

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146341.8 + phase: 0
(268 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
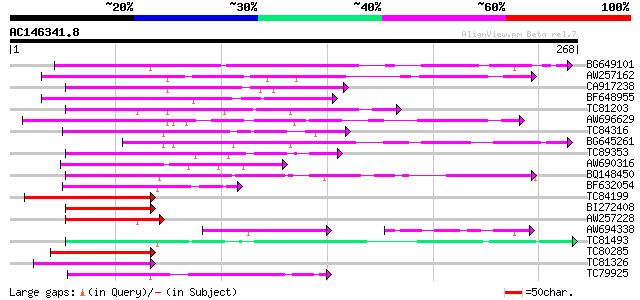
Score E
Sequences producing significant alignments: (bits) Value
BG649101 homologue to GP|6648199|gb|A unknown protein {Arabidops... 134 3e-32
AW257162 102 1e-22
CA917238 weakly similar to PIR|A84538|A845 hypothetical protein ... 91 5e-19
BF648955 similar to GP|18043667|gb| Unknown (protein for IMAGE:3... 82 2e-16
TC81203 weakly similar to GP|10177464|dbj|BAB10855. gb|AAD21700.... 81 5e-16
AW696629 74 8e-14
TC84316 weakly similar to GP|11994612|dbj|BAB02749. gb|AAC24186.... 69 2e-12
BG645261 homologue to PIR|H90254|H902 sulfate ABC transporter p... 66 1e-11
TC89353 weakly similar to GP|9279704|dbj|BAB01261.1 gb|AAF25964.... 62 3e-10
AW690316 60 7e-10
BQ148450 similar to GP|10177464|dbj gb|AAD21700.1~gene_id:MQB2.1... 57 6e-09
BF632054 similar to GP|10177628|dbj gb|AAF30317.1~gene_id:K6M13.... 57 6e-09
TC84199 similar to GP|9294656|dbj|BAB03005.1 gb|AAF30317.1~gene_... 50 7e-07
BI272408 50 1e-06
AW257228 GP|17342711|gb 1-aminocyclopropanecarboxylic acid oxida... 50 1e-06
AW694338 41 2e-06
TC81493 weakly similar to PIR|F86249|F86249 protein F25C20.23 [i... 48 5e-06
TC80285 similar to GP|7300092|gb|AAF55261.1| srp gene product {D... 47 8e-06
TC81326 weakly similar to PIR|T45651|T45651 hypothetical protein... 46 2e-05
TC79925 similar to GP|10177628|dbj|BAB10775. gb|AAF30317.1~gene_... 45 2e-05
>BG649101 homologue to GP|6648199|gb|A unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 799
Score = 134 bits (338), Expect = 3e-32
Identities = 97/252 (38%), Positives = 131/252 (51%), Gaps = 7/252 (2%)
Frame = +1
Query: 22 TLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQ---KQLKRPPRLILKP 78
++ + +P +LIA IL+ L VKTI +L +SKSWNTLI+ P+F++ Q + P IL P
Sbjct: 70 SVPVVIPSDLIAIILTFLPVKTITQLKLVSKSWNTLITSPSFIKIHLNQSSQNPNFILTP 249
Query: 79 PCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNK 138
Y + + S+PI RLL TVSGD ++ + + +VV SCNGLLC +F S+
Sbjct: 250 SRKQYSINNVLSVPIPRLLTGN--TVSGDTYHNILNNDHHFRVVGSCNGLLCLLFKSEFI 423
Query: 139 EYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLK 198
+ + FRIWNPAT T S+ LG Y FG G +
Sbjct: 424 THLKFRFRIWNPATRTISEELGFFRKY------------KPLFG------------GVSR 531
Query: 199 FTFGCGILSGTYKTVEFRAKGDEENKYGPWRSQVRILSL----SDDCWRNIDSFPVIPLI 254
FTFGC L GTYK V D + RS VR+ +L SD CWRNI + P +
Sbjct: 532 FTFGCDYLRGTYKLVALHTVEDGD----VMRSNVRVFNLGNDDSDKCWRNIPN----PFV 687
Query: 255 SPFNNGVYLSGT 266
