

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
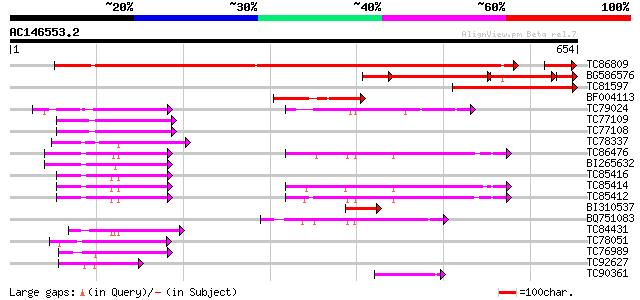
Query= AC146553.2 + phase: 0 /pseudo
(654 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86809 homologue to PIR|S35701|S35701 translation elongation fa... 426 e-125
BG586576 similar to GP|10129885|db mitochondrial elongation fact... 221 9e-72
TC81597 similar to PIR|D96510|D96510 probable mitochondrial elon... 263 2e-70
BF004113 weakly similar to GP|13957623|gb| mitochondrial elongat... 109 3e-24
TC79024 similar to PIR|A84716|A84716 probable GTP-binding protei... 90 3e-18
TC77109 similar to GP|9758030|dbj|BAB08691.1 GTP-binding protein... 87 3e-17
TC77108 similar to GP|9758030|dbj|BAB08691.1 GTP-binding protein... 87 3e-17
TC78337 similar to GP|9759359|dbj|BAB10014.1 GTP-binding protein... 83 3e-16
TC86476 similar to GP|15450763|gb|AAK96653.1 elongation factor E... 79 6e-15
BI265632 homologue to GP|22947038|gb CG2238-PA {Drosophila melan... 78 9e-15
TC85416 homologue to GP|15450763|gb|AAK96653.1 elongation factor... 77 3e-14
TC85414 homologue to GP|15450763|gb|AAK96653.1 elongation factor... 77 3e-14
TC85412 homologue to GP|15450763|gb|AAK96653.1 elongation factor... 76 4e-14
BI310537 similar to GP|10129885|dbj mitochondrial elongation fac... 65 6e-11
BQ751083 homologue to GP|13925370|gb elongation factor 2 {Neuros... 65 8e-11
TC84431 similar to GP|10176983|dbj|BAB10215. GTP-binding membran... 64 2e-10
TC78051 homologue to PIR|T01400|T01400 translation elongation fa... 56 4e-08
TC76989 homologue to SP|O24310|EFTU_PEA Elongation factor Tu ch... 53 3e-07
TC92627 similar to PIR|T49975|T49975 hypothetical protein F12B17... 50 2e-06
TC90361 similar to PIR|S35701|S35701 translation elongation fact... 48 1e-05
>TC86809 homologue to PIR|S35701|S35701 translation elongation factor EF-G
chloroplast - soybean, partial (91%)
Length = 2674
Score = 426 bits (1096), Expect(2) = e-125
Identities = 238/540 (44%), Positives = 331/540 (61%), Gaps = 4/540 (0%)
Frame = +2
Query: 52 MEKVRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITI 111
++ RNIGI AHID+GKTT TE ILFYTG+ + + EV MD+ E GITI
Sbjct: 308 LKDYRNIGIMAHIDAGKTTTTERILFYTGRNYKIGEVHEGTAT---MDWMEQEQERGITI 478
Query: 112 KSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQM 171
SAAT W +I IIDTPGHVDFT+EVERALRVLDGA+ + SV GV+ QS TV RQ
Sbjct: 479 TSAATTTFWDNHRINIIDTPGHVDFTLEVERALRVLDGAICLFDSVAGVEPQSETVWRQA 658
Query: 172 RRYQVPRIAFINKLDRPGADPWKVITQARSKLRHHCAALQVPIGLESDFKGVVDLVKLKA 231
RY VPRI F+NK+DR GA+ ++ + L LQ+PIG E FKGV+DLV++KA
Sbjct: 659 DRYGVPRICFVNKMDRLGANFFRTRDMIVTNLGAKPLVLQLPIGAEDSFKGVIDLVRMKA 838
Query: 232 YCFDG-QYGQNVVVGEVPADMEALVAEKRRELIETVSEVDDVLAEAFLSDDENISAADLE 290
+ G + G ++P D+ + R ++IET+ E+DD E +L E A ++
Sbjct: 839 IVWGGEELGAKFTYEDIPVDLLEQAQDYRSQMIETIVELDDEAMENYLEGVEP-DEATIK 1015
Query: 291 GAIRRATIARKFIPVFMGSAVKNTGVQPLLDGVVSYLPCPIEVSNY-ALDQSKNEEKVQL 349
IR+ +IA F+PV GSA KN GVQPLLD VV YLP P++V D E ++
Sbjct: 1016KLIRKGSIAATFVPVMCGSAFKNKGVQPLLDAVVDYLPSPLDVPPMKGTDPENPEATIER 1195
Query: 350 TGSPDGPLVALAFKLEQTKF-GQLTYLRVYEGVIRKGDFIVNVSTGKKIKVPRLVQMHSN 408
