

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146342.8 - phase: 0
(337 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
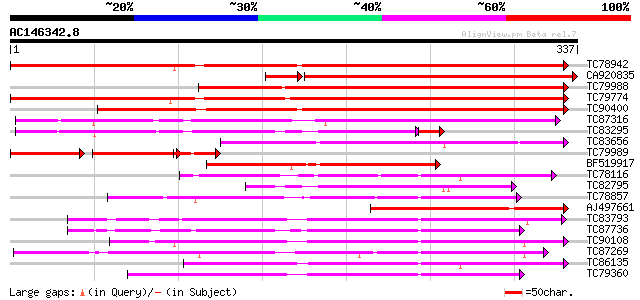
Score E
Sequences producing significant alignments: (bits) Value
TC78942 weakly similar to GP|20146758|gb|AAM12494.1 cytochrome P... 353 7e-98
CA920835 weakly similar to GP|5734639|dbj ESTs AU056036(S20239) ... 338 3e-95
TC79988 similar to PIR|T00864|T00864 cytochrome P450 homolog F17... 336 9e-93
TC79774 weakly similar to GP|20146747|gb|AAM12483.1 cytochrome P... 323 6e-89
TC90400 similar to GP|20146747|gb|AAM12483.1 cytochrome P450-lik... 301 3e-82
TC87316 weakly similar to PIR|G84675|G84675 probable cytochrome ... 225 2e-59
TC83295 similar to SP|O81117|C941_VICSA Cytochrome P450 94A1 (EC... 156 9e-40
TC83656 similar to GP|19698839|gb|AAL91155.1 cytochrome P450 {Ar... 143 8e-35
TC79989 similar to PIR|T00864|T00864 cytochrome P450 homolog F17... 68 5e-29
BF519917 similar to GP|19698839|gb cytochrome P450 {Arabidopsis ... 124 7e-29
TC78116 similar to PIR|F86441|F86441 probable cytochrome P450 [i... 123 9e-29
TC82795 similar to GP|18000068|gb|AAL54885.1 cytochrome P450-dep... 112 2e-25
TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 7... 103 1e-22
AJ497661 similar to GP|10442763|gb cytochrome P450 {Triticum aes... 102 2e-22
TC83793 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P... 100 8e-22
TC87736 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P... 99 2e-21
TC90108 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (E... 97 9e-21
TC87269 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soy... 96 1e-20
TC86135 similar to PIR|A96673|A96673 probable cytochrome P450 F1... 94 6e-20
TC79360 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (E... 94 1e-19
>TC78942 weakly similar to GP|20146758|gb|AAM12494.1 cytochrome P450-like
protein {Oryza sativa (japonica cultivar-group)},
partial (58%)
Length = 1719
Score = 353 bits (905), Expect = 7e-98
Identities = 185/335 (55%), Positives = 235/335 (69%), Gaps = 3/335 (0%)
Frame = +3
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
MK TLDSIF V FG E++ + +S+E F +F++++A RY+D WK+K+ LN GS
Sbjct: 570 MKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLNIGS 749
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKE--DILSRFLLESKKDSSNMTDKYL 118
EAAL+ N +I+++FV LI TR + + K S ++ DILSRFL + D+ KYL
Sbjct: 750 EAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDT-----KYL 914
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSN 178
RDIILNF+IAGKDT+ TLSWF YMLCK P +QEK AQEV T+T S+ EFVS+
Sbjct: 915 RDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCT--EFVSS 1088
Query: 179 ISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGR 238
++D ++KM+Y+HA LTETLRLYPA+P + D LPDGY V K + V Y YAMGR
Sbjct: 1089VTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKRDMVSYQPYAMGR 1268
Query: 239 MPYIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALV 297
M +IWGDDA+EF PERWL ++GIFQPE FKF +F AGPRICLGK+FAYRQMKI S L+
Sbjct: 1269MKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLL 1448