+GV+LSGT
Sbjct: 688 CA--DGVHLSGT 717
>AW257162
Length = 728
Score = 102 bits (255), Expect = 1e-22
Identities = 79/240 (32%), Positives = 109/240 (44%), Gaps = 6/240 (2%)
Frame = +1
Query: 16 KKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPR-- 73
+ L+ S P+++IA+ILS L VKT++++ C+SKSWNTLISD FV+ L R R
Sbjct: 94 RSRMSLSSSPVFPDDIIAEILSWLTVKTLMKMKCVSKSWNTLISDSNFVKMHLNRSARHS 273
Query: 74 --LILKPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCK-VVISCNGLLC 130
++ Y + L+ SIT+ D Y + +C VV SCNGL+C
Sbjct: 274 QSYLVSEHRGDY---NFVPFSVRGLMNGRSITLPKDPYYQ--LIEKDCPGVVGSCNGLVC 438
Query: 131 FIFC-SDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYR 189
C +D +E+ W RIWNPAT T S L Y+
Sbjct: 439 LSGCVADVEEFEEMWLRIWNPATRTISDKL-------------------------YFSAN 543
Query: 190 RRYLLGCLKFTFGCGILSGTYKTVEFRAKGDEENKYGPWRSQVRILSLSDDCWRNIDSFP 249
R L C +F FG + TYK V + K G I+ ++ WRNI SFP
Sbjct: 544 R---LQCWEFMFGYDNTTQTYKVVALYPDSEMTTKVG-------IICFRNNIWRNIQSFP 693
>CA917238 weakly similar to PIR|A84538|A845 hypothetical protein At2g16220
[imported] - Arabidopsis thaliana, partial (6%)
Length = 603
Score = 90.9 bits (224), Expect = 5e-19
Identities = 54/140 (38%), Positives = 79/140 (55%), Gaps = 6/140 (4%)
Frame = +2
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPR----LILKPPCWV 82
LP+ELI ++LS DVK+++R+ C+S+ WN++ISDP FV+ +KR R + +
Sbjct: 146 LPDELITEVLSRGDVKSLMRMRCVSEYWNSMISDPRFVKLHMKRSARNAHLTLSLCKSGI 325
Query: 83 YPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLND-NCKVVI-SCNGLLCFIFCSDNKEY 140
+ P+ L++N IT+ D Y L D C+ VI SCNG LC + S Y
Sbjct: 326 DGDNNVVPYPVRGLIENGLITLPSDPYY---RLKDKECQYVIGSCNGWLCLLGFSSIGAY 496
Query: 141 HNYWFRIWNPATGTRSKALG 160
+ WFR WNPA G ++ LG
Sbjct: 497 RHIWFRFWNPAMGKMTQKLG 556
>BF648955 similar to GP|18043667|gb| Unknown (protein for IMAGE:3661715) {Mus
musculus}, partial (3%)
Length = 514
Score = 82.0 bits (201), Expect = 2e-16
Identities = 51/144 (35%), Positives = 77/144 (53%), Gaps = 4/144 (2%)
Frame = +3
Query: 16 KKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLI 75
K+++ + + FLP ELI +I+S L VK +++ C+SK + TLISDP FVQ L++ R
Sbjct: 24 KRHRSMASAAFLPSELIVEIISWLPVKYLMQFRCVSKFYKTLISDPYFVQMHLEKSARNP 203
Query: 76 LKPPCWVYPM----RTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCF 131
W + +I L ++RLL N T + G +C ++ SCNGLLC
Sbjct: 204 HLALMWQDDLLREDGSIIFLSVSRLLGNKYTTPP----FQSGTFYHSC-IIGSCNGLLCL 368
Query: 132 IFCSDNKEYHNYWFRIWNPATGTR 155
+ Y+ Y+ WNPAT T+
Sbjct: 369 VDFHCPDYYYYYYLYFWNPATRTK 440
>TC81203 weakly similar to GP|10177464|dbj|BAB10855.