D P LAFK+ F G LT++RVY G + G +++N + GKK ++ RL++MH+N
Sbjct: 1196IAGDDEPFSGLAFKIMSDSFVGSLTFVRVYSGKLTAGSYVLNSNKGKKERIGRLLEMHAN 1375
Query: 409 EMNDIEEAHAGQIVAVFGV-DCASSDTFTDGSVKYTMTSMNVPEPVMSLAVQPVSKDSGG 467
D++ A G IVA+ G+ D + +T D + M+ P+PV+ +A++P +K
Sbjct: 1376SREDVKVALTGDIVALAGLKDTITGETLCDPESPVVLERMDFPDPVIKIAIEPKTKADID 1555
Query: 468 KFSKALNRFQREDPTFRVSLDPESGQTIISGMGELHLDIYVKRIKMEYGVDATVGKPRVN 527
K + L + +EDP+F S D E QT+I GMGELHL+I V R+K EY V+A VG P+VN
Sbjct: 1556KMAAGLVKLAQEDPSFHFSRDEEINQTVIEGMGELHLEIIVDRLKREYKVEANVGAPQVN 1735
Query: 528 FRETVTQRADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTKFEFDNMLVGQAIPSNFFP 587
+RE++++ + Y+HKKQSGGQGQ+ + EP+ GSG +EF + + G A+P + P
Sbjct: 1736YRESISKIHEARYVHKKQSGGQGQFADITVRFEPMEPGSG--YEFKSEIKGGAVPKEYIP 1909
Score = 41.2 bits (95), Expect(2) = e-125
Identities = 21/37 (56%), Positives = 27/37 (72%)
Frame = +1
Query: 617 GAAHDVDSSELAFKLASIYAFRECYTASRPVILEPVM 653
G+ HDVDSS LAF+LA+ AFRE + P +LEP+M
Sbjct: 1999 GSYHDVDSSVLAFQLAARGAFREGIRKAGPRMLEPIM 2109
>BG586576 similar to GP|10129885|db mitochondrial elongation factor G {Oryza
sativa (japonica cultivar-group)}, partial (30%)
Length = 745
Score = 221 bits (563), Expect(2) = 9e-72
Identities = 109/120 (90%), Positives = 114/120 (94%)
Frame = +1
Query: 438 GSVKYTMTSMNVPEPVMSLAVQPVSKDSGGKFSKALNRFQREDPTFRVSLDPESGQTIIS 497
G + TMTSM+VPEPVMSLAVQPVSKDSGG+FSKALNRFQREDPTFRV LDPESGQTIIS
Sbjct: 94 GLLNITMTSMSVPEPVMSLAVQPVSKDSGGQFSKALNRFQREDPTFRVGLDPESGQTIIS 273
Query: 498 GMGELHLDIYVKRIKMEYGVDATVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGRVIG 557
GMGELHLDIYV+RI+ EY VDATVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGRVIG
Sbjct: 274 GMGELHLDIYVERIRREYKVDATVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGRVIG 453
Score = 114 bits (285), Expect(2) = 3e-33
Identities = 61/83 (73%), Positives = 66/83 (79%), Gaps = 7/83 (8%)
Frame = +2
Query: 555 VIGYIEPLPAGSG-------TKFEFDNMLVGQAIPSNFFPAIEKGFKEAANSGALIGHPV 607
+I YI P G TKFEF+NMLVGQAIPSNFF AIEKGF EAANSG+LIGHPV
Sbjct: 395 LIIYIRSNPEGKDNMDG*LETKFEFENMLVGQAIPSNFFAAIEKGFIEAANSGSLIGHPV 574
Query: 608 QNLRVVLTDGAAHDVDSSELAFK 630
+NLRVVLTDGAAH VDSSELAF+
Sbjct: 575 ENLRVVLTDGAAHAVDSSELAFR 643
Score = 68.2 bits (165), Expect(2) = 9e-72
Identities = 31/35 (88%), Positives = 33/35 (93%)
Frame = +3
Query: 408 NEMNDIEEAHAGQIVAVFGVDCASSDTFTDGSVKY 442
NEM +I+EAHAGQIVAVFGVDCAS DTFTDGSVKY
Sbjct: 3 NEMEEIDEAHAGQIVAVFGVDCASGDTFTDGSVKY 107
Score = 46.2 bits (108), Expect(2) = 3e-33
Identities = 20/24 (83%), Positives = 23/24 (95%)
Frame = +1
Query: 631 LASIYAFRECYTASRPVILEPVML 654
+ASIYAFR+CYTASRP IL+PVML
Sbjct: 643 MASIYAFRQCYTASRPTILKPVML 714
>TC81597 similar to PIR|D96510|D96510 probable mitochondrial elongation
factor [imported] - Arabidopsis thaliana, partial (31%)
Length = 1065
Score = 263 bits (672), Expect = 2e-70
Identities = 130/144 (90%), Positives = 136/144 (94%)
Frame = +2
Query: 511 IKMEYGVDATVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTKF 570
I+ EY DATVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGRVIGYIEPLPAGS TKF
Sbjct: 2 IRREYRADATVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSETKF 181