Query: 298 RFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
FRFKL +E +VTY+TM TLHID GL + A R
Sbjct: 1449GCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYR 1553
>CA920835 weakly similar to GP|5734639|dbj ESTs AU056036(S20239)
C72753(E2173) AU056035(S20239) correspond to a region
of the predicted gene.~, partial (36%)
Length = 809
Score = 338 bits (868), Expect(2) = 3e-95
Identities = 162/162 (100%), Positives = 162/162 (100%)
Frame = -2
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA
Sbjct: 733 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 554
Query: 236 MGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA 295
MGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA
Sbjct: 553 MGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA 374
Query: 296 LVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPRCDVAE 337
LVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPRCDVAE
Sbjct: 373 LVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPRCDVAE 248
Score = 28.1 bits (61), Expect(2) = 3e-95
Identities = 14/22 (63%), Positives = 15/22 (67%)
Frame = -3
Query: 153 KVAQEVINVTSTSQESNLNLDE 174
K E TST+QESNLNLDE
Sbjct: 801 KSGTESDKCTSTAQESNLNLDE 736
>TC79988 similar to PIR|T00864|T00864 cytochrome P450 homolog F17K2.4 -
Arabidopsis thaliana, partial (39%)
Length = 808
Score = 336 bits (861), Expect = 9e-93
Identities = 157/221 (71%), Positives = 189/221 (85%), Gaps = 1/221 (0%)
Frame = +2
Query: 113 MTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLN- 171
+TDKYLRDIILNFMIAGKDT+ANTLSWFFYMLCKNPL+++K+ QE+ +VT S ES LN
Sbjct: 5 ITDKYLRDIILNFMIAGKDTTANTLSWFFYMLCKNPLVEDKIVQEIRDVT-CSHESELNS 181
Query: 172 LDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYY 231
+DEFV+N++D LDKMHYLHA LTETLRLYP +P+ GRTA+ D+LPDG+ + KG+ VYY
Sbjct: 182 IDEFVANLTDLILDKMHYLHATLTETLRLYPVLPVDGRTADAPDVLPDGHKLEKGDGVYY 361
Query: 232 LSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKI 291
L+YAMGRM IWG+DA EF PERW+ DGIFQPES FKF++FHAGPR+CLGKDFAYRQMKI
Sbjct: 362 LAYAMGRMSSIWGEDADEFRPERWITDGIFQPESPFKFVAFHAGPRMCLGKDFAYRQMKI 541
Query: 292 VSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
V+M ++ FF+FKL N T +VTY+ MFTLH+D GLPL A PR
Sbjct: 542 VAMCVLNFFKFKLANGTQNVTYKVMFTLHLDKGLPLTAIPR 664
>TC79774 weakly similar to GP|20146747|gb|AAM12483.1 cytochrome P450-like
protein {Oryza sativa (japonica cultivar-group)},
partial (52%)
Length = 1609
Score = 323 bits (828), Expect = 6e-89
Identities = 168/335 (50%), Positives = 233/335 (69%), Gaps = 3/335 (0%)
Frame = +2
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
MK TLDS+FK GVEL+ + + +E F AF++++A + RY++ LWK++R LN GS
Sbjct: 605 MKSTLDSVFKGILGVELDTMCGTYREGTQFSNAFDEASAAIMFRYVNFLWKVQRFLNMGS 784
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSD--KEDILSRFLLESKKDSSNMTDKYL 118
EA LKKN+++ID++V+++I+++ E ++ S K DILSRFL ++ DS KYL
Sbjct: 785 EAVLKKNLRVIDEYVYTVIRSKIEQSQKPQNNSSELKGDILSRFLELNETDS-----KYL 949
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSN 178
+D+IL+F+IAGKDT+A TLSWF Y LCK+P +QEK+AQE++ T E +DE +