gb|AAD21700.1~gene_id:MQB2.18~similar to unknown protein
{Arabidopsis thaliana}, partial (9%)
Length = 715
Score = 80.9 bits (198), Expect = 5e-16
Identities = 58/179 (32%), Positives = 87/179 (48%), Gaps = 20/179 (11%)
Frame = +1
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLIS-DPTFVQKQLKRPPR-----LILKPPC 80
L +ELI ILS L VKT+++ C+ KSW TLIS DP+F + L+R PR L+
Sbjct: 199 LLDELIVDILSRLPVKTLMQFKCVCKSWKTLISHDPSFAKLHLQRSPRNTHLTLVSDRSS 378
Query: 81 WVYPMRTIQSLPITRLLKNP-------------SITVSGDFCYDGGGLNDNCKVVISCNG 127
++ P++ L++ P ++ + D Y G + D C ++ SCNG
Sbjct: 379 DDESNFSVVPFPVSHLIEAPLTITPFEPYHLLRNVAIPDDPYYVLGNM-DCCIIIGSCNG 555
Query: 128 LLCF-IFCSDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCY 185
L C + +E + WFR WNPAT T S+ LG +++ R FG + Y
Sbjct: 556 LYCLRCYSLIYEEEEHDWFRFWNPATNTLSEELGCLNEF------FRLTFGYDISNDTY 714
>AW696629
Length = 663
Score = 73.6 bits (179), Expect = 8e-14
Identities = 68/250 (27%), Positives = 107/250 (42%), Gaps = 13/250 (5%)
Frame = +2
Query: 7 MSIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQK 66
M+ K+KK++ + + LP +LI QILS L VK ++R +SK W +LI DP F +
Sbjct: 5 MTEKRKKRESSEEATSSPPILPSDLIMQILSWLPVKLLIRFTSVSKHWKSLILDPNFAKL 184
Query: 67 QLKRPPR---LIL-----KPPCWV---YPMRTIQSLPITRLLKNPSITVSGDFCYDGGGL 115
L++ P+ +IL + WV YP+R++ LL+ S + C+D
Sbjct: 185 HLQKSPKNTHMILTALDDEDDTWVVTPYPVRSL-------LLEQSSFSDEECCCFD---- 331
Query: 116 NDNCKVVISCNGLLCFIF--CSDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLR 173
+ +V S NGL+C +N++Y + + WNP+ RSK ++
Sbjct: 332 YHSYFIVGSTNGLVCLAVEKSLENRKY-ELFIKFWNPSLRLRSK------------KAPS 472
Query: 174 FNFGSEAFGSCYYDYRRRYLLGCLKFTFGCGILSGTYKTVEFRAKGDEENKYGPWRSQVR 233
N G L G + FG L+ TYK V G R
Sbjct: 473 LNIG---------------LYGTARLGFGYDDLNDTYKAVAVFWDHTTHKMEG------R 589
Query: 234 ILSLSDDCWR 243
+ + D CW+
Sbjct: 590 VHCMGDSCWK 619
>TC84316 weakly similar to GP|11994612|dbj|BAB02749.
gb|AAC24186.1~gene_id:MGL6.4~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (10%)
Length = 892
Score = 68.9 bits (167), Expect = 2e-12
Identities = 51/141 (36%), Positives = 67/141 (47%), Gaps = 5/141 (3%)
Frame = +3
Query: 26 FLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRP---PRLILKPPCWV 82
FLPEELI IL L V+++LR C+ KSW TL SD F P+L+
Sbjct: 39 FLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVACESVSA 218
Query: 83 YPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHN 142
Y I++ PI LL+N S TV G + ++ SCNG LC Y N
Sbjct: 219 YRTWEIKTYPIESLLENSSTTV---IPVSNTG-HQRYTILGSCNGFLCL--------YDN 362
Query: 143 Y--WFRIWNPATGTRSKALGT 161
Y R+WNP+ +SK+ T
Sbjct: 363 YQRCVRLWNPSINLKSKSSPT 425
>BG645261 homologue to PIR|H90254|H902 sulfate ABC transporter permease
protein SSO1032 [imported] - Sulfolobus solfataricus,
partial (4%)
Length = 771
Score = 66.2 bits (160), Expect = 1e-11
Identities = 66/224 (29%), Positives = 93/224 (41%), Gaps = 11/224 (4%)
Frame = +1
Query: 54 WNTLISDPTFVQKQLKRP---PRLIL----KPPCWVYPMRTIQSLPITRLLKNPSITV-- 104