Query: 571 EFDNMLVGQAIPSNFFPAIEKGFKEAANSGALIGHPVQNLRVVLTDGAAHDVDSSELAFK 630
EF+NMLVGQAIPSNFF AIEKGF EAANSG+LIGHPV+NLRVVLTDGAAH VDSSELAFK
Sbjct: 182 EFENMLVGQAIPSNFFAAIEKGFIEAANSGSLIGHPVENLRVVLTDGAAHAVDSSELAFK 361
Query: 631 LASIYAFRECYTASRPVILEPVML 654
+ASIYAFR+CYTASRP ILEPVML
Sbjct: 362 MASIYAFRQCYTASRPTILEPVML 433
>BF004113 weakly similar to GP|13957623|gb| mitochondrial elongation factor G
{Oryza sativa}, partial (11%)
Length = 585
Score = 109 bits (273), Expect = 3e-24
Identities = 60/107 (56%), Positives = 78/107 (72%), Gaps = 1/107 (0%)
Frame = -2
Query: 305 VFMGS-AVKNTGVQPLLDGVVSYLPCPIEVSNYALDQSKNEEKVQLTGSPDGPLVALAFK 363
VFMG A K G+Q LLDGV +YLPCP EVSNY+LD+SKN EK LVALAF
Sbjct: 584 VFMGGYAFKYKGLQLLLDGVTNYLPCPTEVSNYSLDKSKNGEK--------DILVALAFT 429
Query: 364 LEQTKFGQLTYLRVYEGVIRKGDFIVNVSTGKKIKVPRLVQMHSNEM 410
L++ + GQ+TYLR+YEGVIRKGDFI +++ GK +VP L+ + ++E+
Sbjct: 428 LKE-RVGQITYLRIYEGVIRKGDFITDLNFGKTFEVPHLIGLRNDEV 291
>TC79024 similar to PIR|A84716|A84716 probable GTP-binding protein
[imported] - Arabidopsis thaliana, partial (67%)
Length = 1513
Score = 89.7 bits (221), Expect = 3e-18
Identities = 60/165 (36%), Positives = 92/165 (55%), Gaps = 3/165 (1%)
Frame = +2
Query: 27 RHFSAGNVARAT---AATIDKDPWWKESMEKVRNIGISAHIDSGKTTLTEWILFYTGKIH 83
R FS+ + A AT AA++D + ++RN+ + AH+D GKTTL + +L G
Sbjct: 134 RFFSSSSAAAATSSPAASLDPN--------RLRNVAVIAHVDHGKTTLMDRLLRQCGA-D 286
Query: 84 LMYEVRSKDGMGPKMDFKPLEIIMGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERA 143
+ +E MD LE GITI S T +WK +++ ++DTPGH DF EVER
Sbjct: 287 IPHE--------RAMDSISLERERGITISSKVTSISWKDNELNMVDTPGHADFGGEVERV 442
Query: 144 LRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQVPRIAFINKLDRP 188
+ +++GAVLV+ + G Q+ V + +Y + I +NK+DRP
Sbjct: 443 VGMVEGAVLVVDAGEGPLAQTKFVLAKALKYGLRPILLLNKVDRP 577
Score = 51.6 bits (122), Expect = 9e-07
Identities = 56/238 (23%), Positives = 104/238 (43%), Gaps = 19/238 (7%)
Frame = +2
Query: 319 LLDGVVSYLPCPIEVSNYALDQSKNEEKVQLTGSPDGPLVALAFKLEQTK-FGQLTYLRV 377
LLD +VS++P P + D P L +E+ FG++ RV
Sbjct: 755 LLDAIVSHVPPP-------------------NANIDAPFQMLVSMMEKDNYFGRILTGRV 877
Query: 378 YEGVIRKGDFIVNV----STGKKI---KVPRLVQMHSNEMNDIEEAHAGQIVAVFGVDCA 430
+ GV+R GD + + S +KI KV ++++ M + A AG I+++ G+
Sbjct: 878 HSGVVRVGDKVHGLRNKDSGAEKIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSP 1057
Query: 431 S-SDTFTDGSVKYTMTSMNVPEPVMS---------LAVQPVSKDSGGKFSKALNRFQRED 480
S T T + + ++ + P +S LA + + +GGK L +
Sbjct: 1058SIGHTVTTVEIMSALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGKIGDRL--MAESE 1231
Query: 481 PTFRVSLDPESGQTI-ISGMGELHLDIYVKRIKMEYGVDATVGKPRVNFRETVTQRAD 537
+++ P +T + G GEL L I ++ ++ E G + +V P+V ++ Q+ +
Sbjct: 1232TNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSPPKVMYKTDKGQKLE 1402
>TC77109 similar to GP|9758030|dbj|BAB08691.1 GTP-binding protein typA
(tyrosine phosphorylated protein A) {Arabidopsis
thaliana}, partial (27%)
Length = 672
Score = 86.7 bits (213), Expect = 3e-17
Identities = 56/138 (40%), Positives = 76/138 (54%)
Frame = +3
Query: 55 VRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSA 114