Sbjct: 950 KDVILSFIIAGKDTTAITLSWFLYQLCKHPHVQEKIAQEIMEATKV--EDGSTIDELAAR 1123
Query: 179 ISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGR 238
+++ +++KM YLHAALTETLRL+P VP+ + D LPDGY V KG+ V + Y MGR
Sbjct: 1124LTEESMEKMQYLHAALTETLRLHPPVPVESKYCFSDDTLPDGYSVTKGDLVSFQPYVMGR 1303
Query: 239 MPYIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALV 297
M ++WG+DA++F PERWL ++G FQ ES FKF +F AGPRICLGK+FAYRQMKI S L+
Sbjct: 1304MKFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLL 1483
Query: 298 RFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
FKL ++ V YRT TL ID GL + A R
Sbjct: 1484GSHSFKLADQNKLVKYRTSLTLQIDDGLHVNAFHR 1588
>TC90400 similar to GP|20146747|gb|AAM12483.1 cytochrome P450-like protein
{Oryza sativa (japonica cultivar-group)}, partial (37%)
Length = 967
Score = 301 bits (770), Expect = 3e-82
Identities = 161/282 (57%), Positives = 199/282 (70%), Gaps = 2/282 (0%)
Frame = +1
Query: 53 KRVLNFGSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKE-DILSRFLLESKKDSS 111
K V F SEAAL+KN +++++FV LI TR + ++ + D K DILSRFL + D++
Sbjct: 1 KEVSQFRSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKEYDTT 180
Query: 112 NMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLN 171
YLRDIILNF+IAGKDT+A TLSWF YMLCK P +QEK A+EV T+T S+
Sbjct: 181 -----YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCT 345
Query: 172 LDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYY 231
EFVS ++D L+KM+YLHA LTETLRLYPAVP+ + D LPDGY V KG+ V Y
Sbjct: 346 --EFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSY 519
Query: 232 LSYAMGRMPYIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMK 290
YAMGRM +IWGDDA+EF P RWL ++G FQ E+ FKF +F AGPRICLGK+FAYRQMK
Sbjct: 520 QPYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMK 699
Query: 291 IVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
I S L+ FRFKL +E +VTY+TM LHID GL + A R
Sbjct: 700 IFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHR 825
>TC87316 weakly similar to PIR|G84675|G84675 probable cytochrome P450
[imported] - Arabidopsis thaliana, partial (77%)
Length = 1678
Score = 225 bits (574), Expect = 2e-59
Identities = 129/330 (39%), Positives = 192/330 (58%), Gaps = 6/330 (1%)
Frame = +2
Query: 4 TLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP---LWKLKRVLNFGS 60
+ D+I K FG++ CL S +N+ AF+ S+ +R L +WK+KR N GS
Sbjct: 641 SFDNICKFSFGLDPCCLVPSLPVSNL-ANAFDLSSTLSAQRALTASPLIWKMKRFFNIGS 817
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
E LK+ IKI++D + +IK RRE+ + ++D+LSRF+ ++ D+YLRD
Sbjct: 818 EKKLKEAIKIVNDLANEMIKQRREI---ENGVESRKDLLSRFM----GALNSHDDEYLRD 976
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
I+++F++AG+DT A+ L+ FF +L KNP ++EK+ E+ V + +QE
Sbjct: 977 IVVSFLLAGRDTVASALTGFFILLSKNPKVEEKIRVELDRVMNPNQEC------------ 1120
Query: 181 DATLDK---MHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMG 237
AT ++ MHYL+ A+ E++RL+P V + A E D+LPDG + KG V Y YAMG
Sbjct: 1121-ATFEQTREMHYLNGAIHESMRLFPPVQFDSKFALEDDVLPDGTFIKKGSRVTYHPYAMG 1297
Query: 238 RMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALV 297
RM IWG D EF PERWLKDG+F P+ FK+ F AG R+CLGK+ A +MK V +LV
