

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
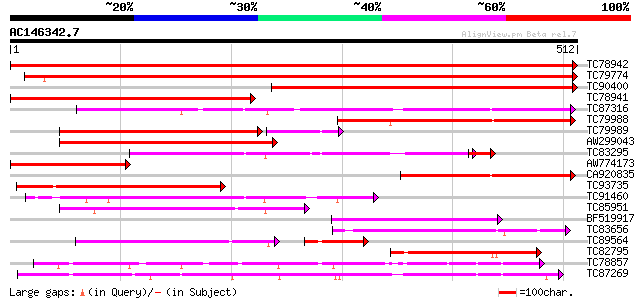
Query= AC146342.7 - phase: 0
(512 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78942 weakly similar to GP|20146758|gb|AAM12494.1 cytochrome P... 950 0.0
TC79774 weakly similar to GP|20146747|gb|AAM12483.1 cytochrome P... 631 0.0
TC90400 similar to GP|20146747|gb|AAM12483.1 cytochrome P450-lik... 546 e-156
TC78941 similar to GP|10334505|emb|CAC10214. hypothetical protei... 441 e-124
TC87316 weakly similar to PIR|G84675|G84675 probable cytochrome ... 283 1e-76
TC79988 similar to PIR|T00864|T00864 cytochrome P450 homolog F17... 267 6e-72
TC79989 similar to PIR|T00864|T00864 cytochrome P450 homolog F17... 215 4e-64
AW299043 weakly similar to PIR|T00864|T00 cytochrome P450 homolo... 226 2e-59
TC83295 similar to SP|O81117|C941_VICSA Cytochrome P450 94A1 (EC... 214 2e-57
AW774173 similar to GP|10334505|emb hypothetical protein {Cicer ... 216 2e-56
CA920835 weakly similar to GP|5734639|dbj ESTs AU056036(S20239) ... 196 2e-50
TC93735 similar to GP|20146758|gb|AAM12494.1 cytochrome P450-lik... 177 7e-45
TC91460 similar to GP|19698839|gb|AAL91155.1 cytochrome P450 {Ar... 156 2e-38
TC85951 similar to GP|18000068|gb|AAL54885.1 cytochrome P450-dep... 142 4e-34
BF519917 similar to GP|19698839|gb cytochrome P450 {Arabidopsis ... 130 9e-31
TC83656 similar to GP|19698839|gb|AAL91155.1 cytochrome P450 {Ar... 127 8e-30
TC89564 weakly similar to PIR|G84675|G84675 probable cytochrome ... 99 2e-29
TC82795 similar to GP|18000068|gb|AAL54885.1 cytochrome P450-dep... 124 9e-29
TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 7... 119 4e-27
TC87269 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soy... 107 1e-23
>TC78942 weakly similar to GP|20146758|gb|AAM12494.1 cytochrome P450-like
protein {Oryza sativa (japonica cultivar-group)},
partial (58%)
Length = 1719
Score = 950 bits (2456), Expect = 0.0
Identities = 472/513 (92%), Positives = 490/513 (95%), Gaps = 1/513 (0%)
Frame = +3
Query: 1 MLFQSIMDFLSNPFLFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMN 60
MLFQSIMDFLSNP LFAALSASLTLLLVQLLLRKLNNKSN+MKKKKYHPVAGTVFNQMMN
Sbjct: 18 MLFQSIMDFLSNPLLFAALSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMN 197
Query: 61 FNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDL 120
FNRLHHYMTDLARKY+TYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDL
Sbjct: 198 FNRLHHYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDL 377
Query: 121 LGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEI 180
LGDGIF VDGEKWREQRKISSHEFST+ML+DFSTSIFRKNAAKVANIVSEAA SN KLEI
Sbjct: 378 LGDGIFTVDGEKWREQRKISSHEFSTRMLKDFSTSIFRKNAAKVANIVSEAANSNTKLEI 557
Query: 181 QDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFL 240
QD+ MKSTLDSIF V FGTE++SMCG+SEEGKNFAN+FD ASALTLYRYVDVFWKIKKFL
Sbjct: 558 QDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFL 737
Query: 241 NIGSEAALRKNTEVLNEFVIKLINTRIQQM-NSKGDSIRKSGDILSRFLQVKEYDTTYLR 299
NIGSEAALR NTE+LNEFVIKLINTRIQQM NSKGDS+RK GDILSRFLQVKEYDT YLR
Sbjct: 738 NIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDTKYLR 917
Query: 300 DIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVSCVTD 359
DIILNFVIAGKDTT TLSWFMYMLCKYPAVQEKAA+EVREATNTKTVSSCTEFVS VTD
Sbjct: 918 DIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTD 1097
Query: 360 EALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKF 419
EA+EKMNY+HA LTETLRLYPA+P DAKICFADDTLPDGYSVKK DMVSYQPYAMGRMKF
Sbjct: 1098EAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKRDMVSYQPYAMGRMKF 1277
Query: 420 IWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCF 479
IWGDDAEEFRP RWLDENG FQ E PFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCF
Sbjct: 1278IWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCF 1457
Query: 480 RFKLNDEKRNVTYKTMINLHIDGGLEIKALHRD 512
RFKLNDEK+NVTYKTMI LHIDGGLEIKAL+R+
Sbjct: 1458RFKLNDEKKNVTYKTMITLHIDGGLEIKALYRN 1556
>TC79774 weakly similar to GP|20146747|gb|AAM12483.1 cytochrome P450-like
protein {Oryza sativa (japonica cultivar-group)},
partial (52%)
Length = 1609
Score = 631 bits (1628), Expect = 0.0
Identities = 305/502 (60%), Positives = 393/502 (77%), Gaps = 3/502 (0%)
Frame = +2
Query: 14 FLFAALSASLTLLLVQL--LLRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMTDL 71
FLFA L+ + L + K++ KK++YHPVAGTV +Q+ NF+RL YMTDL
Sbjct: 86 FLFALKPLFPILIAIALAGFIIKIHGSRFFDKKRRYHPVAGTVLHQLFNFHRLLEYMTDL 265
Query: 72 ARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVDGE 131
K +TYRLL+ RSEVYTS+P+N+E+++ TNF NYGKG Y++ L+DLLGDGIF VDGE
Sbjct: 266 TSKRKTYRLLSFNRSEVYTSDPANIEHMMATNFSNYGKGWYHHSVLEDLLGDGIFTVDGE 445
Query: 132 KWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQDLLMKSTLDS 191
KWR QRK +S++FSTK+LRDFS+S+F+ NA K+A IVSEAATSN +E+QDL MKSTLDS
Sbjct: 446 KWRHQRKSASYQFSTKLLRDFSSSVFKSNAVKLAGIVSEAATSNNIIELQDLFMKSTLDS 625
Query: 192 IFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLNIGSEAALRKN 251
+F+ G EL++MCG+ EG F+NAFD ASA ++RYV+ WK+++FLN+GSEA L+KN
Sbjct: 626 VFKGILGVELDTMCGTYREGTQFSNAFDEASAAIMFRYVNFLWKVQRFLNMGSEAVLKKN 805
Query: 252 TEVLNEFVIKLINTRIQQMNS-KGDSIRKSGDILSRFLQVKEYDTTYLRDIILNFVIAGK 310
V++E+V +I ++I+Q + +S GDILSRFL++ E D+ YL+D+IL+F+IAGK
Sbjct: 806 LRVIDEYVYTVIRSKIEQSQKPQNNSSELKGDILSRFLELNETDSKYLKDVILSFIIAGK 985
Query: 311 DTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVSCVTDEALEKMNYLHA 370
DTTA TLSWF+Y LCK+P VQEK A+E+ EAT + S+ E + +T+E++EKM YLHA
Sbjct: 986 DTTAITLSWFLYQLCKHPHVQEKIAQEIMEATKVEDGSTIDELAARLTEESMEKMQYLHA 1165
Query: 371 TLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRP 430
LTETLRL+P VPV++K CF+DDTLPDGYSV KGD+VS+QPY MGRMKF+WG+DAE+FRP
Sbjct: 1166ALTETLRLHPPVPVESKYCFSDDTLPDGYSVTKGDLVSFQPYVMGRMKFLWGEDAEQFRP 1345
Query: 431 ARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKRNV 490
RWLDENGNFQ E+PFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLG FKL D+ + V
Sbjct: 1346ERWLDENGNFQRESPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGSHSFKLADQNKLV 1525
Query: 491 TYKTMINLHIDGGLEIKALHRD 512
Y+T + L ID GL + A HR+
Sbjct: 1526KYRTSLTLQIDDGLHVNAFHRN 1591
>TC90400 similar to GP|20146747|gb|AAM12483.1 cytochrome P450-like protein
{Oryza sativa (japonica cultivar-group)}, partial (37%)
Length = 967
Score = 546 bits (1406), Expect = e-156
Identities = 270/276 (97%), Positives = 271/276 (97%)
Frame = +1
Query: 237 KKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKEYDTT 296
K+ SEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKEYDTT
Sbjct: 1 KEVSQFRSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKEYDTT 180
Query: 297 YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVSC 356
YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVSC
Sbjct: 181 YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVSC 360
Query: 357 VTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGR 416
VTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGR
Sbjct: 361 VTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGR 540
Query: 417 MKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLL 476
MKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLL
Sbjct: 541 MKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLL 720
Query: 477 GCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHRD 512
GCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHRD
Sbjct: 721 GCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHRD 828
>TC78941 similar to GP|10334505|emb|CAC10214. hypothetical protein {Cicer
arietinum}, partial (77%)
Length = 733
Score = 441 bits (1133), Expect = e-124
Identities = 222/222 (100%), Positives = 222/222 (100%)
Frame = +1
Query: 1 MLFQSIMDFLSNPFLFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMN 60
MLFQSIMDFLSNPFLFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMN
Sbjct: 67 MLFQSIMDFLSNPFLFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMN 246
Query: 61 FNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDL 120
FNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDL
Sbjct: 247 FNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDL 426
Query: 121 LGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEI 180
LGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEI
Sbjct: 427 LGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEI 606
Query: 181 QDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTAS 222
QDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTAS
Sbjct: 607 QDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTAS 732
>TC87316 weakly similar to PIR|G84675|G84675 probable cytochrome P450
[imported] - Arabidopsis thaliana, partial (77%)
Length = 1678
Score = 283 bits (724), Expect = 1e-76
Identities = 174/471 (36%), Positives = 253/471 (52%), Gaps = 20/471 (4%)
Frame = +2
Query: 61 FNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDL 120
F L Y T L ++ T + TS P NVEYILKTNF NY KG L DL
Sbjct: 224 FVNLVDYYTHLLQESPTGTIHVHVLGNTITSNPENVEYILKTNFNNYPKGKQFSTILGDL 403
Query: 121 LGDGIFAVDGEKWREQRKISSHEFSTKMLRDFS----------------TSIFRKNAAKV 164
LG GIF VDG W+ QRK++S E + ++R ++ S+ K A
Sbjct: 404 LGRGIFNVDGHSWKFQRKMASLELGSVVIRSYAMELVIEEIKTRLLPLIASVAEKKTASK 583
Query: 165 ANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASAL 224
A+ SE + L++QD+L + + D+I + +FG + + S N ANAFD +S L
Sbjct: 584 ADNTSE----DVLLDMQDILRRFSFDNICKFSFGLDPCCLVPSLPVS-NLANAFDLSSTL 748
Query: 225 TLYRYVD---VFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSG 281
+ R + + WK+K+F NIGSE L++ +++N+ L N I+Q + +
Sbjct: 749 SAQRALTASPLIWKMKRFFNIGSEKKLKEAIKIVND----LANEMIKQRREIENGVESRK 916
Query: 282 DILSRFL-QVKEYDTTYLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVRE 340
D+LSRF+ + +D YLRDI+++F++AG+DT A+ L+ F +L K P V+EK E+
Sbjct: 917 DLLSRFMGALNSHDDEYLRDIVVSFLLAGRDTVASALTGFFILLSKNPKVEEKIRVELDR 1096
Query: 341 ATNTKTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYS 400
N C T E +M+YL+ + E++RL+P V D+K DD LPDG
Sbjct: 1097VMNPNQ--------ECATFEQTREMHYLNGAIHESMRLFPPVQFDSKFALEDDVLPDGTF 1252
Query: 401 VKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLG 460
+KKG V+Y PYAMGRM+ IWG D EF+P RWL ++G F + PFK+ FQAG R+CLG
Sbjct: 1253IKKGSRVTYHPYAMGRMENIWGPDCLEFKPERWL-KDGVFVPKCPFKYPVFQAGSRVCLG 1429
Query: 461 KEFAYRQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHR 511
KE A +MK A L+ F ++ + + + GGL +K R
Sbjct: 1430KELAIVEMKSVVASLVKRFDVRVVGPNQEPQFAPGLTASFRGGLPVKIYER 1582
>TC79988 similar to PIR|T00864|T00864 cytochrome P450 homolog F17K2.4 -
Arabidopsis thaliana, partial (39%)
Length = 808
Score = 267 bits (683), Expect = 6e-72
Identities = 131/217 (60%), Positives = 164/217 (75%), Gaps = 2/217 (0%)
Frame = +2
Query: 297 YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREAT--NTKTVSSCTEFV 354