+RNI I AH+D GKTTL + +L T V+ + MD LE GITI S
Sbjct: 186 IRNIAIVAHVDHGKTTLVDAMLKQTKVFRDNQTVQERI-----MDSNDLERERGITILSK 350
Query: 115 ATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRY 174
T +K +KI IIDTPGH DF EVER L +++G +LV+ SV G Q+ V ++ +
Sbjct: 351 NTSVTYKDAKINIIDTPGHSDFGGEVERILNMVEGILLVVDSVEGPMPQTRFVLKKALEF 530
Query: 175 QVPRIAFINKLDRPGADP 192
+ +NK+DR A P
Sbjct: 531 GHAVVVVVNKIDRSSARP 584
>TC77108 similar to GP|9758030|dbj|BAB08691.1 GTP-binding protein typA
(tyrosine phosphorylated protein A) {Arabidopsis
thaliana}, partial (98%)
Length = 2589
Score = 86.7 bits (213), Expect = 3e-17
Identities = 56/138 (40%), Positives = 76/138 (54%)
Frame = +3
Query: 55 VRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSA 114
+RNI I AH+D GKTTL + +L T V+ + MD LE GITI S
Sbjct: 396 IRNIAIVAHVDHGKTTLVDAMLKQTKVFRDNQTVQERI-----MDSNDLERERGITILSK 560
Query: 115 ATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRY 174
T +K +KI IIDTPGH DF EVER L +++G +LV+ SV G Q+ V ++ +
Sbjct: 561 NTSVTYKDAKINIIDTPGHSDFGGEVERILNMVEGILLVVDSVEGPMPQTRFVLKKALEF 740
Query: 175 QVPRIAFINKLDRPGADP 192
+ +NK+DR A P
Sbjct: 741 GHAVVVVVNKIDRSSARP 794
Score = 36.6 bits (83), Expect = 0.031
Identities = 30/126 (23%), Positives = 57/126 (44%), Gaps = 9/126 (7%)
Frame = +2
Query: 444 MTSMNVPEPVMSLAVQPVSKDSGGKFSKALNRFQREDPTFR-----VSLDPESGQT---- 494
+ S+ V EP + ++ + G+ K + D +R +++ E G+T
Sbjct: 1274 LPSIKVEEPTVKMSFSINTSPFVGREGKYVTSRNLRDRLYRELERNLAMKVEDGETADTF 1453
Query: 495 IISGMGELHLDIYVKRIKMEYGVDATVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGR 554
I+SG G LH+ I ++ ++ E G + VG P+V + ++ D L ++ G GR
Sbjct: 1454 IVSGRGTLHITILIENMRRE-GYEFMVGPPKV-----INKKVDDKVLEPYENATCGATGR 1615
Query: 555 VIGYIE 560
I+
Sbjct: 1616 KYSIID 1633
>TC78337 similar to GP|9759359|dbj|BAB10014.1 GTP-binding protein LepA
homolog {Arabidopsis thaliana}, partial (40%)
Length = 1035
Score = 83.2 bits (204), Expect = 3e-16
Identities = 56/164 (34%), Positives = 85/164 (51%), Gaps = 4/164 (2%)
Frame = +1
Query: 49 KESMEKVRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMG 108
K + +RN I AHID GK+TL + +L TG + + K+ MD LE G
Sbjct: 241 KVPVSNIRNFSIIAHIDHGKSTLADKLLQVTGTVP---QREMKEQFLDNMD---LERERG 402
Query: 109 ITIKSAATYCNWKGSK----ITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQS 164
ITIK A + + +IDTPGHVDF+ EV R+L +GA+LV+ + GV+ Q+
Sbjct: 403 ITIKLQAARMRYVFENEPYCLNLIDTPGHVDFSYEVSRSLAACEGALLVVDASQGVEAQT 582
Query: 165 ITVDRQMRRYQVPRIAFINKLDRPGADPWKVITQARSKLRHHCA 208
+ + I +NK+D PGA+P +V+ + + C+
Sbjct: 583 LANVYLALDNNLEIIPVLNKIDLPGAEPDRVLKEIEEVIGLDCS 714
>TC86476 similar to GP|15450763|gb|AAK96653.1 elongation factor EF-2
{Arabidopsis thaliana}, complete
Length = 2974
Score = 79.0 bits (193), Expect = 6e-15
Identities = 56/163 (34%), Positives = 84/163 (51%), Gaps = 16/163 (9%)
Frame = +2
Query: 41 TIDKDPWWKESMEKVRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDF 100
T+D+ + +RN+ + AH+D GK+TLT+ ++ G I + G D
Sbjct: 257 TVDELRRIMDLKHNIRNMSVIAHVDHGKSTLTDSLVAAAGII-----AQEVAGDVRMTDT 421
Query: 101 KPLEIIMGITIKSAATYC----------NWKGSK------ITIIDTPGHVDFTIEVERAL 144
+ E GITIKS N+KG + I +ID+PGHVDF+ EV AL
