSNPLFNGLKVPKFNLEKGGVHSGHIQQGLRDPLCFVENGKG 181
Query: 334 RCQDGYTKEVNGQESVKVLVATKAKPNKDIRSSLDREYFKSFLNVLKQPQQAGKFNFTTQ 393
+CQDGYTKE G SV+VLVATKAKPN+D+ SSLDREYF FL+VLKQP+QAG++NFTTQ
Sbjct: 182 KCQDGYTKEEGGPGSVRVLVATKAKPNRDVGSSLDREYFIRFLDVLKQPRQAGRYNFTTQ 361
Query: 394 FPYYEEVTHKPDFWNKILGKPVVFDMDMSAGDFLALSYLLKVPVEVINLKAIIVSPTGWA 453
FPYY+EVT+KP+F NK LGKPVVFDMDMSAGDFLAL YLLKVPV+VI+LKAIIVSPTGWA
Sbjct: 362 FPYYKEVTYKPNFQNKKLGKPVVFDMDMSAGDFLALFYLLKVPVQVIDLKAIIVSPTGWA 541
Query: 454 NAATIDVIYDLLHMMGRDDIKVGIGDFFAMNQSN--FSPVGDCKYVKAIPHGNGGFLDSD 511
NAATID+IYD+LHMMGRDDI VG+GD FAMNQ + F VG CKYVKAIPHGNGG++DSD
Sbjct: 542 NAATIDIIYDILHMMGRDDIPVGLGDVFAMNQRDPIFGAVGGCKYVKAIPHGNGGYIDSD 721
Query: 512 TLFGLARDLPRSPRRYTAENSKKFGAPRDTDHPELRQPQAMEIWESILQTLKPGSKVTVL 571
TL+GLAR LPRSPRRYT ENS KFGAPRDTDHPELRQP AME+WES+LQT+KPGS +TVL
Sbjct: 722 TLYGLARYLPRSPRRYTGENSVKFGAPRDTDHPELRQPLAMEVWESVLQTMKPGSNITVL 901
Query: 572 TNGPLTNLAKVVSIKNISSIIQ 593
TNGPLTNLA VVS+KNISS IQ
Sbjct: 902 TNGPLTNLANVVSVKNISSRIQ 967
Score = 120 bits (300), Expect = 3e-27
Identities = 60/89 (67%), Positives = 73/89 (81%)
Frame = +2
Query: 704 GAVVLANGHSSLLDAKFELKSVKLLAEGIESTDGKMVVDEKYGKLVRILRHVDAKTYHEI 763
GAVVLA+ SS L+ KFE+K +K+LA GIESTDGK+VVDEK+GKLVRIL +V+ K Y+ +
Sbjct: 1298 GAVVLAD-RSSSLNPKFEVKPIKVLASGIESTDGKIVVDEKHGKLVRILSNVEEKAYYNM 1474
Query: 764 YAKRLGDPNQSAKVGSFKEQKRKWSHPHD 792
Y +LGD QSAKVGSF+EQ R WSHPHD
Sbjct: 1475 YVNKLGDLYQSAKVGSFEEQMRNWSHPHD 1561
Score = 104 bits (260), Expect(2) = 0.0
Identities = 58/116 (50%), Positives = 72/116 (62%)
Frame = +1
Query: 601 SGGLCNGRTYQQE*Q*QRKCIFCSFQQVCRIQYVLGSFSSKDSVSIRS*HHTHSTRYPAQ 660
SG C+G +QQ + QRKC+F SF +CRIQ+V SFS ++SV IRS* HTHS + A
Sbjct: 964 SGSFCSGGAHQQ*CRRQRKCVFSSF*PICRIQHVPRSFSRQNSVRIRS*DHTHSIEHSAP 1143
Query: 661 SKFIFKYFKLA*SDRKNTRSSVFQASSVKATPFEENPPQISPYGAVVLANGHSSLL 716
S+FI Y++ D+KN R V+Q S VK PFE N QI PYG V N S L
Sbjct: 1144 SEFICNYYR*IRRDKKNIRGCVYQESIVKPKPFEANQ*QILPYGHVPGRNSRCSSL 1311
Score = 90.5 bits (223), Expect = 2e-18
Identities = 84/298 (28%), Positives = 132/298 (44%), Gaps = 20/298 (6%)
Frame = +2
Query: 1 MMGRDDIAVGVGGEGGILPNGTILPNVGGYLPIIEQGMTTAGYCRYRQAIPVGFGGRLDI 60
MMGRDDI VG+G + I VGG C+Y +AIP G GG +D
Sbjct: 581 MMGRDDIPVGLGDVFAMNQRDPIFGAVGG--------------CKYVKAIPHGNGGYIDS 718
Query: 61 DTNLGIRKAFLPQGKRKYT----------------PLRQPTTQQV---LIDKISAGP-IT 100
DT G+ + +LP+ R+YT LRQP +V ++ + G IT
Sbjct: 719 DTLYGLAR-YLPRSPRRYTGENSVKFGAPRDTDHPELRQPLAMEVWESVLQTMKPGSNIT 895
Query: 101 LIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCGDTGNVL 160
++ G TNLA +++ ++ ++ V+++GG I S NA D GNV
Sbjct: 896 VLTNGPLTNLAN-VVSVKNISSRIQEVFVVGGHISS------NAE---------DKGNVF 1027
Query: 161 TNYNANPYAEYNIFGDPFAAYKVIHSGIPITLVPLDATNTIPISEEFFDEFEKSQDTYEA 220
+ +N YAE+N+F DP AA V S + ITL+PL + E ++ T E
