

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
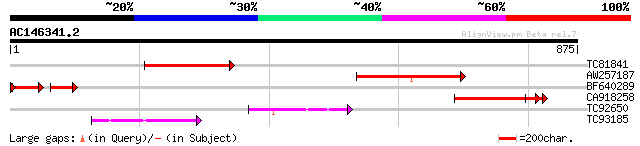
Query= AC146341.2 - phase: 0 /pseudo
(875 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC81841 273 2e-73
AW257187 similar to GP|15810133|gb| unknown protein {Arabidopsis... 240 2e-63
BF640289 88 3e-36
CA918258 weakly similar to PIR|T02457|T024 hypothetical protein ... 119 5e-27
TC92650 homologue to GP|2252842|gb|AAB62841.1| A_IG005I10.23 gen... 97 3e-20
TC93185 similar to GP|2252842|gb|AAB62841.1| A_IG005I10.23 gene ... 92 1e-18
>TC81841
Length = 422
Score = 273 bits (699), Expect = 2e-73
Identities = 140/140 (100%), Positives = 140/140 (100%)
Frame = -3
Query: 208 PTKPETEACCVRKTFKRCDS*ICESDDIERQRSGGS*KVSLL**NNGSTSAYKFR*GIVS 267
PTKPETEACCVRKTFKRCDS*ICESDDIERQRSGGS*KVSLL**NNGSTSAYKFR*GIVS
Sbjct: 420 PTKPETEACCVRKTFKRCDS*ICESDDIERQRSGGS*KVSLL**NNGSTSAYKFR*GIVS 241
Query: 268 CISTKSKSTCIEMCSRV*EFSSSERQRFQLLGTRSWK*RTNQ*DC*PQEA*LFQEKGEVS 327
CISTKSKSTCIEMCSRV*EFSSSERQRFQLLGTRSWK*RTNQ*DC*PQEA*LFQEKGEVS
Sbjct: 240 CISTKSKSTCIEMCSRV*EFSSSERQRFQLLGTRSWK*RTNQ*DC*PQEA*LFQEKGEVS 61
Query: 328 IEKFNK*KCKYQHFK*NCHS 347
IEKFNK*KCKYQHFK*NCHS
Sbjct: 60 IEKFNK*KCKYQHFK*NCHS 1
>AW257187 similar to GP|15810133|gb| unknown protein {Arabidopsis thaliana},
partial (1%)
Length = 536
Score = 240 bits (612), Expect = 2e-63
Identities = 133/178 (74%), Positives = 140/178 (77%), Gaps = 9/178 (5%)
Frame = +3
Query: 535 YQ*ITRN*VRLKSLRRSNSR**RRKLLSC*R*DSY*RGCRICKRDCYCRIFLTISWPQCR 594
YQ ITRN*VRLKSLR SNSR**RRKL SC*R*DSY RGC ICKRDCYCRIFLTISWPQCR
Sbjct: 3 YQSITRN*VRLKSLRLSNSR**RRKLWSC*R*DSYCRGCIICKRDCYCRIFLTISWPQCR 182
Query: 595 KC*PKP*YLRKF*LCRIFSMFGAG---------GISVCTTILHKLMIYVRVKYNFSPYKI 645
KC*PKP*YL+KF*LCRIFSMFGAG +S+ H+ + + +
Sbjct: 183 KC*PKP*YLKKF*LCRIFSMFGAGCNRTKPITISLSLFAHWEHQGARNYHICIRAAKSGV 362
Query: 646 TISQTILRG*RLLWKLQMSTC*STSTTTACSIRRTRFFSCESNQKRKILYRRKRIDI* 703
TILRG*RLLWKLQMSTC*STSTTTACSIRRTRFFSCES QKRKILYRRKRIDI*
Sbjct: 363 CSGCTILRG*RLLWKLQMSTC*STSTTTACSIRRTRFFSCESXQKRKILYRRKRIDI* 536
>BF640289
Length = 337
Score = 87.8 bits (216), Expect(3) = 3e-36
Identities = 40/47 (85%), Positives = 41/47 (87%)
Frame = +3
Query: 5 SQRRAVRYEKDKAGCMWGFISMLDFRHGHSTRKLIADKRRSSKHSEG 51
S+ R YEKDKAGCMWGFISMLDFRHGHSTRKLIAD RR SKHSEG
Sbjct: 75 SKTRXCXYEKDKAGCMWGFISMLDFRHGHSTRKLIADXRRXSKHSEG 215
Score = 83.2 bits (204), Expect(3) = 3e-36
Identities = 41/42 (97%), Positives = 42/42 (99%)
Frame = +2
Query: 63 KCCAF*E*V*GVERYG*SLSWHF*RWRE*KTNSENYC**A*C 104
+CCAF*E*V*GVERYG*SLSWHF*RWRE*KTNSENYC**A*C
Sbjct: 212 RCCAF*E*V*GVERYG*SLSWHF*RWRE*KTNSENYC**A*C 337
Score = 20.8 bits (42), Expect(3) = 3e-36
Identities = 9/9 (100%), Positives = 9/9 (100%)
Frame = +1
Query: 1 MAKKSQRRA 9
MAKKSQRRA
Sbjct: 61 MAKKSQRRA 87
>CA918258 weakly similar to PIR|T02457|T024 hypothetical protein At2g45900
[imported] - Arabidopsis thaliana, partial (10%)
Length = 557
Score = 119 bits (298), Expect = 5e-27
Identities = 65/132 (49%), Positives = 85/132 (64%)
Frame = +3
Query: 687 SNQKRKILYRRKRIDI*LHQCSSASLWIDKRSIVDEMPFLRQDARPFLV*SGRVITKPAM 746
S+ ++L+ K IDI*+H+C W+D RS +DEM F+ QD R F+V*SGRV+ K A+
Sbjct: 114 SDSIEEMLF*TK*IDI*VHKCCHPHRWLDTRSTIDEMSFVGQDTRSFVV*SGRVLLKHAL 293