Sbjct: 1298RMENIWGPDCLEFKPERWLKDGVFVPKCPFKYPVFQAGSRVCLGKELAIVEMKSVVASLV 1477
Query: 298 RFFRFKLENETNDVTYRTMFTLHIDHGLPL 327
+ F ++ + + T GLP+
Sbjct: 1478KRFDVRVVGPNQEPQFAPGLTASFRGGLPV 1567
>TC83295 similar to SP|O81117|C941_VICSA Cytochrome P450 94A1 (EC 1.14.-.-)
(P450-dependent fatty acid omega-hydroxylase)., partial
(62%)
Length = 971
Score = 156 bits (394), Expect(2) = 9e-40
Identities = 86/244 (35%), Positives = 143/244 (58%), Gaps = 3/244 (1%)
Frame = +2
Query: 4 TLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPL---WKLKRVLNFGS 60
T D+I K+ FG + L S+K F +AF D+ +R+ PL WK+K+ N GS
Sbjct: 245 TFDNICKIAFGFDPEYLTPSAKRTK-FAQAFEDATEISSRRFRLPLPVIWKMKKRFNIGS 421
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
E L++ + I ++ + +++ +++ L + D +D+LSRFL S + + ++ D
Sbjct: 422 EKRLREAVTEIREYANKIVREKKKELK-ENDSLHTQDMLSRFL-----SSGHSEEDFVTD 583
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
I+++F++AGKDT++ L+WFF++L KNP ++E++ +E+ + +L+ DE
Sbjct: 584 IVISFILAGKDTTSAALTWFFWLLWKNPRVEEEILKEI-----NKKSESLDYDE------ 730
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
+ M Y HAAL+E++RLYP VPM + A D+L DG +V KG + Y YAMGRM
Sbjct: 731 ---VKTMVYTHAALSESMRLYPPVPMDSKEAINDDVLLDGIVVKKGTMITYHVYAMGRME 901
Query: 241 YIWG 244
+WG
Sbjct: 902 SLWG 913
Score = 25.0 bits (53), Expect(2) = 9e-40
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = +3
Query: 244 GDDAQEFLPERWLKD 258
G+D EF PERWL++
Sbjct: 912 GEDWAEFKPERWLEN 956
>TC83656 similar to GP|19698839|gb|AAL91155.1 cytochrome P450 {Arabidopsis
thaliana}, partial (39%)
Length = 929
Score = 143 bits (361), Expect = 8e-35
Identities = 84/211 (39%), Positives = 120/211 (56%), Gaps = 4/211 (1%)
Frame = +3
Query: 126 MIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTS-TSQESNLNLDEFVSNISDATL 184
+I + T L FF+++ +P ++EK+ +E+ V T E + S+A
Sbjct: 33 LILLEGTRHQLLQLFFWLVMNHPKVEEKIIKELTTVLEETRGGEKRKWTEDPLDFSEA-- 206
Query: 185 DKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWG 244
D+M YL AAL ETLRLYP+VP + A D+ PDG ++ G TV Y Y++GRM IWG
Sbjct: 207 DQMVYLKAALAETLRLYPSVPQDIKQAVVDDVFPDGTVIPAGSTVTYSIYSVGRMEKIWG 386
Query: 245 DDAQEFLPERWLK---DGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFR 301
+D EF PERWL D P+ F F++F+AGPR CLGKD AY QMK V+ A++ +R
Sbjct: 387 EDCLEFKPERWLSVRGDRFEPPKEGFMFVAFNAGPRTCLGKDLAYLQMKSVAAAVLLRYR 566
Query: 302 FKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
L + V + TL + +GL ++ PR
Sbjct: 567 L-LPVPGHVVEQKMSLTLFMKNGLKVFLQPR 656
>TC79989 similar to PIR|T00864|T00864 cytochrome P450 homolog F17K2.4 -
Arabidopsis thaliana, partial (22%)
Length = 806
Score = 67.8 bits (164), Expect(3) = 5e-29
Identities = 31/44 (70%), Positives = 36/44 (81%)
Frame = +2
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKR 44
M+C LDSIFKVGFG ELNCLE SSKE FMKAF++SNA ++ R
Sbjct: 434 MRCALDSIFKVGFGTELNCLEGSSKEGTEFMKAFDESNALIYWR 565
Score = 61.6 bits (148), Expect(3) = 5e-29
Identities = 28/53 (52%), Positives = 37/53 (68%)
Frame = +1
Query: 50 WKLKRVLNFGSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRF 102
W LKR LN G EA LK N+K+IDDFV+ +I T++ L +Q+D + KED S F