Sbjct: 1028S-VPSNQYAEFNMFLDPLAAKTVFESEVKITLIPLSTQRQVSSFATIIGRLEGTRKTSEV 1204
Query: 221 QYCFKSLKMAHDTWFDNQFYTSYFMWDSFTSGVAVSIMRNSNRKKGENEFAEMEYINI 278
+ L + Q Y+ D+F + +++ ++R N E++ I +
Sbjct: 1205VFTKSLLSSLNRL---KQTNNRYYHMDTFLGEILGAVVL-ADRSSSLNPKFEVKPIKV 1366
>TC80780 similar to GP|22137232|gb|AAM91461.1 AT5g18860/F17K4_110
{Arabidopsis thaliana}, partial (19%)
Length = 657
Score = 219 bits (559), Expect = 2e-57
Identities = 104/122 (85%), Positives = 111/122 (90%)
Frame = +2
Query: 1 MMGRDDIAVGVGGEGGILPNGTILPNVGGYLPIIEQGMTTAGYCRYRQAIPVGFGGRLDI 60
MMGRDD+AVG+GGEGGIL NGTILPNVGGYLPIIEQGMTT G CRYRQAIPVG GGRLDI
Sbjct: 287 MMGRDDVAVGIGGEGGILSNGTILPNVGGYLPIIEQGMTTIGGCRYRQAIPVGLGGRLDI 466
Query: 61 DTNLGIRKAFLPQGKRKYTPLRQPTTQQVLIDKISAGPITLIVTGAHTNLAIFLMNNPHL 120
D N GIRK+FLPQGKRKYTPL QPT QQVLI+K+SAGP TL + GAHTN+AIFLMNNPHL
Sbjct: 467 DANYGIRKSFLPQGKRKYTPLEQPTAQQVLIEKVSAGPTTLFMMGAHTNVAIFLMNNPHL 646
Query: 121 KK 122
KK
Sbjct: 647 KK 652
Score = 89.4 bits (220), Expect = 5e-18
Identities = 61/189 (32%), Positives = 92/189 (48%), Gaps = 20/189 (10%)
Frame = +2
Query: 412 GKP--VVFDMDMSAGDFLALSYLLKVPVEVINLKAIIVSPTGWANAA-TIDVIYDLLHMM 468
GKP +V D D+ D AL YLLK+ L+A+ +S W +A ++ IYDLL+MM
Sbjct: 113 GKPQRIVVDTDVDTDDLFALLYLLKLNTSQFQLEAVTISANSWTSAGHAVNQIYDLLYMM 292
Query: 469 GRDDIKVGI----------------GDFFAMNQSNFSPVGDCKYVKAIPHGNGGFLDSDT 512
GRDD+ VGI G + + + + +G C+Y +AIP G GG LD D
Sbjct: 293 GRDDVAVGIGGEGGILSNGTILPNVGGYLPIIEQGMTTIGGCRYRQAIPVGLGGRLDIDA 472
Query: 513 LFGLARD-LPRSPRRYTAENSKKFGAPRDTDHPELRQPQAMEIWESILQTLKPGSKVTVL 571
+G+ + LP+ R+YT L QP A ++ +++ + G T+
Sbjct: 473 NYGIRKSFLPQGKRKYT----------------PLEQPTAQQV---LIEKVSAG-PTTLF 592
Query: 572 TNGPLTNLA 580
G TN+A
Sbjct: 593 MMGAHTNVA 619
>TC85258 similar to GP|21593497|gb|AAM65464.1 unknown {Arabidopsis
thaliana}, partial (92%)
Length = 1193
Score = 57.8 bits (138), Expect = 2e-08
Identities = 44/163 (26%), Positives = 70/163 (41%)
Frame = +2
Query: 93 KISAGPITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSK 152
K + G IT++ G TN+A+ + +P KN+ + ++GG
Sbjct: 293 KANPGKITVVALGPLTNIALAIQMDPEFAKNIGQIVLLGG-------------------- 412
Query: 153 CGDTGNVLTNYNANPYAEYNIFGDPFAAYKVIHSGIPITLVPLDATNTIPISEEFFDEFE 212
+ N N NP AE NIFGDP AA V SG I V ++ T+ + +S ++
Sbjct: 413 -----SFAVNGNVNPAAEANIFGDPDAADVVFTSGADILAVGINVTHQVVLSGSDREKLA 577
Query: 213 KSQDTYEAQYCFKSLKMAHDTWFDNQFYTSYFMWDSFTSGVAV 255
S+ + AQY + L++ D ++ D T AV
Sbjct: 578 SSKGKF-AQYLTQILEVYFSYHHDAYNTKGVYLHDPTTLLAAV 703
>TC83696 similar to GP|21593497|gb|AAM65464.1 unknown {Arabidopsis
thaliana}, partial (83%)
Length = 975
Score = 51.2 bits (121), Expect = 2e-06
Identities = 47/212 (22%), Positives = 82/212 (38%), Gaps = 24/212 (11%)
Frame = +1
Query: 93 KISAGPITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSK 152