W ++I PTFV+ L R P L + W S + RLL+NP + +
Sbjct: 130 WKSIIDHPTFVKLHLNRSSQKPDLTFVSSARITLWTLESTCTVSFTVFRLLENPPVIINL 309
Query: 105 SGDFCYDGGGLNDNCKVVISCNGLLCF--IFCSDNKEYHNYWFRIWNPATGTRSKALGTN 162
S D Y D ++ SCNGLLC + +D+ + + R WNPA+ SK +G
Sbjct: 310 SDDDPYYQLKDKDCVHIIGSCNGLLCLFGLKINDSCGHMDISIRFWNPASRKISKKVGYC 489
Query: 163 HDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLKFTFGCGILSGTYKTVEFRAKGDEE 222
D G SC+ LKF FG S TYK V F
Sbjct: 490 GD------------GVNNDPSCFPGKG-----DLLKFVFGYVNSSDTYKVVYFIVG---- 606
Query: 223 NKYGPWRSQVRILSLSDDCWRNIDSFPVIPLISPFNNGVYLSGT 266
+ R+ SL D+ WR I++ PV+ I + V+LSG+
Sbjct: 607 ------TTSARVFSLGDNVWRKIENSPVV--IHHVSGFVHLSGS 714
>TC89353 weakly similar to GP|9279704|dbj|BAB01261.1
gb|AAF25964.1~gene_id:MYA6.2~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 1482
Score = 61.6 bits (148), Expect = 3e-10
Identities = 46/134 (34%), Positives = 66/134 (48%), Gaps = 3/134 (2%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVYPMR 86
+PEE+I +IL L V+++L+ C+ K W TLISDP F +K + +V +
Sbjct: 105 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 284
Query: 87 -TIQSLPITRLLKNPSI--TVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNY 143
+ S P+ LL NPS DF + ++ SCNGLLC SD Y
Sbjct: 285 CNLVSYPLKPLLDNPSAHRVEPADFEMIH---TTSMTIIGSCNGLLCL---SD-----FY 431
Query: 144 WFRIWNPATGTRSK 157
F +WNP+ +SK
Sbjct: 432 QFTLWNPSIKLKSK 473
>AW690316
Length = 606
Score = 60.5 bits (145), Expect = 7e-10
Identities = 41/118 (34%), Positives = 64/118 (53%), Gaps = 11/118 (9%)
Frame = +2
Query: 25 MFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVY- 83
+FLP+++I + L+ L+VK ++R+ C+ KSWNT+ISDP F + LK+ R K P +
Sbjct: 221 IFLPDDVIIEFLTFLEVKDLIRMKCVCKSWNTIISDPIFAKTHLKKKKR--TKKPHLAFL 394
Query: 84 -----PMRTIQSLPITRLL---KNPSITVSGDFC--YDGGGLNDNCKVVISCNGLLCF 131
+++PI+RLL K S ++ D Y + +V SCNGL F
Sbjct: 395 SDKSEGSGDCRAVPISRLLQEMKKSSSNLTHDDLPYYYRFNYKNYSDIVGSCNGLGLF 568
>BQ148450 similar to GP|10177464|dbj gb|AAD21700.1~gene_id:MQB2.18~similar to
unknown protein {Arabidopsis thaliana}, partial (9%)
Length = 666
Score = 57.4 bits (137), Expect = 6e-09
Identities = 61/227 (26%), Positives = 98/227 (42%), Gaps = 4/227 (1%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLK--RPPRLILKPPCWVYP 84
LP E+ +ILS L VK +++ C+ K W + IS P FV+K L+ L L + P
Sbjct: 3 LPFEIQVEILSRLPVKYLMQFQCVCKLWKSQISKPDFVKKHLRVSNTRHLFLLTFSKLSP 182
Query: 85 MRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNYW 144
I+S P++ + + T + Y +++ +V SC+G+LC I C N
Sbjct: 183 ELVIKSYPLSSVFTEMTPTFT-QLEYPLNNRDESDSMVGSCHGILC-IQC-------NLS 335
Query: 145 FRI-WNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLKFTFGC 203
F + WNP+ +K N +F + AFG YD+
Sbjct: 336 FPVLWNPSIRKFTKLPSFEFPQN------KFINPTYAFG---YDHS-------------- 446
Query: 204 GILSGTYKTVEFRAKGDEENKYGPWRSQVRILSLSDDCWRNIDS-FP 249
S TYK V + +N ++ V + ++ +CWR I + FP
Sbjct: 447 ---SDTYKVVAVFCTSNIDNGVYQLKTLVNVHTMGTNCWRRIQTEFP 578