Query: 747 P*SETPIRMYQ*NSHRSLLRLLRRLTFCIICKS*HKANSKYENGYSHGLGRSVLVFTSHA 806
P*+E PIR+YQ SH LL LL L IICK *+KANSKYE GYS GLG+ + +
Sbjct: 294 P*TEAPIRLYQRGSHGGLLALLWSLALGIICKP*YKANSKYEKGYSQGLGKECVGMSFLC 473
Query: 807 PAAHVGKNC*ER 818
P + K E+
Sbjct: 474 PHRTLWKQLSEK 509
Score = 46.2 bits (108), Expect = 5e-05
Identities = 20/33 (60%), Positives = 23/33 (69%)
Frame = +1
Query: 797 RSVLVFTSHAPAAHVGKNC*ERHGKMWIMDGPS 829
RSVL S AP AH G NC +RHG+ W +DGPS
Sbjct: 445 RSVLACPSFAPTAHFGNNCQKRHGEKWNVDGPS 543
>TC92650 homologue to GP|2252842|gb|AAB62841.1| A_IG005I10.23 gene product
{Arabidopsis thaliana}, partial (2%)
Length = 760
Score = 97.1 bits (240), Expect = 3e-20
Identities = 82/177 (46%), Positives = 94/177 (52%), Gaps = 16/177 (9%)
Frame = +1
Query: 369 FS*YCRVQFSF-CER*FTFFSYRNKKEIKACYRKGETW---------------KSHW*RQ 412
FS*YC +Q SF CE FTFF YR+KKEI CYRKGE W K+HW*R
Sbjct: 52 FS*YCPLQRSFFCESWFTFFPYRDKKEI*TCYRKGEAWKPRTKC*T*KQRLKRKNHW*R* 231
Query: 413 CRNEIS**RSFLY*KNRKAFSRCY*RRQDWY**RFGSDCGA*KGYLSETRCF*FIY*G*E 472
NEIS**R FL+*KN K + C RRQD Y R C *K + + F I *G E
Sbjct: 232 I*NEIS**RPFLH*KNCKTYV*CCKRRQDCYSERLQI*CTT*KRFY-*GKGFKHIR*GKE 408
Query: 473 TSERDGR**R*EHNGLVK*AKFQNPRKNNIVS*LQLFST**S*KGLGGSFCDRKNKT 529
T D R* R* H + NP +N S*+QL S+ S + G S CD KT
Sbjct: 409 TFI*DAR*WR**HRHF*Q-PDS*NPWQNTSSS*VQLLSSWESWREFGASSCDCTFKT 576
>TC93185 similar to GP|2252842|gb|AAB62841.1| A_IG005I10.23 gene product
{Arabidopsis thaliana}, partial (2%)
Length = 696
Score = 91.7 bits (226), Expect = 1e-18
Identities = 74/170 (43%), Positives = 95/170 (55%)
Frame = +2
Query: 127 KA*SLPEVRFQKEKEIFREKTQYCR*FKFGCSIEI*NFASSALKEAI*R*S*CGYNN*RV 186
+ * PE RF K+K+I +EK +Y C EI SALK I*R* * G +N R
Sbjct: 95 RV*RFPEDRF*KKKKISQEKPRYGHQ*S-ECDFEIRILT*SALKPTI*R*Y*FGQDNGRF 271
Query: 187 L*SKRCLFHDA*R*WRSRKAWPTKPETEACCVRKTFKRCDS*ICESDDIERQRSGGS*KV 246
L + CL DA*R* + +++ C + KRC+S* CES DIE +R G *K+
Sbjct: 272 LSN*TCLLLDA*R***QKS*---SVKSKECQF*RACKRCNS*FCESKDIEWERYGRG*KI 442
Query: 247 SLL**NNGSTSAYKFR*GIVSCISTKSKSTCIEMCSRV*EFSSSERQRFQ 296
L *+NG+TS YK *GIVS ST+SK T E+ S + E S +R Q
Sbjct: 443 PLFE*SNGNTSGYKLG*GIVSKTSTRSKFTFTEVYSGIGECSREN*KRIQ 592
Score = 38.5 bits (88), Expect = 0.011
Identities = 25/52 (48%), Positives = 29/52 (55%)
Frame = +1
Query: 97 NYC**A*CEETHRRRDVH*SKCNEGYR*F*KA*SLPEVRFQKEKEIFREKTQ 148
N C* A*CEETHRRRDVH* N+ R +P + KEKE K +
Sbjct: 4 N*C*QA*CEETHRRRDVH*PG*NKESRYLI*GLKIP*RQILKEKENLARKAE 159
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.374 0.166 0.667
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,071,222
Number of Sequences: 36976
Number of extensions: 477903
Number of successful extensions: 7291
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1204
Number of HSP's successfully gapped in prelim test: 206
Number of HSP's that attempted gapping in prelim test: 6007
Number of HSP's gapped (non-prelim): 1594
length of query: 875
length of database: 9,014,727
effective HSP length: 105
effective length of query: 770
effective length of database: 5,132,247
effective search space: 3951830190
effective search space used: 3951830190
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.5 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146341.2