Sbjct: 583 WSLKRFLNIGGEAKLKHNVKLIDDFVNGVINTKKSQLELQQDSNVKEDYYSSF 741
Score = 36.2 bits (82), Expect(3) = 5e-29
Identities = 18/28 (64%), Positives = 23/28 (81%)
Frame = +3
Query: 98 ILSRFLLESKKDSSNMTDKYLRDIILNF 125
+L +FL+ESK + + TDKYLRDIILNF
Sbjct: 726 LLFKFLMESKM-AKHYTDKYLRDIILNF 806
>BF519917 similar to GP|19698839|gb cytochrome P450 {Arabidopsis thaliana},
partial (26%)
Length = 505
Score = 124 bits (310), Expect = 7e-29
Identities = 64/141 (45%), Positives = 95/141 (66%), Gaps = 2/141 (1%)
Frame = +2
Query: 118 LRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQ--ESNLNLDEF 175
LR I LNF++AG+DTS+ LSWFF+++ +P ++EK+ E+ V + ++ +S +E
Sbjct: 41 LRHIALNFVLAGRDTSSVALSWFFWLVMNHPSVEEKILAELTAVLAETRGGDSRRWTEEA 220
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
V + +A +K+ YL AAL ETLRLYP+VP + A + D+LPDG +V G TV Y Y+
Sbjct: 221 V-DFEEA--EKLVYLKAALAETLRLYPSVPEDFKYAVDDDVLPDGTVVPAGSTVTYSIYS 391
Query: 236 MGRMPYIWGDDAQEFLPERWL 256
+GRM +WG+D EF P+RWL
Sbjct: 392 VGRMKSVWGEDCMEFKPDRWL 454
>TC78116 similar to PIR|F86441|F86441 probable cytochrome P450 [imported] -
Arabidopsis thaliana, partial (80%)
Length = 1547
Score = 123 bits (309), Expect = 9e-29
Identities = 80/226 (35%), Positives = 120/226 (52%), Gaps = 2/226 (0%)
Frame = +3
Query: 102 FLLESKKDSSNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINV 161
FLL S D +T K LRD ++ +IAG +TSA L+W FY+L K P + K+ +EV +V
Sbjct: 891 FLLASGDD---VTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSV 1061
Query: 162 TSTSQESNLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGY 221
L + I D + K+ Y + E+LRLYP P+ R + E D+L + Y
Sbjct: 1062 ----------LGDRFPTIED--MKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVLGE-Y 1202
Query: 222 IVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESS--FKFISFHAGPRIC 279
+ +GE ++ + + R P +W +DA +F PERW DG E++ FK++ F GPR C
Sbjct: 1203 PIKRGEDIFISVWNLHRSPTLW-NDADKFEPERWPLDGPNPNETNQGFKYLPFGGGPRKC 1379
Query: 280 LGKDFAYRQMKIVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGL 325
+G FA ++ + LVR F F++ V T T+H GL
Sbjct: 1380 IGDMFASYEVVVALAMLVRRFNFQMAVGAPPVVMTTGATIHTTQGL 1517
>TC82795 similar to GP|18000068|gb|AAL54885.1 cytochrome P450-dependent
fatty acid hydroxylase {Vicia sativa}, partial (36%)
Length = 845
Score = 112 bits (281), Expect = 2e-25
Identities = 66/166 (39%), Positives = 94/166 (55%), Gaps = 5/166 (3%)
Frame = +1
Query: 141 FYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISDATLDKMHYLHAALTETLRL 200
F++L K+ ++ ++ +E+ T + ++ DE + M Y HA+L E++RL
Sbjct: 10 FWLLSKHSHVENEILKEI-----TGKSEIVSYDE---------VKDMVYTHASLCESMRL 147
Query: 201 YPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWL---K 257
YP VP+ + A +D+LPDG V KG V Y YAMGR IWG D EF PERWL +
Sbjct: 148 YPPVPVDTKEAAYNDVLPDGTFVKKGWRVAYHIYAMGRSEKIWGLDWAEFRPERWLSQDE 327
Query: 258 DG--IFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFR 301
DG F + + F AGPR+CLGK+ A+ QMK V ++R FR
Sbjct: 328 DGKWSFIGMDPYSYAVFQAGPRVCLGKEMAFLQMKRVVAGIMRQFR 465
>TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 71D8 (EC
