

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
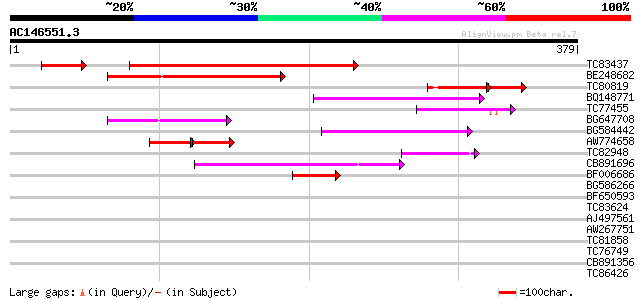
Query= AC146551.3 - phase: 1 /pseudo
(379 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 160 9e-43
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 107 1e-23
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 72 6e-19
BQ148771 66 3e-11
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 55 3e-08
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 54 1e-07
BG584442 49 2e-06
AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsut... 35 4e-05
TC82948 42 4e-04
CB891696 42 4e-04
BF006686 42 4e-04
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 39 0.004
BF650593 38 0.007
TC83624 homologue to PIR|G84581|G84581 copia-like retroelement p... 37 0.013
AJ497561 weakly similar to GP|10140689|gb putative non-LTR retro... 32 0.53
AW267751 29 2.6
TC81858 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 29 3.4
TC76749 similar to GP|15293289|gb|AAK93755.1 unknown protein {Ar... 28 4.5
CB891356 similar to GP|20269067|em protease inhibitor {Sesbania ... 28 4.5
TC86426 similar to PIR|T07634|T07634 pollen-specific protein hom... 28 5.8
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 160 bits (405), Expect(2) = 9e-43
Identities = 79/153 (51%), Positives = 112/153 (72%)
Frame = +2
Query: 81 VLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNVRTMMAVLLLFEEISG 140
VLM ++V T ++ + G + V V+HLQFA+DTL++ K W N+R + A L++F +SG
Sbjct: 470 VLMKSLVQTQLFTRYSFGVVNPVVVSHLQFANDTLLLETKNWANIRALRAALVIF*AMSG 649
Query: 141 LKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIGGDPRKLSFWKPVIDRI 200
LKVNFHKS L VNI+ SWL EA+++++ G F+YLG+PI G+ R+LSFW+P+++RI
Sbjct: 650 LKVNFHKSGLVCVNIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGNSRRLSFWEPIVNRI 829
Query: 201 VGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFL 233
RL+ WN++ LS GGRLVLLK VL+SL VY L
Sbjct: 830 KARLTGWNSRFLSFGGRLVLLKSVLTSLSVYAL 928
Score = 31.2 bits (69), Expect(2) = 9e-43
Identities = 13/30 (43%), Positives = 19/30 (63%)
Frame = +1
Query: 22 GKYEFSNTLAEMDK*VCWDSNGIDPCKWES 51
G+Y FS ++ ++D VC SN C+WES
Sbjct: 373 GRYVFSGSVEKVD*RVC*YSNNFSACEWES 462
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 107 bits (266), Expect = 1e-23
Identities = 57/119 (47%), Positives = 75/119 (62%)
Frame = +3
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PL+PFLFLL AEG + LM V +++ F V G RV+HLQ+ADDTL IG N+
Sbjct: 45 PLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGG-TRVSHLQYADDTLCIGMPTVDNL 221
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIG 184
T+ A+L FE SGLKVNFHKS L G+N+ ++ A +NC + F+YLG+P G
Sbjct: 222 WTLKALLQGFEMASGLKVNFHKSSLIGINVPRDFMEAACRFLNCREESIPFIYLGLPGG 398
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 71.6 bits (174), Expect(2) = 6e-19
Identities = 31/43 (72%), Positives = 38/43 (88%)
Frame = +1
Query: 280 GLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKAKYGEEGG 322
GLGV GAFN+SLLGKWCWR+L +++ LW+RVLKA+YGEEGG
Sbjct: 31 GLGV---GAFNLSLLGKWCWRLLVDKEGLWHRVLKARYGEEGG 150
Score = 40.0 bits (92), Expect(2) = 6e-19
Identities = 15/25 (60%), Positives = 19/25 (76%)
Frame = +3
Query: 321 GGRIREGGRLSSAWWKMVCHIREGL 345
GG + EGGR SS WWK +C +REG+
Sbjct: 144 GGWLCEGGRQSSMWWKTICKVREGV 218
>BQ148771
Length = 680
Score = 65.9 bits (159), Expect = 3e-11
Identities = 33/114 (28%), Positives = 59/114 (50%)
Frame = -3
Query: 204 LSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIESIFKSFFWGGGEENRK 263
L++W LS+ R+ L K V+ ++P+Y + P I I+ + + F WG E +R+
Sbjct: 573 LANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDTEVSRR 394
Query: 264 IAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKAKY 317