K + G IT++ G TN+A+ + +P KN+ + ++GG
Sbjct: 274 KANRGKITVVALGPLTNIALAIQMDPEFSKNIGQIVLLGGAF------------------ 399
Query: 153 CGDTGNVLTNYNANPYAEYNIFGDPFAAYKVIHSGIPITLVPLDATNTIPISEEFFDEFE 212
N + NP AE NI GDP AA V SG + V ++ T + + ++
Sbjct: 400 -------AVNGSVNPSAETNILGDPDAADVVFTSGADVLAVGINVTQQVVFTGSDREKLA 558
Query: 213 KSQDTYEAQYCFKSLKMAHDTWFDNQFYTSYFMWD-------------SFTSGV------ 253
S+ + AQY L++ + ++ D +FT GV
Sbjct: 559 SSKGKF-AQYLNGILEVYFSNYCKAYNIKGIYLHDPIALLAVVDPSLVTFTEGVVRVQTN 735
Query: 254 ----AVSIMRNSNRKKGE-NEFAEMEYINITV 280
++I+ N ++ GE E+ M + + V
Sbjct: 736 GITRGLTILENKQKRFGEVTEWCNMPTVKVAV 831
>TC80987 similar to GP|15809830|gb|AAL06843.1 At2g36310/F2H17.8 {Arabidopsis
thaliana}, partial (76%)
Length = 1089
Score = 38.9 bits (89), Expect = 0.008
Identities = 19/52 (36%), Positives = 31/52 (59%)
Frame = +3
Query: 164 NANPYAEYNIFGDPFAAYKVIHSGIPITLVPLDATNTIPISEEFFDEFEKSQ 215
N NP AE NI+GDP AA V SG + +V ++ T + +++ E ++S+
Sbjct: 798 NVNPAAEANIYGDPEAADVVFTSGADVVVVGINITTQVQLTDADLLELKESK 953
>BF640960 similar to GP|22137232|gb| AT5g18860/F17K4_110 {Arabidopsis
thaliana}, partial (4%)
Length = 429
Score = 34.7 bits (78), Expect = 0.15
Identities = 18/50 (36%), Positives = 26/50 (52%), Gaps = 2/50 (4%)
Frame = +2
Query: 408 NKILGKP--VVFDMDMSAGDFLALSYLLKVPVEVINLKAIIVSPTGWANA 455
N + GKP ++ D D+ D AL YLLK+ L+ + +S W NA
Sbjct: 278 NIVEGKPQRILLDTDVDTDDLFALLYLLKLNRSQFLLQGVTLSANAWTNA 427
>TC77101 similar to GP|15148920|gb|AAK84887.1 homeodomain leucine zipper
protein HDZ3 {Phaseolus vulgaris}, complete
Length = 1532
Score = 29.3 bits (64), Expect = 6.2
Identities = 20/65 (30%), Positives = 28/65 (42%), Gaps = 4/65 (6%)
Frame = -1
Query: 492 GDCKYVKAIPHGNGGFLDSDTLFG----LARDLPRSPRRYTAENSKKFGAPRDTDHPELR 547
GDC + + + G L LF L DLPR PR NS PR + + R
Sbjct: 626 GDCSS*YSSSYSSSGELKKGLLFEPSSMLIIDLPRKPRSAFPRNSMFMLFPRQAEESK*R 447
Query: 548 QPQAM 552
+P ++
Sbjct: 446 RPDSI 432
>BF641883 similar to GP|15983384|gb AT5g09860/MYH9_7 {Arabidopsis thaliana},
partial (32%)
Length = 646
Score = 28.9 bits (63), Expect = 8.1
Identities = 12/24 (50%), Positives = 14/24 (58%)
Frame = +1
Query: 304 KFNLKKGGVHSGHIQQGLTDPFCF 327
+F L GG+ GHI G T FCF
Sbjct: 10 QFKLHNGGI*KGHITTGAT*KFCF 81
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,865,828
Number of Sequences: 36976
Number of extensions: 334871
Number of successful extensions: 1574
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1546
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1568
length of query: 799
length of database: 9,014,727
effective HSP length: 104
effective length of query: 695
effective length of database: 5,169,223
effective search space: 3592609985
effective search space used: 3592609985
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146553.6