1.14.-.-) (P450 CP7). [Soybean] {Glycine max}, partial
(38%)
Length = 1724
Score = 103 bits (256), Expect = 1e-22
Identities = 71/249 (28%), Positives = 122/249 (48%), Gaps = 3/249 (1%)
Frame = +3
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKD---SSNMTD 115
G+ A + K +D+F ++K + +KD D+E + LLE K S ++T+
Sbjct: 786 GAIARVDNTFKALDEFFEQVLKEHLNPNNRKKD--DEEKDIVDVLLELKNQGRLSIDLTN 959
Query: 116 KYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEF 175
+++ +++N ++A DTSA T W L KNP +K +E+ N+ EF
Sbjct: 960 NHIKAVVMNLLVAATDTSAATSVWVMTGLIKNPRAMKKAQEEIRNIKK----------EF 1109
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
I + + K Y A + ETLR Y P++ R + L +GY + +V+ ++
Sbjct: 1110 ---IDEDDIQKFVYFKAVIKETLRFYSPAPLAPRETSKSFTL-NGYKIEPKTSVFVSIWS 1277
Query: 236 MGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA 295
+ R P W D EF PER+L + I +F+FI F AG RIC G +++++
Sbjct: 1278 IHRDPETW-KDPDEFYPERFLNNDIDFKGQNFEFIPFGAGRRICPGIPLGIATVEMITAN 1454
Query: 296 LVRFFRFKL 304
L+ F +++
Sbjct: 1455 LLNSFDWEM 1481
>AJ497661 similar to GP|10442763|gb cytochrome P450 {Triticum aestivum},
partial (23%)
Length = 490
Score = 102 bits (255), Expect = 2e-22
Identities = 54/119 (45%), Positives = 74/119 (61%), Gaps = 1/119 (0%)
Frame = +2
Query: 215 DILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHA 274
D+LP+G V G V Y Y++GRM +IWG+D EF PERW + +SS+KF+SF+A
Sbjct: 38 DVLPNGTFVPAGSMVTYSIYSVGRMKFIWGEDCLEFKPERWFSTDGEKMQSSYKFVSFNA 217
Query: 275 GPRICLGKDFAYRQMKIVSMALVRFFRFKLENETND-VTYRTMFTLHIDHGLPLYATPR 332
GPRICLGKD AY QMK ++ A++ R +LE V + TL + +GL + PR
Sbjct: 218 GPRICLGKDLAYLQMKSIAAAVL--LRHRLEVVPGHCVEQKMSLTLFMKYGLKVNVHPR 388
>TC83793 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P450
monooxygenase {Cicer arietinum}, partial (48%)
Length = 923
Score = 100 bits (249), Expect = 8e-22
Identities = 79/299 (26%), Positives = 132/299 (43%), Gaps = 2/299 (0%)
Frame = +2
Query: 35 NDSNAFVFKRYLDPLWKLKRVLNFGSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSD 94
N ++ F + LDP+ +R + K + I V +K R +
Sbjct: 17 NIADCFPVLKVLDPIGIRRRTGEY-----FGKLLGIFQGLVDQRLKMRE-----LNGYCG 166
Query: 95 KEDILSRFLLESKKDSSNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKV 154
K D+L +L+ +K++ M + + ++ +AG DT +TL W L NP I K
Sbjct: 167 KSDMLDA-MLDDEKNAGEMYKDKIERLSVDLFVAGTDTVTSTLEWAMAELLHNPNIMSKA 343
Query: 155 AQEVINVTSTSQESNLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEH 214
E+ + +++ ++ + K+ YL A + ET RL+PAVP+ E
Sbjct: 344 KSELNQIIGKG-----------NSVEESDIGKLPYLQAIIKETFRLHPAVPLLLPRKAEI 490
Query: 215 DILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHA 274
D+ +GY V KG V +A+GR +W ++ EFLPER+L I +F+ F
Sbjct: 491 DLEINGYKVPKGAQVLINVWAIGRDSNLW-ENPNEFLPERFLGSDIDFKGRNFELTPFGG 667
Query: 275 GPRICLGKDFAYRQMKIVSMALVRFFRFKLEN--ETNDVTYRTMFTLHIDHGLPLYATP 331
G RIC G A R + ++ + F ++L + D+ F L ++ PL P
Sbjct: 668 GRRICPGLPLAIRVLFLMLGLFINCFDWELVGGIKPEDMNMDDKFELTLEKAQPLLVVP 844
>TC87736 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P450
monooxygenase {Cicer arietinum}, partial (55%)
Length = 1367
Score = 99.4 bits (246), Expect = 2e-21
Identities = 74/272 (27%), Positives = 129/272 (47%)