>BF632054 similar to GP|10177628|dbj gb|AAF30317.1~gene_id:K6M13.17~similar
to unknown protein {Arabidopsis thaliana}, partial (8%)
Length = 462
Score = 57.4 bits (137), Expect = 6e-09
Identities = 39/94 (41%), Positives = 52/94 (54%), Gaps = 9/94 (9%)
Frame = +3
Query: 26 FLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQL---------KRPPRLIL 76
FLPEELI +IL L +K++LR C+ KSW +IS+P F++KQL R+IL
Sbjct: 141 FLPEELILEILIKLPIKSLLRFRCVCKSWLHIISNPYFIKKQLHFSTQNTHFTTNHRIIL 320
Query: 77 KPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCY 110
+ ++ S IT L NPS TVS D Y
Sbjct: 321 SATTAEFHLK---SCSITSLFNNPS-TVSDDLNY 410
>TC84199 similar to GP|9294656|dbj|BAB03005.1
gb|AAF30317.1~gene_id:F14O13.6~similar to unknown
protein {Arabidopsis thaliana}, partial (14%)
Length = 673
Score = 50.4 bits (119), Expect = 7e-07
Identities = 26/62 (41%), Positives = 38/62 (60%)
Frame = +1
Query: 8 SIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQ 67
S ++K++ K K+ + +LP ELI QIL L VK++LR C+ KSW +LISD F
Sbjct: 52 SCREKRRTVKKKEEKTAPYLPLELIIQILLWLPVKSLLRFKCVCKSWFSLISDTHFANSH 231
Query: 68 LK 69
+
Sbjct: 232 FQ 237
>BI272408
Length = 706
Score = 49.7 bits (117), Expect = 1e-06
Identities = 25/43 (58%), Positives = 32/43 (74%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLK 69
LP EL+A+IL L V +L+L CLSK +NTLISDP F +K L+
Sbjct: 291 LPFELVAEILCRLPVMFLLQLRCLSKFFNTLISDPKFAKKHLR 419
>AW257228 GP|17342711|gb 1-aminocyclopropanecarboxylic acid oxidase {Medicago
truncatula}, partial (11%)
Length = 702
Score = 49.7 bits (117), Expect = 1e-06
Identities = 26/48 (54%), Positives = 33/48 (68%), Gaps = 1/48 (2%)
Frame = +1
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLIS-DPTFVQKQLKRPPR 73
L +ELI +ILS L VKT+++ C+ KSW TLIS DP F + L R PR
Sbjct: 346 LLDELIVEILSRLPVKTLMQFKCVCKSWKTLISDDPVFAKFHLHRSPR 489
>AW694338
Length = 493
Score = 40.8 bits (94), Expect(2) = 2e-06
Identities = 23/64 (35%), Positives = 32/64 (49%), Gaps = 3/64 (4%)
Frame = +1
Query: 92 PITRLLKNPSITVSGDFCYD---GGGLNDNCKVVISCNGLLCFIFCSDNKEYHNYWFRIW 148
PIT++ +NPS TV + Y G +V SCNGL+ F + Y R+W
Sbjct: 85 PITQIFENPSFTVVDNPNYHHLINKGSQRGSFLVASCNGLILFSNSFPQIGFKEYRLRVW 264
Query: 149 NPAT 152
NPA+
Sbjct: 265 NPAS 276
Score = 27.3 bits (59), Expect(2) = 2e-06
Identities = 25/72 (34%), Positives = 34/72 (46%), Gaps = 1/72 (1%)
Frame = +2
Query: 178 SEAFGSCYYDYRRRYLLGCLKFTFGCGILSGTYKTVEFRAKGDEENKYGPWRSQVRILSL 237
SE G Y+ + +L G FGC + TYK V A ++E S+VR+L+L
Sbjct: 290 SEEIG--YFCHNDNFLFG-----FGCDNSNETYKVV---AYCNQET-----ASEVRVLNL 424
Query: 238 -SDDCWRNIDSF 248
DD WR F
Sbjct: 425 GGDDVWRKY*KF 460
>TC81493 weakly similar to PIR|F86249|F86249 protein F25C20.23 [imported] -
Arabidopsis thaliana, partial (9%)
Length = 728
Score = 47.8 bits (112), Expect = 5e-06
Identities = 59/244 (24%), Positives = 92/244 (37%), Gaps = 2/244 (0%)
Frame = +1
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQL--KRPPRLILKPPCWVYP 84
LP E+ +ILS + K +LRL K W LI F+ L R +IL+ +Y