YLRDIILNF+IAGKDTTA TLSWF YMLCK P V++K +E+R+ T + ++S EFV
Sbjct: 17 YLRDIILNFMIAGKDTTANTLSWFFYMLCKNPLVEDKIVQEIRDVTCSHESELNSIDEFV 196
Query: 355 SCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAM 414
+ +TD L+KM+YLHATLTETLRLYP +PVD + A D LPDG+ ++KGD V Y YAM
Sbjct: 197 ANLTDLILDKMHYLHATLTETLRLYPVLPVDGRTADAPDVLPDGHKLEKGDGVYYLAYAM 376
Query: 415 GRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAV 474
GRM IWG+DA+EFRP RW+ + G FQ E+PFKF AF AGPR+CLGK+FAYRQMKI +
Sbjct: 377 GRMSSIWGEDADEFRPERWITD-GIFQPESPFKFVAFHAGPRMCLGKDFAYRQMKIVAMC 553
Query: 475 LLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHR 511
+L F+FKL + +NVTYK M LH+D GL + A+ R
Sbjct: 554 VLNFFKFKLANGTQNVTYKVMFTLHLDKGLPLTAIPR 664
>TC79989 similar to PIR|T00864|T00864 cytochrome P450 homolog F17K2.4 -
Arabidopsis thaliana, partial (22%)
Length = 806
Score = 215 bits (548), Expect(2) = 4e-64
Identities = 103/183 (56%), Positives = 137/183 (74%)
Frame = +2
Query: 46 KYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFE 105
KY PV GTVFNQ+ NF +LH Y LA+ + T RLL P +SE+YT + N+E+ILKTNF+
Sbjct: 17 KYAPVKGTVFNQLFNFKKLHDYHAQLAKTHPTMRLLAPNQSELYTIDVRNIEHILKTNFD 196
Query: 106 NYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVA 165
Y KG YN + DLLG+GIFAVDG+KW++QRKI+S+EFST++LRDFS S+FRKNA+K+
Sbjct: 197 KYSKGKYNKDIITDLLGEGIFAVDGDKWKQQRKIASYEFSTRVLRDFSCSVFRKNASKLV 376
Query: 166 NIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALT 225
++SE + ++QDL M+ LDSIF+V FGTELN + GSS+EG F AFD ++AL
Sbjct: 377 RVISEFSHEGLVFDMQDLQMRCALDSIFKVGFGTELNCLEGSSKEGTEFMKAFDESNALI 556
Query: 226 LYR 228
+R
Sbjct: 557 YWR 565
Score = 48.1 bits (113), Expect(2) = 4e-64
Identities = 27/69 (39%), Positives = 40/69 (57%)
Frame = +1
Query: 233 FWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKE 292
FW +K+FLNIG EA L+ N +++++FV +INT+ Q+ + DS K D S F
Sbjct: 580 FWSLKRFLNIGGEAKLKHNVKLIDDFVNGVINTKKSQLELQQDSNVKE-DYYSSF*WKAR 756
Query: 293 YDTTYLRDI 301
+ T L I
Sbjct: 757 WPNTILISI 783
>AW299043 weakly similar to PIR|T00864|T00 cytochrome P450 homolog F17K2.4 -
Arabidopsis thaliana, partial (34%)
Length = 594
Score = 226 bits (575), Expect = 2e-59
Identities = 107/197 (54%), Positives = 142/197 (71%)
Frame = +2
Query: 46 KYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFE 105
KY PV GTVFNQ+ F +L+ Y LA+ + TYRLL P SE+YT + N+E++LKTNF+
Sbjct: 2 KYTPVKGTVFNQLFYFKKLYDYHAQLAKIHPTYRLLAPNHSELYTIDVRNIEHVLKTNFD 181
Query: 106 NYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVA 165
Y +G Y+ + DL G+GIF VDG KW++QRK++S+EFST++LRDFS S+FRKNAAK+
Sbjct: 182 KYSRGKYSQDIMTDLFGEGIFVVDGNKWKQQRKVASYEFSTRVLRDFSCSVFRKNAAKLV 361
Query: 166 NIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALT 225
++SE ++QDL M+ LDSIF+V FGTELN + GSS+EG F AFD ++AL
Sbjct: 362 RVISEFYHEGLVFDMQDLQMRCALDSIFKVGFGTELNCLEGSSKEGTEFMKAFDESNALI 541
Query: 226 LYRYVDVFWKIKKFLNI 242
RYVD W +KKFLNI
Sbjct: 542 YLRYVDPIWSLKKFLNI 592
>TC83295 similar to SP|O81117|C941_VICSA Cytochrome P450 94A1 (EC 1.14.-.-)
(P450-dependent fatty acid omega-hydroxylase)., partial
(62%)
Length = 971
Score = 214 bits (544), Expect(2) = 2e-57
Identities = 117/318 (36%), Positives = 192/318 (59%), Gaps = 4/318 (1%)
Frame = +2
Query: 109 KGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAA-KVANI 167
KG + L D LG+GIF +GE W+ QR+++SHEF+TK LR F + ++ I
Sbjct: 5 KGTIFTKTLSDFLGNGIFNTNGENWKFQRQVASHEFNTKSLRKFVEDVVDTELTNRLIPI 184
Query: 168 VSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLY 227
+S + + L+ QD+L + T D+I ++AFG + + S++ K FA AF+ A+ ++
Sbjct: 185 LSSSTQTKQILDFQDILQRFTFDNICKIAFGFDPEYLTPSAKRTK-FAQAFEDATEISSR 361