+ W+T+ K GLG+RR+ N + + K W + + + L V++ KY
Sbjct: 393 YHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEVMRGKY 232
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 55.5 bits (132), Expect = 3e-08
Identities = 31/78 (39%), Positives = 43/78 (54%), Gaps = 12/78 (15%)
Frame = -3
Query: 273 CLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKAKYGEE-------GGRI- 324
CL + +GGLGVR + NVSLL KW WR+L ++ LW VL+ YG GRI
Sbjct: 969 CLPRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIYGPRVSSRTMMVGRIG 790
Query: 325 ----REGGRLSSAWWKMV 338
++GG++ W + V
Sbjct: 789 RLMLQDGGKI**VWRRWV 736
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 53.5 bits (127), Expect = 1e-07
Identities = 31/83 (37%), Positives = 43/83 (51%)
Frame = +1
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L A + L+ H V S+ ++THL FADD+L+
Sbjct: 31 PLSPYLFILCANVLSGLLKREGNKQNLHGIQVAR-SDPKITHLLFADDSLLFARANLTEA 207
Query: 126 RTMMAVLLLFEEISGLKVNFHKS 148
T+M VL ++ SG VNF KS
Sbjct: 208 ATIMQVLHSYQSASGQLVNFEKS 276
>BG584442
Length = 775
Score = 49.3 bits (116), Expect = 2e-06
Identities = 33/102 (32%), Positives = 48/102 (46%), Gaps = 1/102 (0%)
Frame = +1
Query: 209 NKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIESIFKSFFWGGGEENRK-IAWI 267
NK LS V++K L S+ Y +S F + IE I +F W ENRK + W+
Sbjct: 400 NKCLSKVM*EVMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVGENRKGMHWM 579
Query: 268 KWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELW 309
+ + + K GG+G FN+ +LGK L R L+
Sbjct: 580 S*EKLFVHKNYGGMGFTDFTTFNIPMLGKQV*SFLLNRTTLF 705
>AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsutum}, partial
(4%)
Length = 665
Score = 35.4 bits (80), Expect(2) = 4e-05
Identities = 15/31 (48%), Positives = 20/31 (64%)
Frame = -1
Query: 94 SFGVGNGSEVRVTHLQFADDTLIIGEKCWLN 124
+ +G S +HLQFADDTL++G K W N
Sbjct: 419 NLSIGMHSLTVFSHLQFADDTLLLGVKSWAN 327
Score = 29.3 bits (64), Expect(2) = 4e-05
Identities = 11/28 (39%), Positives = 20/28 (71%)
Frame = -3
Query: 123 LNVRTMMAVLLLFEEISGLKVNFHKSML 150
L R + ++L++FE +SGLKVN + ++
Sbjct: 336 LGERALRSILVIFENMSGLKVNLREEVI 253
>TC82948
Length = 705
Score = 42.0 bits (97), Expect = 4e-04
Identities = 16/52 (30%), Positives = 29/52 (55%)
Frame = +3
Query: 263 KIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLK 314
K+ + W+ +C EG LG+R + N +L K CW M+ +++ WY ++
Sbjct: 255 KVVKVSWEKVCRPIKEGSLGIRSLSKLNEALNLKLCWDMMISKEQ-WYAFMR 407
>CB891696
Length = 638
Score = 42.0 bits (97), Expect = 4e-04
Identities = 38/142 (26%), Positives = 62/142 (42%), Gaps = 1/142 (0%)
Frame = +1
Query: 124 NVRTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPI 183
N+ TM ++ FE S L VNF KS L +N+ + + + C F YLG+ +
Sbjct: 10 NILTMKTIVSYFELASSLWVNFLKSGLINLNVIGHF*GW*NIYIKCKVH*VIFKYLGILV 189
Query: 184 GGDPRKLSFWKPVIDRIVGRLSS-WNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGI 242
G +P +++ + ++ + L S WN K L + S +YF S K P +
Sbjct: 190 GENPCRVNM*ELLLKLLTN*LGSWWNTK*LWTQNGFSQIHAK*ISQNIYF-SLMKIPVKV 366
Query: 243 ISSIESIFKSFFWGGGEENRKI 264
I + F G + +KI
Sbjct: 367 *ELISQLKTQFL*GNLKVTKKI 432
>BF006686
Length = 325
Score = 42.0 bits (97), Expect = 4e-04
Identities = 17/32 (53%), Positives = 23/32 (71%)
Frame = +3
Query: 190 LSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLL 221
L W+P+++ + L SW NKLLS GGR+VLL
Sbjct: 228 LPMWEPLLEHVNKMLKSWGNKLLSFGGRIVLL 323
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 38.5 bits (88), Expect = 0.004
Identities = 24/86 (27%), Positives = 40/86 (45%)
Frame = -3
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF+L E + L + V + HL FADDT+ G+ +
Sbjct: 367 PLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPP-INHLLFADDTMFFGKSNASSC 191