Sbjct: 422 RQDEAERGITIKSTGISLYYEMSDGDLKNFKGEREGNKYLINLIDSPGHVDFSSEVTAAL 601
Query: 145 RVLDGAVLVLCSVGGVQCQSITVDRQMRRYQVPRIAFINKLDR 187
R+ DGA++V+ V GV Q+ TV RQ ++ + +NK+DR
Sbjct: 602 RITDGALVVVDCVEGVCVQTETVLRQALGERIKPVLTVNKMDR 730
Score = 63.9 bits (154), Expect = 2e-10
Identities = 71/284 (25%), Positives = 122/284 (42%), Gaps = 24/284 (8%)
Frame = +2
Query: 319 LLDGVVSYLPCPIEVSNYALDQSKNEEKVQLTGS------PDGPLVALAFKL--EQTKFG 370
LL+ ++ +LP P + Y ++ S P+GPL+ K+ K
Sbjct: 1238 LLEMMIFHLPSPTKAQKYRVENLYEGPLDDPYASAIRNCDPEGPLMLYVSKMIPASDKGR 1417
Query: 371 QLTYLRVYEGVIRKGDFI----VNVSTGKKI-----KVPRLVQMHSNEMNDIEEAHAGQI 421
+ RV+ G + G + N G+K V R V + +E+ G
Sbjct: 1418 FYAFGRVFSGKVSTGMKVRIMGPNYIPGEKKDLYVKSVQRTVIWMGKKQETVEDVPCGNT 1597
Query: 422 VAVFGVDCASSDTFTDGSVK----YTMTSMNVP-EPVMSLAVQPVSKDSGGKFSKALNRF 476
VA+ G+D + T + K + + +M PV+S+AV K + L R
Sbjct: 1598 VAMVGLDQFITKNATLTNEKEVDAHPIRAMKFSVSPVVSVAVTCKVASDLPKLVEGLKRL 1777
Query: 477 QREDPTFRVSLDPESGQTIISGMGELHLDIYVKRIKMEY--GVDATVGKPRVNFRETVTQ 534
+ DP ++ E+G+ II+ GELHL+I +K ++ ++ G + T P V+FRETV +
Sbjct: 1778 AKSDPMVVCTIS-ETGEHIIAAAGELHLEICLKDLQDDFMNGAEITKSDPIVSFRETVLE 1954
Query: 535 RADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTKFEFDNMLVG 578
++ H S ++ R+ Y+E P G D+ +G
Sbjct: 1955 KSS----HTVMSKSPNKHNRL--YMEARPMEEGLAEAIDDGRIG 2068
>BI265632 homologue to GP|22947038|gb CG2238-PA {Drosophila melanogaster},
partial (21%)
Length = 593
Score = 78.2 bits (191), Expect = 9e-15
Identities = 57/167 (34%), Positives = 83/167 (49%), Gaps = 20/167 (11%)
Frame = +3
Query: 41 TIDKDPWWKESMEKVRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDF 100
T+D+ + +RN+ + AH+D GK+TLT+ ++ G I ++ G D
Sbjct: 69 TVDEIRQMMDKKRNIRNMSVIAHVDHGKSTLTDSLVSKAGII-----AGARAGETRFTDT 233
Query: 101 KPLEIIMGITIKSAATYCNW--------------------KGSKITIIDTPGHVDFTIEV 140
+ E ITIKS A + KG I +ID+PGHVDF+ EV
Sbjct: 234 RKDEQDRCITIKSTAISMFFELEEKDLVFITNPDQREKSEKGFLINLIDSPGHVDFSSEV 413
Query: 141 ERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQVPRIAFINKLDR 187
ALRV DGA++V+ V GV Q+ TV RQ ++ F+NK+DR
Sbjct: 414 TAALRVTDGALVVVDCVSGVCVQTETVLRQAIAERIKPXLFMNKMDR 554
>TC85416 homologue to GP|15450763|gb|AAK96653.1 elongation factor EF-2
{Arabidopsis thaliana}, partial (21%)
Length = 660
Score = 76.6 bits (187), Expect = 3e-14
Identities = 53/149 (35%), Positives = 80/149 (53%), Gaps = 16/149 (10%)
Frame = +3
Query: 55 VRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSA 114
+RN+ + AH+D GK+TLT+ ++ G I + G D + E GITIKS
Sbjct: 165 IRNMSVIAHVDHGKSTLTDSLVAAAGII-----AQEVAGDVRMTDTRADEAERGITIKST 329
Query: 115 A----------TYCNWKGSK------ITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVG 158
+ ++KG + I +ID+PGHVDF+ EV ALR+ DGA++V+ V
Sbjct: 330 GISLYYEMSDESLKSFKGERNGNEYLINLIDSPGHVDFSSEVTAALRITDGALVVVDCVE 509
Query: 159 GVQCQSITVDRQMRRYQVPRIAFINKLDR 187
GV Q+ TV RQ ++ + +NK+DR
Sbjct: 510 GVCVQTETVLRQALGERIRPVLTVNKMDR 596
>TC85414 homologue to GP|15450763|gb|AAK96653.1 elongation factor EF-2
{Arabidopsis thaliana}, complete
Length = 2892
Score = 76.6 bits (187), Expect = 3e-14