Query: 228 RY---VDVFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDIL 284
R+ + V WK+KK NIGSE LR+ + E+ K++ + +++ + DS+ D+L
Sbjct: 362 RFRLPLPVIWKMKKRFNIGSEKRLREAVTEIREYANKIVREKKKELK-ENDSLHTQ-DML 535
Query: 285 SRFLQVKEYDTTYLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNT 344
SRFL + ++ DI+++F++AGKDTT+A L+WF ++L K P V+E+ +E+ + + +
Sbjct: 536 SRFLSSGHSEEDFVTDIVISFILAGKDTTSAALTWFFWLLWKNPRVEEEILKEINKKSES 715
Query: 345 KTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKG 404
+ + ++ M Y HA L+E++RLYP VP+D+K DD L DG VKKG
Sbjct: 716 ------------LDYDEVKTMVYTHAALSESMRLYPPVPMDSKEAINDDVLLDGIVVKKG 859
Query: 405 DMVSYQPYAMGRMKFIWG 422
M++Y YAMGRM+ +WG
Sbjct: 860 TMITYHVYAMGRMESLWG 913
Score = 27.3 bits (59), Expect(2) = 2e-57
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = +3
Query: 415 GRMKFIWGDDAEEFRPARWLDENG 438
G + G+D EF+P RWL+ +G
Sbjct: 891 GGWRVYGGEDWAEFKPERWLENDG 962
>AW774173 similar to GP|10334505|emb hypothetical protein {Cicer arietinum},
partial (65%)
Length = 639
Score = 216 bits (549), Expect = 2e-56
Identities = 106/109 (97%), Positives = 108/109 (98%)
Frame = +3
Query: 1 MLFQSIMDFLSNPFLFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMN 60
MLFQSIMDFLSNP LFAALSASLTLLLVQLLLRKLNNKSN+MKKKKYHPVAGTVFNQMMN
Sbjct: 18 MLFQSIMDFLSNPLLFAALSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMN 197
Query: 61 FNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGK 109
FNRLHHYMTDLARKY+TYRLLNPFRSEVYTSEPSNVEYILKTNFENYGK
Sbjct: 198 FNRLHHYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGK 344
>CA920835 weakly similar to GP|5734639|dbj ESTs AU056036(S20239)
C72753(E2173) AU056035(S20239) correspond to a region
of the predicted gene.~, partial (36%)
Length = 809
Score = 196 bits (497), Expect = 2e-50
Identities = 97/158 (61%), Positives = 116/158 (73%)
Frame = -2
Query: 354 VSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYA 413
VS ++D L+KM+YLHA LTETLRLYPAVP+ + D LPDGY V KG+ V Y YA
Sbjct: 733 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 554
Query: 414 MGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSA 473
MGRM +IWGDDA+EF P RWL ++G FQ E+ FKF +F AGPRICLGK+FAYRQMKI S
Sbjct: 553 MGRMPYIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSM 377
Query: 474 VLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHR 511
L+ FRFKL +E +VTY+TM LHID GL + A R
Sbjct: 376 ALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 263
>TC93735 similar to GP|20146758|gb|AAM12494.1 cytochrome P450-like protein
{Oryza sativa (japonica cultivar-group)}, partial (18%)
Length = 647
Score = 177 bits (450), Expect = 7e-45
Identities = 92/189 (48%), Positives = 126/189 (65%)
Frame = +3
Query: 7 MDFLSNPFLFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHH 66
M FL+ + L L LL+ + L +S + KY PV GTVF+Q++NF L H
Sbjct: 48 MLFLTMFYFILILFLCLFLLISIITLSTFIGQS--IGNPKYPPVKGTVFHQLINFKTLFH 221
Query: 67 YMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIF 126
Y T +A+ T+RLL P +S++YT + NVE+ILKT F Y KG +N+ + DLLGDGIF
Sbjct: 222 YFTKIAQTSPTFRLLAPKKSKIYTCDTKNVEHILKTQFHKYSKGKFNHDTIFDLLGDGIF 401
Query: 127 AVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQDLLMK 186
AVDG KWR QRK+SS EFST++LRDFS ++FR+NA K+ +V + + +IQD++MK
Sbjct: 402 AVDGAKWRHQRKLSSLEFSTRVLRDFSCNVFRRNAVKLVRVVLGFSNAGLAFDIQDIVMK 581
Query: 187 STLDSIFQV 195
TLDSIF+V
Sbjct: 582 CTLDSIFKV 608
>TC91460 similar to GP|19698839|gb|AAL91155.1 cytochrome P450 {Arabidopsis
thaliana}, partial (59%)
Length = 976
Score = 156 bits (395), Expect = 2e-38
Identities = 108/338 (31%), Positives = 178/338 (51%), Gaps = 19/338 (5%)