Frame = +3
Query: 35 NDSNAFVFKRYLDPLWKLKRVLNFGSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSD 94
N ++ F R +DP +KR F + K I D+ + +K R F
Sbjct: 255 NMADFFPLLRLIDPQG-IKRTYVF----YVGKLFGIFDNIIDQKLKLREG-----DGFVT 404
Query: 95 KEDILSRFLLESKKDSSNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKV 154
D+L L E K + + ++ ++ + ++ G DT+ TL W L NP + KV
Sbjct: 405 NNDMLDSLLAEENK--KELDREKIQHLLHDLLVGGTDTTTYTLEWAMAELLHNPNVMSKV 578
Query: 155 AQEVINVTSTSQESNLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEH 214
+E+ E + + + I ++ + ++ YL A + ETLRL+P P+ +
Sbjct: 579 KKEL--------EETIGIG---NPIEESDVTRLPYLQAIIKETLRLHPIAPLLLPRKAKE 725
Query: 215 DILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHA 274
D+ +GY++ KG ++ +A+GR P +W D+ F PER+L + +F+ F +
Sbjct: 726 DVEVNGYLIPKGAQIFVNVWAIGRDPKVW-DNPNLFSPERFLGTKLDIKGQNFQLTPFGS 902
Query: 275 GPRICLGKDFAYRQMKIVSMALVRFFRFKLEN 306
G RIC G A R + ++ +L+ F +KLEN
Sbjct: 903 GRRICPGLPLAMRMLHMMLGSLLISFDWKLEN 998
>TC90108 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (41%)
Length = 958
Score = 97.1 bits (240), Expect = 9e-21
Identities = 77/280 (27%), Positives = 131/280 (46%), Gaps = 7/280 (2%)
Frame = +1
Query: 60 SEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKE---DILSRFLLESKKDSSNMTDK 116
++A ++K K ID + +I + S+ K+ S++E D+L + E+ +TD
Sbjct: 97 AKAKMEKLHKEIDMILQDIIDDHK---SIHKEDSNEEYLVDVLLKIQQENYHSQHPLTDD 267
Query: 117 YLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFV 176
++ II + AG +TS+ T+ W + KNP + E+ EV V
Sbjct: 268 NIKSIIQDIFDAGTETSSTTVLWAISEMVKNPKVMEEAQAEVRRVFDRK----------- 414
Query: 177 SNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAM 236
+ + L ++ YL + + ET+RL+P VP+ +GY + V ++A+
Sbjct: 415 GFVDETELHQLIYLKSVIKETMRLHPTVPLLLPRESRERCQINGYEIPAKTRVMVNAWAI 594
Query: 237 GRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMAL 296
GR P W DA+ F PER++ I + F++I F AG ++C G FA +++ +L
Sbjct: 595 GRDPRYW-VDAESFKPERFVNSPIDFKGTDFEYIPFGAGRKMCPGIAFALPNVELPLASL 771
Query: 297 VRFFRFKL----ENETNDVTYRTMFTLHIDHGLPLYATPR 332
+ F +KL +NE D+T T H L L R
Sbjct: 772 LYHFDWKLPNKMKNEELDMTESFGITAGRKHNLCLIPITR 891
>TC87269 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soybean
(fragment), partial (79%)
Length = 1878
Score = 96.3 bits (238), Expect = 1e-20
Identities = 82/328 (25%), Positives = 147/328 (44%), Gaps = 10/328 (3%)
Frame = +1
Query: 3 CTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGSEA 62
C + ++G G + + L+ EA + F S+ F ++D ++K G+
Sbjct: 805 CDYEEEVELGSGQKRSRLQVLLNEAQALLAEFYFSDNFPLLGWID---RVK-----GTLG 960
Query: 63 ALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSS---NMTDKYLR 119
L K K +D +I + + K + D + LL+ D S ++T +++
Sbjct: 961 KLDKTFKELDLIYQRVIDDHMDNSARPKTKEQEVDDIIDILLQMMNDHSLSFDLTLDHIK 1140
Query: 120 DIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNI 179
+++N IAG DTS+ + W L NP + KV E+ NL E I
Sbjct: 1141 AVLMNIFIAGTDTSSAIVVWAMTTLMNNPRVMNKVQMEI-----------RNLYEDKDFI 1287
Query: 180 SDATLDKMHYLHAALTETLRLYPAVPM--SGRTAEEHDILPDGYIVNKGETVYYLSYAMG 237
++ ++K+ YL A + ET+RL+P P+ T E +I DGY + VY ++A+