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLT 151
+++++ + SG +N KS +T
Sbjct: 190 AILLSIMDKYRAASGRCIN*TKSAIT 113
>BF650593
Length = 486
Score = 37.7 bits (86), Expect = 0.007
Identities = 18/40 (45%), Positives = 22/40 (55%)
Frame = +1
Query: 308 LWYRVLKAKYGEEGGRIREGGRLSSAWWKMVCHIREGLVD 347
LW+R L KYG G I R S WWK +C+I G V+
Sbjct: 88 LWFRALVNKYGLNRGSITIENRGVSLWWKDICYIDFGGVE 207
>TC83624 homologue to PIR|G84581|G84581 copia-like retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(1%)
Length = 831
Score = 37.0 bits (84), Expect = 0.013
Identities = 16/56 (28%), Positives = 31/56 (54%)
Frame = +1
Query: 145 FHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIGGDPRKLSFWKPVIDRI 200
F + +N+ +S++ + + C F +LG+PIG +P++ S KPV+D +
Sbjct: 256 FSRVNFMALNLEESFVEASPNFLLCNVNEVPFCFLGLPIGANPKRSSTRKPVLDSL 423
>AJ497561 weakly similar to GP|10140689|gb putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 621
Score = 31.6 bits (70), Expect = 0.53
Identities = 13/37 (35%), Positives = 26/37 (70%)
Frame = +3
Query: 268 KWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTE 304
+++ + K+ GGLG++ FN+S+LGK W++L++
Sbjct: 300 EFNNLARHKSVGGLGLQNQ-EFNISMLGKQRWKLLSQ 407
>AW267751
Length = 708
Score = 29.3 bits (64), Expect = 2.6
Identities = 8/28 (28%), Positives = 19/28 (67%)
Frame = +3
Query: 242 IISSIESIFKSFFWGGGEENRKIAWIKW 269
+++ + I ++FF+ G+E +K+ W+ W
Sbjct: 531 VLTILIGI*RTFFFSSGDETKKVDWVTW 614
>TC81858 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (9%)
Length = 748
Score = 28.9 bits (63), Expect = 3.4
Identities = 12/21 (57%), Positives = 12/21 (57%)
Frame = -2
Query: 280 GLGVRRMGAFNVSLLGKWCWR 300
G G R G FN S G WCWR
Sbjct: 357 GSGWRLGGRFNGSFSGGWCWR 295
>TC76749 similar to GP|15293289|gb|AAK93755.1 unknown protein {Arabidopsis
thaliana}, partial (64%)
Length = 860
Score = 28.5 bits (62), Expect = 4.5
Identities = 22/66 (33%), Positives = 31/66 (46%), Gaps = 3/66 (4%)
Frame = +1
Query: 168 NCCRGNFTFVYLGVPIGGDPRKLSFWKPVIDRIVGRL-SSWNNKLLSI--GGRLVLLKVV 224
NCC T+ +LG IGG S + V D R WN ++ + GG L L ++
Sbjct: 157 NCCHVCCTYSWLGPEIGGRMASYSLPRAVSDSQAERH*VPWNRRI*QVLWGGSLQLCRLR 336
Query: 225 LSSLPV 230
S+L V
Sbjct: 337 DSTLQV 354
>CB891356 similar to GP|20269067|em protease inhibitor {Sesbania rostrata},
partial (51%)
Length = 829
Score = 28.5 bits (62), Expect = 4.5
Identities = 22/66 (33%), Positives = 31/66 (46%), Gaps = 3/66 (4%)
Frame = +2
Query: 168 NCCRGNFTFVYLGVPIGGDPRKLSFWKPVIDRIVGRL-SSWNNKLLSI--GGRLVLLKVV 224
NCC T+ +LG IGG S + V D R WN ++ + GG L L ++
Sbjct: 512 NCCHVCCTYSWLGPEIGGRMASYSLPRAVSDSQAERH*VPWNRRI*QVLWGGSLQLCRLR 691
Query: 225 LSSLPV 230
S+L V
Sbjct: 692 DSTLQV 709
>TC86426 similar to PIR|T07634|T07634 pollen-specific protein homolog T1P17.10
- Arabidopsis thaliana, partial (91%)
Length = 2220
Score = 28.1 bits (61), Expect = 5.8
Identities = 19/56 (33%), Positives = 25/56 (43%)
Frame = +3
Query: 255 WGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWY 310
+G EN + + KWD I S A+ G A VSL W + TE + WY
Sbjct: 1575 FGDWSENSRGTYNKWDGIARSTAQVYPGA--WTAVLVSLDNVGVWNLRTENLDSWY 1736
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.331 0.146 0.494
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,403,378
Number of Sequences: 36976
Number of extensions: 233717
Number of successful extensions: 1463
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 1447
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1457
length of query: 379
length of database: 9,014,727
effective HSP length: 98
effective length of query: 281
effective length of database: 5,391,079
effective search space: 1514893199
effective search space used: 1514893199
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146551.3