Frame = +2
Query: 15 LFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYM------ 68
L AALSA L LL R L K P G++ N NR+H ++
Sbjct: 14 LTAALSAYF--LWFHLLARTLTGP-------KAWPFIGSLPGLFKNRNRVHDWIAENLRA 166
Query: 69 TDLARKYRTYRLLNPFRSE-----VYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGD 123
T ++ Y+T + PF + T P N+E+IL+T F+NY KG DLLG
Sbjct: 167 TGVSATYQTSIMPFPFLAHKQGFYTVTCHPKNLEHILRTRFDNYPKGPKWQTAFHDLLGQ 346
Query: 124 GIFAVDGEKWREQRKISSHEFSTKMLR-DFSTSIFRKNAAKVANIVSEAATSNFKLEIQD 182
GIF DGE W QRK ++ EF+ + L+ + + R ++ I+ ++ N +++QD
Sbjct: 347 GIFNSDGETWIMQRKTAALEFTARTLKLAMARWVNRSIKNRLWCILDKSVKDNVYVDLQD 526
Query: 183 LLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYR--YVDVFWKIKKFL 240
LL++ T D+I + G + ++ + E F+ AFDTA+ T+YR Y + W+ +K
Sbjct: 527 LLLRLTFDNICGLTLGKDPETLSPALPENP-FSVAFDTATEATMYRFLYPGLIWRFQKLF 703
Query: 241 NIGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKEYD-----T 295
IGSE L+++ +++ ++ I+ R + S D++SRF++ ++ D
Sbjct: 704 GIGSEKMLKQSLQIVETYMNNAISDRKE---------TPSDDLMSRFMKKRDIDGKPINA 856
Query: 296 TYLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEK 333
T L+ IILNF++AG+DT++ LSWF +++ +P V+EK
Sbjct: 857 TILQHIILNFILAGRDTSSVALSWFFWLVMNHPKVEEK 970
>TC85951 similar to GP|18000068|gb|AAL54885.1 cytochrome P450-dependent
fatty acid hydroxylase {Vicia sativa}, partial (45%)
Length = 742
Score = 142 bits (357), Expect = 4e-34
Identities = 85/233 (36%), Positives = 139/233 (59%), Gaps = 8/233 (3%)
Frame = +1
Query: 46 KYHPVAGTVFNQMMNFNRLHHYMTDLARKY--RTYRLLNPFRS-EVYTSEPSNVEYILKT 102
K +P+ G+VF+ NF+R H+++D+ + T+ L F S +V+T+ P+ V++ILKT
Sbjct: 43 KTYPIIGSVFSIYANFHRRVHWISDILQTIPSSTFILHRSFGSRQVFTANPAVVQHILKT 222
Query: 103 NFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNA- 161
NF Y KGL +++ D LGDGIF+ DGE W+ QR+ISSHEF+TK LR F ++
Sbjct: 223 NFPCYRKGLTLKRSVGDFLGDGIFSADGETWKFQRQISSHEFNTKSLRKFVETVVEVELN 402
Query: 162 AKVANIVSEAA-TSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDT 220
++ I+SEA+ L+ QD+L + T DSI ++AFG + + SS F AFD
Sbjct: 403 DRLLPILSEASKNQTVLLDFQDILQRFTFDSICRIAFGFDPEYL-HSSLPKTMFVKAFDD 579
Query: 221 ASALTLYRY---VDVFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQM 270
+S ++ R+ + + WK+KK LNIG E L++ + +++ + +++
Sbjct: 580 SSLISSVRFNAAIPLIWKVKKILNIGIERRLKEAVAEVRRLAYRIVREKKKEL 738
>BF519917 similar to GP|19698839|gb cytochrome P450 {Arabidopsis thaliana},
partial (26%)
Length = 505
Score = 130 bits (328), Expect = 9e-31
Identities = 70/157 (44%), Positives = 92/157 (58%), Gaps = 2/157 (1%)
Frame = +2
Query: 291 KEYDTTYLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREA-TNTKTVSS 349
K +D LR I LNFV+AG+DT++ LSWF +++ +P+V+EK E+ T+ S
Sbjct: 20 KPFDAGKLRHIALNFVLAGRDTSSVALSWFFWLVMNHPSVEEKILAELTAVLAETRGGDS 199
Query: 350 CTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSY 409
V E EK+ YL A L ETLRLYP+VP D K DD LPDG V G V+Y
Sbjct: 200 RRWTEEAVDFEEAEKLVYLKAALAETLRLYPSVPEDFKYAVDDDVLPDGTVVPAGSTVTY 379
Query: 410 QPYAMGRMKFIWGDDAEEFRPARWLD-ENGNFQAENP 445
Y++GRMK +WG+D EF+P RWL G + E P
Sbjct: 380 SIYSVGRMKSVWGEDCMEFKPDRWLSVHEGQTRFEPP 490
>TC83656 similar to GP|19698839|gb|AAL91155.1 cytochrome P450 {Arabidopsis
thaliana}, partial (39%)
Length = 929
Score = 127 bits (320), Expect = 8e-30
Identities = 80/220 (36%), Positives = 118/220 (53%), Gaps = 5/220 (2%)
Frame = +3
Query: 292 EYDTTYLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCT 351
++ T +L+ I+L + T L F +++ +P V+EK +E+
Sbjct: 9 DFTTHHLKLILL------EGTRHQLLQLFFWLVMNHPKVEEKIIKELTTVLEETRGGEKR 170
Query: 352 EFVSCVTD-EALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQ 410