Identities = 53/149 (35%), Positives = 80/149 (53%), Gaps = 16/149 (10%)
Frame = +3
Query: 55 VRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSA 114
+RN+ + AH+D GK+TLT+ ++ G I + G D + E GITIKS
Sbjct: 180 IRNMSVIAHVDHGKSTLTDSLVAAAGII-----AQEVAGDVRMTDTRADEAERGITIKST 344
Query: 115 A----------TYCNWKGSK------ITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVG 158
+ ++KG + I +ID+PGHVDF+ EV ALR+ DGA++V+ V
Sbjct: 345 GISLYYEMTDDSLKSFKGERNGNEYLINLIDSPGHVDFSSEVTAALRITDGALVVVDCVE 524
Query: 159 GVQCQSITVDRQMRRYQVPRIAFINKLDR 187
GV Q+ TV RQ ++ + +NK+DR
Sbjct: 525 GVCVQTETVLRQALGERIRPVLTVNKMDR 611
Score = 63.5 bits (153), Expect = 2e-10
Identities = 69/284 (24%), Positives = 121/284 (42%), Gaps = 24/284 (8%)
Frame = +3
Query: 319 LLDGVVSYLPCPIEVSNYALDQ------SKNEEKVQLTGSPDGPLVALAFKL--EQTKFG 370
LL+ ++ +LP P Y ++ P+GPL+ K+ K
Sbjct: 1119 LLEMMIFHLPSPSTAQRYRVENLYEGPLDDQYATAIRNCDPEGPLMLYVSKMIPASDKGR 1298
Query: 371 QLTYLRVYEGVIRKG--------DFIVNVSTGKKIK-VPRLVQMHSNEMNDIEEAHAGQI 421
+ RV+ G + G +F+ +K V R V +E+ G
Sbjct: 1299 FFAFGRVFSGKVSTGLKVRIMGPNFVPGEKKDLYVKSVQRTVIWMGKRQETVEDVPCGNT 1478
Query: 422 VAVFGVDCASSDTFTDGSVK----YTMTSMNVP-EPVMSLAVQPVSKDSGGKFSKALNRF 476
VA+ G+D + T + K + + +M PV+ +AVQ K + L R
Sbjct: 1479 VAMVGLDQFITKNATLTNEKEVDAHPIRAMKFSVSPVVRVAVQCKVASDLPKLVEGLKRL 1658
Query: 477 QREDPTFRVSLDPESGQTIISGMGELHLDIYVKRIKMEY--GVDATVGKPRVNFRETVTQ 534
+ DP +++ ESG+ I++G GELHL+I +K ++ ++ G + P V+FRETV +
Sbjct: 1659 AKSDPMVVCTIE-ESGEHIVAGAGELHLEICLKDLQDDFMGGAEIIKSDPVVSFRETVLE 1835
Query: 535 RADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTKFEFDNMLVG 578
R+ + K + ++ R+ Y+E P G D+ +G
Sbjct: 1836 RSCRTVMSKSPN----KHNRL--YMEARPLEDGLAEAIDDGKIG 1949
>TC85412 homologue to GP|15450763|gb|AAK96653.1 elongation factor EF-2
{Arabidopsis thaliana}, complete
Length = 3229
Score = 76.3 bits (186), Expect = 4e-14
Identities = 53/149 (35%), Positives = 79/149 (52%), Gaps = 16/149 (10%)
Frame = +3
Query: 55 VRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSA 114
+RN+ + AH+D GK+TLT+ ++ G I + G D + E GITIKS
Sbjct: 222 IRNMSVIAHVDHGKSTLTDSLVAAAGII-----AQEVAGDVRMTDTRADEAERGITIKST 386
Query: 115 A----------TYCNWKGSK------ITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVG 158
+ +KG + I +ID+PGHVDF+ EV ALR+ DGA++V+ V
Sbjct: 387 GISLYYEMTDESLKRFKGERNGNEYLINLIDSPGHVDFSSEVTAALRITDGALVVVDCVE 566
Query: 159 GVQCQSITVDRQMRRYQVPRIAFINKLDR 187
GV Q+ TV RQ ++ + +NK+DR
Sbjct: 567 GVCVQTETVLRQALGERIRPVLTVNKMDR 653
Score = 62.8 bits (151), Expect = 4e-10
Identities = 73/287 (25%), Positives = 121/287 (41%), Gaps = 27/287 (9%)
Frame = +3
Query: 319 LLDGVVSYLPCPIEVSNYAL---------DQSKNEEKVQLTGSPDGPLVALAFKL--EQT 367
LL+ ++ +LP P Y + DQ N + P+GPL+ K+
Sbjct: 1161 LLEMMIFHLPSPSTAQRYRVENLYEGPLDDQYANAIR---NCDPEGPLMLYVSKMIPASD 1331
Query: 368 KFGQLTYLRVYEGVIRKGDFI----VNVSTGKKI-----KVPRLVQMHSNEMNDIEEAHA 418
K + RV+ G + G + N G+K V R V +E+
Sbjct: 1332 KGRFFAFGRVFAGKVSTGLKVRIMGPNYIPGEKKDLYTKSVQRTVIWMGKRQETVEDVPC 1511
Query: 419 GQIVAVFGVDCASSDTFTDGSVK----YTMTSMNVP-EPVMSLAVQPVSKDSGGKFSKAL 473
G VA+ G+D + T + K + + +M PV+ +AVQ K + L
Sbjct: 1512 GNTVALVGLDQFITKNATLTNEKEVDAHPIRAMKFSVSPVVRVAVQCKVASDLPKLVEGL 1691
