

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146343.3 - phase: 0
(670 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
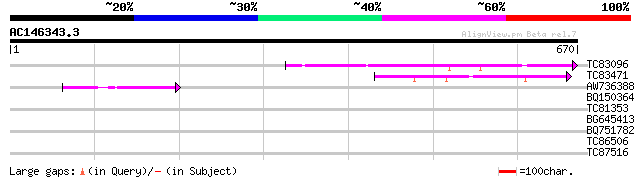
TC83096 weakly similar to PIR|T45912|T45912 hypothetical protein... 117 1e-26
TC83471 similar to PIR|T45912|T45912 hypothetical protein F5K20.... 102 6e-22
AW736388 similar to GP|10178015|db Na+/H+ antiporter-like protei... 59 5e-09
BQ150364 31 1.8
TC81353 similar to GP|21593362|gb|AAM65311.1 microtubule-associa... 31 1.8
BG645413 similar to GP|15341529|gb ferric-chelate reductase {Pis... 29 5.1
BQ751782 similar to GP|20145247|e probable isoleucyl-trna synthe... 29 6.7
TC86506 similar to GP|3075391|gb|AAC14523.1| unknown protein {Ar... 29 6.7
TC87516 PIR|T09591|T09591 probable cdc2-like protein kinase cdc2... 28 8.7
>TC83096 weakly similar to PIR|T45912|T45912 hypothetical protein F5K20.20 -
Arabidopsis thaliana, partial (21%)
Length = 1227
Score = 117 bits (294), Expect = 1e-26
Identities = 97/372 (26%), Positives = 176/372 (47%), Gaps = 28/372 (7%)
Frame = +2
Query: 327 LYDPSRKYAGYQKRNILSLKPNSELKIVSCILKPSHIIPIKNVLDICSPTS-SNPLVIHI 385
+Y PSR+ + + +L+I++CI +I + N ++ T+ S+ + +++
Sbjct: 11 IYKPSRQRRSGNPPPLTDTQ--EKLRILACIHGTGNIPSLINFIESVRATNKSSKIKLYV 184
Query: 386 LHLLELVGRSSPVFISHRLQERVGSSSHTFSEAVIVTFDLFEH-DNAGTASVSTYTAISP 444
+ L EL SS + + ++ + F + + + F G +V T+IS
Sbjct: 185 MQLTELTDSSSSILMVRSSRKSGFPFINRFQKGTMQ--EAFRACGQVGQVTVHHLTSISS 358
Query: 445 VRFMHDDICYLALDKLASIIILPFHLRW--SEDGSVESADETTRSLNTKVLERAPCSVAI 502
+ +H+DIC++A +K ++IILPFH RW ++ ++E + R +N +VL+ APCSVA+
Sbjct: 359 LSTIHEDICHIAEEKGVAMIILPFHKRWRGEDEETIEDIGQRWREVNQRVLQSAPCSVAV 538
Query: 503 LVNRGHSSPFNHNENS-----KQIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHL--- 554
LVNRG + + K++ +IF+GG DDR+ L L R + L V
Sbjct: 539 LVNRGVGRRYEQRVETSATPGKKVCIIFVGGPDDRKVLELGSRMAEHPAIRLSVVRFNLH 718
Query: 555 -------------VSTIKNDEFTSWDVMLDDELLKGVKGVY-GSVDNVTYEKVEVENTSD 600
ST +D + LD+ L K + G+V+ + + V + N
Sbjct: 719 NEGTFRDQEHSYNTSTSASDNNMENEKELDEVALNEFKTKWLGAVEYIENDTVNIANEVL 898
Query: 601 TTEFISDIAIQHDFIIVGRRNGI--KSPQTQALASWTEYPELGVLGDLLASPDTNTKASI 658
+ +++ +IVG+ + + + S E+ ELG +GDLL S +S+
Sbjct: 899 AIGRVK----EYELVIVGKGHQLLNSTGMIDIKDSQLEHAELGPIGDLLTSSAQGITSSV 1066
Query: 659 LVVQQQVMPKAS 670
LV+Q Q + +S
Sbjct: 1067LVIQGQHLINSS 1102
>TC83471 similar to PIR|T45912|T45912 hypothetical protein F5K20.20 -
Arabidopsis thaliana, partial (18%)
Length = 983
Score = 102 bits (253), Expect = 6e-22
Identities = 78/271 (28%), Positives = 129/271 (46%), Gaps = 38/271 (14%)
Frame = +3
Query: 432 GTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSEDGS----------VESA 481
G V + TAIS + MH+DIC+ A +K ++IILPFH W + +E+A
Sbjct: 9 GRVIVRSTTAISSLSTMHEDICHAAEEKRVTMIILPFHKHWRMEVDDENDKEAHEVLENA 188
Query: 482 DETTRSLNTKVLERAPCSVAILVNRGHSSPFNH----NENSKQIAMIFLGGSDDREALCL 537
R +N +VL+ APCSVA+LV+RG+ + +++I ++F GG DDREAL L
Sbjct: 189 GHGWRGVNQRVLKNAPCSVAVLVDRGYGLGLKNLGSDGRVAQRICIVFFGGPDDREALEL 368
Query: 538 AKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGVYG-SVDNVTYEKVEVE 596
K+ ++ + VV +V ++ +E + + +L K + Y S+ + +K +V
Sbjct: 369 GKKMVE---HPAVVVTVVRFVEQNELSGNNFVLRQSPGKSTEENYSFSIAKINRQKEQVL 539
Query: 597 NTSDTTEFISDI-----------------------AIQHDFIIVGRRNGIKSPQTQALAS 633
+ + EF S + +D I+VG+ + +
Sbjct: 540 DENAMEEFRSKCGETVKYIEKGSGNVVEEVIALGESADYDLIVVGKGRFPSTMVAELAER 719
Query: 634 WTEYPELGVLGDLLASPDTNTKASILVVQQQ 664
E+ ELG +GD+L S + AS + V QQ
Sbjct: 720 EAEHAELGPIGDILTSSMGHKMASSVFVIQQ 812
>AW736388 similar to GP|10178015|db Na+/H+ antiporter-like protein
{Arabidopsis thaliana}, partial (21%)
Length = 558
Score = 59.3 bits (142), Expect = 5e-09
Identities = 40/141 (28%), Positives = 72/141 (50%), Gaps = 1/141 (0%)