++ D ++M YL A L ETLRLYP+VP D K DD PDG + G V+Y
Sbjct: 171 KWTEDPLDFSEADQMVYLKAALAETLRLYPSVPQDIKQAVVDDVFPDGTVIPAGSTVTYS 350
Query: 411 PYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENP---FKFTAFQAGPRICLGKEFAYRQ 467
Y++GRM+ IWG+D EF+P RWL G+ + E P F F AF AGPR CLGK+ AY Q
Sbjct: 351 IYSVGRMEKIWGEDCLEFKPERWLSVRGD-RFEPPKEGFMFVAFNAGPRTCLGKDLAYLQ 527
Query: 468 MKIFSAVLLGCFRFKLNDEKRNVTYKTM-INLHIDGGLEI 506
MK +A +L R++L +V + M + L + GL++
Sbjct: 528 MKSVAAAVL--LRYRLLPVPGHVVEQKMSLTLFMKNGLKV 641
>TC89564 weakly similar to PIR|G84675|G84675 probable cytochrome P450
[imported] - Arabidopsis thaliana, partial (46%)
Length = 1041
Score = 99.0 bits (245), Expect(2) = 2e-29
Identities = 65/189 (34%), Positives = 101/189 (53%), Gaps = 5/189 (2%)
Frame = +3
Query: 60 NFNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKD 119
+FN L + L +K T + + T+ NVEYILKT FENY KG L D
Sbjct: 252 DFNNLCDWYAHLLKKSPTKTIHIHVLRNIITANCENVEYILKTKFENYPKGKPFSSILGD 431
Query: 120 LLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAA-KVANIVSEAATSNFKL 178
LG GIF VDG+ W+ Q+K++S E + + +R F+ + ++ ++ +
Sbjct: 432 FLGRGIFNVDGDLWKFQKKMASLELNKQSIRSFAFEVVNNEIENRLIPLLQNQNLNQVVF 611
Query: 179 EIQDLLMKSTLDSIFQVAFGTELNSMC-GSSEEGKNFANAFDTASALTLYRYVDV---FW 234
++QD+ + + DSI + +FG L+ MC +S +FA +FD AS L+ R + V W
Sbjct: 612 DLQDVFKRFSFDSICRFSFG--LDPMCLETSLPMSDFALSFDLASKLSAERAMVVSPLIW 785
Query: 235 KIKKFLNIG 243
KIK+F N+G
Sbjct: 786 KIKRFFNMG 812
Score = 48.9 bits (115), Expect(2) = 2e-29
Identities = 24/58 (41%), Positives = 39/58 (66%)
Frame = +2
Query: 267 IQQMNSKGDSIRKSGDILSRFLQVKEYDTTYLRDIILNFVIAGKDTTAATLSWFMYML 324
I Q G S K D+LSRF+ +D +LRDI+++F++AG+DT A++L+ F ++L
Sbjct: 872 INQKRKLGFSHHK--DLLSRFMSTTIHDDMFLRDIVISFLLAGRDTVASSLTSFFWLL 1039
>TC82795 similar to GP|18000068|gb|AAL54885.1 cytochrome P450-dependent
fatty acid hydroxylase {Vicia sativa}, partial (36%)
Length = 845
Score = 124 bits (311), Expect = 9e-29
Identities = 69/140 (49%), Positives = 90/140 (64%), Gaps = 4/140 (2%)
Frame = +1
Query: 345 KTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKG 404
K ++ +E VS + ++ M Y HA+L E++RLYP VPVD K +D LPDG VKKG
Sbjct: 55 KEITGKSEIVSY---DEVKDMVYTHASLCESMRLYPPVPVDTKEAAYNDVLPDGTFVKKG 225
Query: 405 DMVSYQPYAMGRMKFIWGDDAEEFRPARWL--DENG--NFQAENPFKFTAFQAGPRICLG 460
V+Y YAMGR + IWG D EFRP RWL DE+G +F +P+ + FQAGPR+CLG
Sbjct: 226 WRVAYHIYAMGRSEKIWGLDWAEFRPERWLSQDEDGKWSFIGMDPYSYAVFQAGPRVCLG 405
Query: 461 KEFAYRQMKIFSAVLLGCFR 480
KE A+ QMK A ++ FR
Sbjct: 406 KEMAFLQMKRVVAGIMRQFR 465
>TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 71D8 (EC
1.14.-.-) (P450 CP7). [Soybean] {Glycine max}, partial
(38%)
Length = 1724
Score = 119 bits (297), Expect = 4e-27
Identities = 123/488 (25%), Positives = 220/488 (44%), Gaps = 26/488 (5%)
Frame = +3
Query: 22 SLTLLLVQLLLRKLNNKSNNM-----KKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYR 76
+L LL+ L ++ N N+ K K P+ G + ++ + LH +L++ Y
Sbjct: 108 ALPLLVFFLFMKYKTNIKNSSSSTFPKGPKGLPIIGNL--HQLDTSNLHLQFWNLSKIYG 281
Query: 77 T-YRLLNPFRSEVYTSEPSNVEYILKTNFEN-------YGKGLYNYQNLKDLLGDGIFAV 128
+ L F+ + + ILK + + +G +Y + D IF+
Sbjct: 282 PLFSLQIGFKKAIVVCSSKLAQEILKDHDHDVSSRPPSHGPKTLSYNGI-----DMIFSP 446
Query: 129 DGEKWREQRKISS-HEFSTKMLRDFS---TSIFRKNAAKVANIVSEAATSNFKLEIQDLL 184
+ WRE RKI H FS+K + F+ S + K++N V + SN + ++L
Sbjct: 447 YNDCWREIRKICVVHFFSSKKISSFAHVRKSEVKLMIEKISNHVCSSKISN----LSEVL 614
Query: 185 MKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLN--I 242
M + + ++AFG G E F + A+ L +V + +++
Sbjct: 615 MSVSSSIVCRIAFGKSYEHEGG---EKSRFHGLLNETQAIFLSFFVSDYIPFMGWVDKLT 785