Query: 474 NRFQREDPTFRVSLDPESGQTIISGMGELHLDIYVKRIKMEY--GVDATVGKPRVNFRET 531
R + DP +++ ESG+ I++G GELHL+I +K ++ ++ G + P V+FRET
Sbjct: 1692 KRLAKSDPMVVCTIE-ESGEHIVAGAGELHLEICLKDLQDDFMGGAEIIKSDPVVSFRET 1868
Query: 532 VTQRADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTKFEFDNMLVG 578
V R+ + S ++ R+ Y+E P G D +G
Sbjct: 1869 VLDRS----VRTVMSKSPNKHNRL--YMEARPLEEGLAEAIDEGTIG 1991
>BI310537 similar to GP|10129885|dbj mitochondrial elongation factor G {Oryza
sativa (japonica cultivar-group)}, partial (5%)
Length = 124
Score = 65.5 bits (158), Expect = 6e-11
Identities = 31/41 (75%), Positives = 36/41 (87%)
Frame = +1
Query: 388 IVNVSTGKKIKVPRLVQMHSNEMNDIEEAHAGQIVAVFGVD 428
I+NV+TGKK KVP L +MHSNEM +I+EAHAGQI AVFGVD
Sbjct: 1 IINVNTGKKNKVPLLGRMHSNEMEEIDEAHAGQIFAVFGVD 123
>BQ751083 homologue to GP|13925370|gb elongation factor 2 {Neurospora
crassa}, partial (28%)
Length = 721
Score = 65.1 bits (157), Expect = 8e-11
Identities = 63/237 (26%), Positives = 102/237 (42%), Gaps = 20/237 (8%)
Frame = +3
Query: 290 EGAIRRATIARKFIPVFMGSAVKNTGVQPLLDGVVSYLPCPIEVSNY---ALDQSKNEEK 346
EG + R F+P LL+ ++ +LP P+ Y L + +++
Sbjct: 21 EGKQLLKAVMRTFLPA----------ADALLEMMIIHLPSPVTAQKYRSETLYEGPPDDE 170
Query: 347 VQLT---GSPDGPLVALAFKLEQTKFGQLTYL--RVYEGVIRKGDFI----VNVSTGKK- 396
+ P GPL+ K+ T Y RV+ G +R G + N GKK
Sbjct: 171 AAIAIRDCDPKGPLMLYVSKMVPTSDKGRFYAFGRVFAGTVRSGLKVRIQGPNYVPGKKE 350
Query: 397 ---IK-VPRLVQMHSNEMNDIEEAHAGQIVAVFGVD--CASSDTFTDGSVKYTMTSMNVP 450
IK + R V M ++ I++ AG IV + G+D S T T + + M
Sbjct: 351 DLFIKAIQRTVLMMGGKVEPIDDMPAGNIVGLVGIDQFLLKSGTLTTSDTAHNLKVMKFS 530
Query: 451 -EPVMSLAVQPVSKDSGGKFSKALNRFQREDPTFRVSLDPESGQTIISGMGELHLDI 506
PV+ +VQ + K + L R + DP +++ ESG+ +++G GELHL+I
Sbjct: 531 VSPVVQRSVQVKNAQDLPKLVEGLKRLSKSDPCV-LTMTNESGEHVVAGAGELHLEI 698
>TC84431 similar to GP|10176983|dbj|BAB10215. GTP-binding membrane protein
LepA homolog {Arabidopsis thaliana}, partial (46%)
Length = 976
Score = 63.9 bits (154), Expect = 2e-10
Identities = 52/150 (34%), Positives = 74/150 (48%), Gaps = 17/150 (11%)
Frame = +3
Query: 69 TTLTEWILFYTGKIHLMYEVRSKDGMGPK--MDFKPLEIIMGITIKSAAT---YCN---- 119
TTL + +L TG I K G+G +D +E GIT+K+ Y N
Sbjct: 3 TTLADRLLELTGTI--------KKGLGQPQYLDKLQVERERGITVKAQTATMFYKNIING 158
Query: 120 --WKGSK------ITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQM 171
+K K + +IDTPGHVDF+ EV R+L G +LV+ + GVQ Q++
Sbjct: 159 DDFKDGKESSNYLLNLIDTPGHVDFSYEVSRSLAACQGVLLVVDAAQGVQAQTVANFYLA 338
Query: 172 RRYQVPRIAFINKLDRPGADPWKVITQARS 201
+ I INK+D+P ADP +V Q +S
Sbjct: 339 FESNLAIIPVINKIDQPTADPDRVKGQLKS 428
Score = 29.6 bits (65), Expect = 3.8
Identities = 21/84 (25%), Positives = 42/84 (50%)
Frame = +3
Query: 370 GQLTYLRVYEGVIRKGDFIVNVSTGKKIKVPRLVQMHSNEMNDIEEAHAGQIVAVFGVDC 429
G + ++ V +G +RKGD I + +TGK + + MH E+ GQ+ V
Sbjct: 594 GVICHVAVVDGALRKGDKISSAATGKSYEAMDIGIMHP-ELTPTGILFTGQVGYVITGMR 770
Query: 430 ASSDTFTDGSVKYTMTSMNVPEPV 453
+ + ++ +T ++++V EP+
Sbjct: 771 TTKEARIGDTIYHTKSTVDV-EPL 839
>TC78051 homologue to PIR|T01400|T01400 translation elongation factor EF-Tu
precursor mitochondrial - Arabidopsis thaliana, partial
(87%)