Sbjct: 1288 NEDDIEKLPYLKAVVKETIRLFPPSPLLVPRETIENCNI--DGYEIKPKTLVYVNAWAIA 1461
Query: 238 RMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALV 297
R P W D +EF PER++ + +F+ I F +G R+C + +++ L+
Sbjct: 1462 RDPENW-KDPEEFYPERFIISSVDFKGKNFELIPFGSGRRMCPAMNMGVVTVELTLANLL 1638
Query: 298 RFFRFKL-----ENETNDVTYRTMFTLH 320
+ F + L + + D + T+H
Sbjct: 1639 QSFDWNLPHGFDKEQVLDTQVKPGITMH 1722
>TC86135 similar to PIR|A96673|A96673 probable cytochrome P450 F13O11.25
[imported] - Arabidopsis thaliana, partial (62%)
Length = 1842
Score = 94.4 bits (233), Expect = 6e-20
Identities = 65/232 (28%), Positives = 110/232 (47%), Gaps = 3/232 (1%)
Frame = +2
Query: 104 LESKKDSSNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTS 163
LE ++ +++ + ++ F+ G DT++ L W L K P +Q ++ +E+ V
Sbjct: 860 LELPEEKRKLSENEMVNLCSEFLNGGTDTTSTALQWIMANLVKYPEVQGRLVEEIREVMV 1039
Query: 164 TSQESNLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIV 223
+ + + L K+ YL + E LR +P A D++ +GY+V
Sbjct: 1040 SDDNGEKE------EVKEENLQKLRYLKCVVLEGLRRHPPGHFVLPHAVSEDVVLNGYLV 1201
Query: 224 NKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESS--FKFISFHAGPRICLG 281
K TV ++ MG P +W +D EF PER+LKD F S K + F AG RIC G
Sbjct: 1202 PKDGTVNFMVAEMGLDPRVW-EDPMEFKPERFLKDETFDITGSKEIKMMPFGAGRRICPG 1378
Query: 282 KDFAYRQMKIVSMALVRFFRFKL-ENETNDVTYRTMFTLHIDHGLPLYATPR 332
+ A ++ LV F +K+ E D++ + FT+ + + L + +PR
Sbjct: 1379 YNLALLHLEYFVANLVWNFDWKVPEGGRVDLSEKQEFTMVMKNPLQAHISPR 1534
>TC79360 similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (52%)
Length = 1134
Score = 93.6 bits (231), Expect = 1e-19
Identities = 64/236 (27%), Positives = 105/236 (44%)
Frame = +1
Query: 71 IDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRDIILNFMIAGK 130
ID + +I R S D D+L + E+ +TD+ ++ +I + IAG
Sbjct: 139 IDRILQDIINDHRNNHSKTSKDEDLVDVLLKVQHENVHSQQPLTDENIKSVIQDLFIAGS 318
Query: 131 DTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNISDATLDKMHYL 190
+TS+ + W + KNP++ E+ EV V + + L ++ YL
Sbjct: 319 ETSSGIVLWAMSEMIKNPIVMEEAQVEVRRVFDKK-----------GYVDETELQQLTYL 465
Query: 191 HAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEF 250
+ ET RL+P VP+ +GY + V +A+GR P W +A+ F
Sbjct: 466 KCVIKETFRLHPTVPLLVPRESRERCEINGYEIPAKTRVAVNVWAIGRDPKYW-VEAESF 642
Query: 251 LPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFFRFKLEN 306
PER++ I + F+ I F AG R+C G FA +++ L+ F +KL N
Sbjct: 643 KPERFVNSSIDFKGTDFELIPFGAGRRMCPGIAFALPNVELPLAKLLYHFDWKLPN 810
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.138 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,982,244
Number of Sequences: 36976
Number of extensions: 126448
Number of successful extensions: 838
Number of sequences better than 10.0: 161
Number of HSP's better than 10.0 without gapping: 731
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 731
length of query: 337
length of database: 9,014,727
effective HSP length: 97
effective length of query: 240
effective length of database: 5,428,055
effective search space: 1302733200
effective search space used: 1302733200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146342.8