Frame = +3
Query: 63 FAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGKGT 122
F V+A +L++L++L + +GR+A+S A V D I+ + A +G+ +
Sbjct: 156 FPVLARILAELKLLTTSVGRMAMSAAAVNDVAAWILLALAVAL----------SGNSQSP 305
Query: 123 LKAFLNVFYYLC-FMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGLL 181
L + L VF C F+V + L++ PI KW ++ EG P+ ++Y+ LA G +
Sbjct: 306 LVS-LWVFLAGCGFVVCSILIVLPIFKWMAQQCHEGEPVDELYICATLASVLAAGFVTDA 482
Query: 182 TKQSVLGGICIVGLIVPEGPP 202
+ G + G++VP+ P
Sbjct: 483 IGIHAMFGAFVFGILVPKDGP 545
>BQ150364
Length = 714
Score = 30.8 bits (68), Expect = 1.8
Identities = 26/71 (36%), Positives = 36/71 (50%)
Frame = +2
Query: 324 VKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVI 383
V LYD R+ YQ R +L+P + LKI S K +HI P ++C P + PL+
Sbjct: 437 VSTLYDGGRQTPSYQGRLPTTLRPVTVLKI-SLGFKAAHIEP----FNLCIPI-TEPLLF 598
Query: 384 HILHLLELVGR 394
+LE V R
Sbjct: 599 IACSVLESV*R 631
>TC81353 similar to GP|21593362|gb|AAM65311.1 microtubule-associated protein
EB1-like protein {Arabidopsis thaliana}, partial (41%)
Length = 669
Score = 30.8 bits (68), Expect = 1.8
Identities = 17/69 (24%), Positives = 33/69 (47%)
Frame = -2
Query: 360 PSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSHTFSEAV 419
P+H+ P++ ++ + S + + H LH L+++ RSS H+L S F E +
Sbjct: 503 PAHLSPLQRIITVLSRINRITVSFHPLHELQIIQRSS----FHQLANLNMLSDFEFVENI 336
Query: 420 IVTFDLFEH 428
+ + H
Sbjct: 335 LKNLIILNH 309
>BG645413 similar to GP|15341529|gb ferric-chelate reductase {Pisum sativum},
partial (29%)
Length = 815
Score = 29.3 bits (64), Expect = 5.1
Identities = 15/52 (28%), Positives = 29/52 (54%), Gaps = 6/52 (11%)
Frame = +3
Query: 103 TAFISSIKTDSHDNGDGKGTLKAFLNVFYYL------CFMVVTPLVLRPILK 148
TA +++ TD+H + TLK + N+ + L + V+ P+++R I+K
Sbjct: 498 TATLATTTTDTHITSTLRNTLKGYENILFVL*SIGKM*YRVMIPIMIRKIMK 653
>BQ751782 similar to GP|20145247|e probable isoleucyl-trna synthetase
{Aspergillus fumigatus}, partial (20%)
Length = 695
Score = 28.9 bits (63), Expect = 6.7
Identities = 13/39 (33%), Positives = 20/39 (50%)
Frame = +2
Query: 114 HDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVK 152
H G K + + L Y +C+ TPL+ R + WFV+
Sbjct: 83 HLKGADKIVVDSSLKHSYPMCYRSDTPLIYRAVPSWFVR 199
>TC86506 similar to GP|3075391|gb|AAC14523.1| unknown protein {Arabidopsis
thaliana}, partial (6%)
Length = 840
Score = 28.9 bits (63), Expect = 6.7
Identities = 17/41 (41%), Positives = 22/41 (53%), Gaps = 5/41 (12%)
Frame = +3
Query: 466 LPFHLRWSEDGSVESADETTR-----SLNTKVLERAPCSVA 501
+PF LR GS S D TTR SL VL+R+ S++
Sbjct: 423 IPFSLRSLPSGSTSSVDATTRLTTAGSLGKAVLDRSESSIS 545
>TC87516 PIR|T09591|T09591 probable cdc2-like protein kinase cdc2MsF -
alfalfa, complete
Length = 1251
Score = 28.5 bits (62), Expect = 8.7
Identities = 21/72 (29%), Positives = 31/72 (42%), Gaps = 3/72 (4%)
Frame = +2
Query: 573 ELLKGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQ---HDFIIVGRRNGIKSPQTQ 629
E+L G +VD + + E + T F D +Q H F ++G N P
Sbjct: 641 EVLLGATHYSMAVDMWSVACIFAELVTKTALFPGDSELQQLLHIFRLLGTPNEDVWPGVS 820
Query: 630 ALASWTEYPELG 641
L +W EYP+ G
Sbjct: 821 KLMNWHEYPQWG 856
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.139 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,684,524
Number of Sequences: 36976
Number of extensions: 351759
Number of successful extensions: 2545
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 2523
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2538
length of query: 670
length of database: 9,014,727
effective HSP length: 103
effective length of query: 567
effective length of database: 5,206,199
effective search space: 2951914833
effective search space used: 2951914833
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146343.3