Length = 1681
Score = 56.2 bits (134), Expect = 4e-08
Identities = 44/147 (29%), Positives = 64/147 (42%), Gaps = 6/147 (4%)
Frame = +3
Query: 46 PWWKESMEKVR-----NIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDF 100
PWW+ R N+G H+D GKTTLT I L E ++K ++D
Sbjct: 189 PWWRSMATFSRTKPHVNVGTIGHVDHGKTTLTAAITRV-----LADEGKAKAVAFDEIDK 353
Query: 101 KPLEIIMGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGV 160
P E GITI +A +D PGH D+ + +DG +LV+ + G
Sbjct: 354 APEEKKRGITIATAHVEYETAKRHYAHVDCPGHADYVKNMITGAAQMDGGILVVSAPDGP 533
Query: 161 QCQSITVDRQMRRYQVPR-IAFINKLD 186
Q+ R+ VP + F+NK+D
Sbjct: 534 MPQTKEHILLARQVGVPSLVCFLNKVD 614
>TC76989 homologue to SP|O24310|EFTU_PEA Elongation factor Tu chloroplast
precursor (EF-Tu). [Garden pea] {Pisum sativum}, partial
(92%)
Length = 1907
Score = 53.1 bits (126), Expect = 3e-07
Identities = 41/136 (30%), Positives = 58/136 (42%), Gaps = 5/136 (3%)
Frame = +3
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPK----MDFKPLEIIMGITIK 112
NIG H+D GKTTLT L + S PK +D P E GITI
Sbjct: 423 NIGTIGHVDHGKTTLTA---------ALTMALASLGNSAPKKYDEIDAAPEERARGITIN 575
Query: 113 SAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMR 172
+A + +D PGH D+ + +DGA+LV+ G Q+ +
Sbjct: 576 TATVEYETETRHYAHVDCPGHADYVKNMITGAAQMDGAILVVSGADGPMPQTKEHILLAK 755
Query: 173 RYQVPR-IAFINKLDR 187
+ VP + F+NK D+
Sbjct: 756 QVGVPSVVVFLNKQDQ 803
>TC92627 similar to PIR|T49975|T49975 hypothetical protein F12B17.10 -
Arabidopsis thaliana, partial (25%)
Length = 827
Score = 50.4 bits (119), Expect = 2e-06
Identities = 36/108 (33%), Positives = 52/108 (47%), Gaps = 10/108 (9%)
Frame = +2
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKIHLM----YEVRSK-DGMGP-----KMDFKPLEII 106
N+ I H+DSGK+TL+ +L G+I YE +K G G +D E
Sbjct: 224 NLAIVGHVDSGKSTLSGRLLHLLGRISKKEMHKYEKEAKLQGKGSFAYAWALDESSEERE 403
Query: 107 MGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVL 154
GIT+ A Y + K + ++D+PGH DF + D AVLV+
Sbjct: 404 RGITMTVAVAYFDTKKYHVVVLDSPGHKDFIPNMISGATQADAAVLVI 547
>TC90361 similar to PIR|S35701|S35701 translation elongation factor EF-G
chloroplast - soybean, partial (15%)
Length = 790
Score = 48.1 bits (113), Expect = 1e-05
Identities = 28/83 (33%), Positives = 44/83 (52%), Gaps = 1/83 (1%)
Frame = +2
Query: 421 IVAVFGV-DCASSDTFTDGSVKYTMTSMNVPEPVMSLAVQPVSKDSGGKFSKALNRFQRE 479
IV + G+ D + +T D + + M+ P V+ +A++P +K K L + E
Sbjct: 365 IVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAEE 544
Query: 480 DPTFRVSLDPESGQTIISGMGEL 502
DP+FRVS D E +TII G G +
Sbjct: 545 DPSFRVSQDKED-RTIIQGTGRI 610
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,301,195
Number of Sequences: 36976
Number of extensions: 187579
Number of successful extensions: 897
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 858
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 869
length of query: 654
length of database: 9,014,727
effective HSP length: 102
effective length of query: 552
effective length of database: 5,243,175
effective search space: 2894232600
effective search space used: 2894232600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146553.2