Sbjct: 16 LPTEVTTEILSRVPAKPLLRLRSTCKWWRNLIDSTDFIFLHLSKSRDSVIILRQHSRLYE 195
Query: 85 MRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNYW 144
+ + S+ + L +P + CY ++ KV+ SCNGLLC +D+ + N
Sbjct: 196 L-DLNSMDRVKELDHPLM------CY-----SNRIKVLGSCNGLLCICNIADDIAFWNPT 339
Query: 145 FRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLKFTFGCG 204
R TN + + LL + FG
Sbjct: 340 IRKHRIIPSEPLIRKETNENNTITT-----------------------LLAAHVYGFGYD 450
Query: 205 ILSGTYKTVEFRAKGDEENKYGPWRSQVRILSLSDDCWRNIDSFPVIPLISPFNNGVYLS 264
+ YK V D N+ + S V I ++ D W + P L GV++S
Sbjct: 451 SATDDYKLVSISNFVDLHNR--SYDSHVTIYTMGSDVWMPLPGVP-YALCCARPMGVFVS 621
Query: 265 GTIY 268
G ++
Sbjct: 622 GALH 633
>TC80285 similar to GP|7300092|gb|AAF55261.1| srp gene product {Drosophila
melanogaster}, partial (2%)
Length = 1055
Score = 47.0 bits (110), Expect = 8e-06
Identities = 22/50 (44%), Positives = 35/50 (70%)
Frame = -2
Query: 20 KLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLK 69
+L ++ LPEELI +I+S +D + L+L + K WN+LI+DP FV+K +
Sbjct: 1018 ELPKTVVLPEELIIEIISRVDSRNTLQLRPVCKRWNSLITDPEFVKKHFQ 869
>TC81326 weakly similar to PIR|T45651|T45651 hypothetical protein F13I12.200
- Arabidopsis thaliana, partial (12%)
Length = 838
Score = 45.8 bits (107), Expect = 2e-05
Identities = 24/58 (41%), Positives = 33/58 (56%)
Frame = +3
Query: 12 KKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLK 69
+ ++K KK T +LP ELI QIL L VK++ R + KSW +LIS P F +
Sbjct: 123 ESSEEKKKKTTTLPYLPHELIFQILLRLPVKSLTRFKSVRKSWFSLISAPHFANSHFQ 296
>TC79925 similar to GP|10177628|dbj|BAB10775.
gb|AAF30317.1~gene_id:K6M13.17~similar to unknown
protein {Arabidopsis thaliana}, partial (34%)
Length = 652
Score = 45.4 bits (106), Expect = 2e-05
Identities = 40/127 (31%), Positives = 58/127 (45%), Gaps = 2/127 (1%)
Frame = +3
Query: 28 PEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQ--KQLKRPPRLILKPPCWVYPM 85
P+E++ QIL+ L VK++ R + K W L D F+Q + R +IL +
Sbjct: 267 PDEVVMQILARLPVKSLFRSKTVCKLWYRLTLDKYFIQLYNDVSRKNPMIL--------V 422
Query: 86 RTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNYWF 145
SL ++ + G F + LND KV SCNGLLC CS + Y+
Sbjct: 423 EISDSLLESKSSLICVDNLRGVFEFSLNFLNDRVKVRASCNGLLC---CSSIPDMGVYY- 590
Query: 146 RIWNPAT 152
+ NP T
Sbjct: 591 -VCNPVT 608
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.142 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,269,852
Number of Sequences: 36976
Number of extensions: 201524
Number of successful extensions: 1341
Number of sequences better than 10.0: 116
Number of HSP's better than 10.0 without gapping: 1306
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1319
length of query: 268
length of database: 9,014,727
effective HSP length: 94
effective length of query: 174
effective length of database: 5,538,983
effective search space: 963783042
effective search space used: 963783042
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146341.8