Query: 243 GSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVK-------EYDT 295
G+ A + + L+EF +++ + N K D K DI+ L++K +
Sbjct: 786 GAIARVDNTFKALDEFFEQVLKEHLNPNNRKKDDEEK--DIVDVLLELKNQGRLSIDLTN 959
Query: 296 TYLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVS 355
+++ +++N ++A DT+AAT W M L K P +KA EE+R N K EF+
Sbjct: 960 NHIKAVVMNLLVAATDTSAATSVWVMTGLIKNPRAMKKAQEEIR---NIK-----KEFID 1115
Query: 356 CVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMG 415
++ ++K Y A + ETLR Y P+ + TL +GY ++ V +++
Sbjct: 1116---EDDIQKFVYFKAVIKETLRFYSPAPLAPRETSKSFTL-NGYKIEPKTSVFVSIWSIH 1283
Query: 416 RMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVL 475
R W D +EF P R+L+ + +F+ +N F+F F AG RIC G +++ +A L
Sbjct: 1284RDPETW-KDPDEFYPERFLNNDIDFKGQN-FEFIPFGAGRRICPGIPLGIATVEMITANL 1457
Query: 476 LGCFRFKL 483
L F +++
Sbjct: 1458LNSFDWEM 1481
>TC87269 similar to PIR|T05940|T05940 cytochrome P450 83D1p - soybean
(fragment), partial (79%)
Length = 1878
Score = 107 bits (266), Expect = 1e-23
Identities = 119/525 (22%), Positives = 221/525 (41%), Gaps = 32/525 (6%)
Frame = +1
Query: 8 DFLSNPFLFAALSASLTLLLVQLL-LRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHH 66
DF+ + +L L L ++ + +R + S+ K P+ G + ++ + HH
Sbjct: 214 DFIMGFLVLLSLLPILILFIIHIYKIRSTSRASSTPPGPKPLPLIGNL--HQLDPSSPHH 387
Query: 67 YMTDLARKYRTYRLLN-PFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGI 125
+ L++ Y L + + S E +LKT+ + ++ L+ L +G+
Sbjct: 388 SLWKLSKHYGPIMSLQLGYIPTLIVSSAKMAEQVLKTHDLKFASRP-SFLGLRKLSYNGL 564
Query: 126 ---FAVDGEKWREQRKIS-SHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQ 181
FA WRE RK+ H FS++ + F + A++ +S+ +
Sbjct: 565 DLAFAPYSPYWREMRKLCVQHLFSSQRVHSFRP-VRENEVAQLIQKLSQYGGDEKGANLS 741
Query: 182 DLLMKSTLDSIFQVAFGT------ELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWK 235
++LM T I ++AFG E GS ++ + A AL Y +
Sbjct: 742 EILMSLTNTIICKIAFGKTYVCDYEEEVELGSGQKRSRLQVLLNEAQALLAEFYFSDNFP 921
Query: 236 IKKFLNI--GSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKE- 292
+ +++ G+ L K + L+ ++I+ + ++ DI+ LQ+
Sbjct: 922 LLGWIDRVKGTLGKLDKTFKELDLIYQRVIDDHMDNSARPKTKEQEVDDIIDILLQMMND 1101
Query: 293 ----YDTT--YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKT 346
+D T +++ +++N IAG DT++A + W M L P V K E+R K
Sbjct: 1102HSLSFDLTLDHIKAVLMNIFIAGTDTSSAIVVWAMTTLMNNPRVMNKVQMEIRNLYEDK- 1278
Query: 347 VSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDM 406
+ ++ +EK+ YL A + ET+RL+P P+ ++ DGY +K +
Sbjct: 1279--------DFINEDDIEKLPYLKAVVKETIRLFPPSPLLVPRETIENCNIDGYEIKPKTL 1434
Query: 407 VSYQPYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYR 466
V +A+ R W D EEF P R++ + +F+ +N F+ F +G R+C
Sbjct: 1435VYVNAWAIARDPENW-KDPEEFYPERFIISSVDFKGKN-FELIPFGSGRRMCPAMNMGVV 1608
Query: 467 QMKIFSAVLLGCFRFKL-----------NDEKRNVTYKTMINLHI 500
+++ A LL F + L K +T I+LH+
Sbjct: 1609TVELTLANLLQSFDWNLPHGFDKEQVLDTQVKPGITMHKKIDLHL 1743
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.135 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,472,229
Number of Sequences: 36976
Number of extensions: 187872
Number of successful extensions: 1253
Number of sequences better than 10.0: 196
Number of HSP's better than 10.0 without gapping: 1123
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1129
length of query: 512
length of database: 9,014,727
effective HSP length: 100
effective length of query: 412
effective length of database: 5,317,127
effective search space: 2190656324
effective search space used: 2190656324
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146342.7


