

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.8 + phase: 0
(402 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
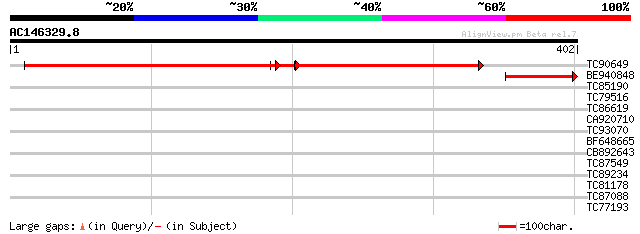
Sequences producing significant alignments: (bits) Value
TC90649 similar to GP|20197028|gb|AAC27454.3 putative heme A:far... 353 4e-99
BE940848 similar to GP|21328147|dbj putative heme A farnesyltran... 104 7e-23
TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 32 0.33
TC79516 similar to GP|8777406|dbj|BAA96996.1 gb|AAF13095.1~gene_... 32 0.57
TC86619 similar to GP|16648824|gb|AAL25602.1 AT4g18950/F13C5_120... 32 0.57
CA920710 similar to PIR|S64314|S643 probable membrane protein YG... 30 1.3
TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown prot... 30 1.7
BF648665 weakly similar to GP|15294296|gb AT3g56820/T8M16_150 {A... 29 3.7
CB892643 similar to PIR|B96582|B96 hypothetical protein F15I1.21... 28 4.8
TC87549 similar to PIR|T01734|T01734 hypothetical protein A_IG00... 28 6.3
TC89234 similar to GP|22945532|gb|AAF51469.2 CG2839-PA {Drosophi... 28 6.3
TC81178 28 6.3
TC87088 similar to PIR|F86399|F86399 protein F17L21.22 [imported... 28 8.2
TC77193 homologue to GP|21593495|gb|AAM65462.1 glycine rich prot... 28 8.2
>TC90649 similar to GP|20197028|gb|AAC27454.3 putative heme
A:farnesyltransferase {Arabidopsis thaliana}, partial
(64%)
Length = 1209
Score = 353 bits (905), Expect(2) = 4e-99
Identities = 182/182 (100%), Positives = 182/182 (100%)
Frame = +2
Query: 11 LYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSLL 70
LYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSLL
Sbjct: 89 LYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSLL 268
Query: 71 VVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLP 130
VVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLP
Sbjct: 269 VVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLP 448
Query: 131 SGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWV 190
SGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWV
Sbjct: 449 SGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWV 628
Query: 191 GA 192
GA
Sbjct: 629 GA 634
Score = 265 bits (677), Expect = 2e-71
Identities = 129/134 (96%), Positives = 129/134 (96%)
Frame = +1
Query: 203 WAAASGDISLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSLADPSGRKTAMVA 262
WAAASGDISLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSLADPSGRKTAMVA
Sbjct: 805 WAAASGDISLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSLADPSGRKTAMVA 984
Query: 263 MRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAAAFSFYRDRTKERARRMFHASLL 322
MRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAA FSFY DRTKERAR MFHA LL
Sbjct: 985 MRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAAXFSFYXDRTKERARXMFHAXLL 1164
Query: 323 YLPVFMAGLLIHRR 336
YLPVFMAGL IHRR
Sbjct: 1165YLPVFMAGLXIHRR 1206
Score = 29.6 bits (65), Expect = 2.2
Identities = 14/31 (45%), Positives = 20/31 (64%), Gaps = 3/31 (9%)
Frame = +1
Query: 1 MFKRNWNCATLY---SRLVNSSSSSSLHNFT 28
MFKRNWNCAT+ + + SS LH+++
Sbjct: 58 MFKRNWNCATVIFQTRQFLLFFVSSQLHSYS 150
Score = 26.9 bits (58), Expect(2) = 4e-99
Identities = 13/21 (61%), Positives = 15/21 (70%)
Frame = +3
Query: 186 VNTWVGAVVGAIPPLLGWAAA 206
++ W G VVGAIPPLLG A
Sbjct: 618 IHGW-GXVVGAIPPLLGTPVA 677
>BE940848 similar to GP|21328147|dbj putative heme A farnesyltransferase
homolog {Oryza sativa (japonica cultivar-group)},
partial (4%)
Length = 365
Score = 104 bits (259), Expect = 7e-23
Identities = 50/51 (98%), Positives = 50/51 (98%)
Frame = +1
Query: 352 KSPSSVETSEMDDKNGNQKIKGRQGTRARPPVAYASIAPFPFLPAPSYDIV 402
KSPSSVETSEMDD NGNQKIKGRQGTRARPPVAYASIAPFPFLPAPSYDIV
Sbjct: 1 KSPSSVETSEMDDMNGNQKIKGRQGTRARPPVAYASIAPFPFLPAPSYDIV 153
>TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (50%)
Length = 1496
Score = 32.3 bits (72), Expect = 0.33
Identities = 44/168 (26%), Positives = 70/168 (41%), Gaps = 8/168 (4%)
Frame = -2
Query: 9 ATLYSRLVNSSSSSSLHNFTHIRSTFSSSPS-VSASKSNDLISLARHYGNCYSQLSKARL 67
+TL S V SSS+ + T S FSS+ S V S S+ S
Sbjct: 685 STLGSSEVEPFSSSTFFSSTTASSFFSSTTSSVFFSSSSGFFSS---------------- 554
Query: 68 SLLVVATSGTGFVLGS-GGAVDLSMLS---YTCLGTMMV---AASASTLNQVFEIKNDAI 120
ATSG+G + G+ GGAV+ +LS +CL + + +S V +K
Sbjct: 553 ---FTATSGSGALTGAGGGAVNSCLLSAS*ISCLSSSSAFF*TSGSSETTLVSSLKETD* 383
Query: 121 MRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAAS 168
S + S ++ + G+++S AG +++AA AA+
Sbjct: 382 ETLLSSTTVSSATLSATTSAGFSASSATAGGGSATASLSVVAAVTAAA 239
>TC79516 similar to GP|8777406|dbj|BAA96996.1
gb|AAF13095.1~gene_id:MIF21.5~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (29%)
Length = 1077
Score = 31.6 bits (70), Expect = 0.57
Identities = 32/110 (29%), Positives = 53/110 (48%), Gaps = 4/110 (3%)
Frame = -3
Query: 5 NWNCATLYSRLVNSSSSSSLHNFTHIRSTFSSSPSV--SASKSNDLISLARHYGNCYSQL 62
NWN + + SSSS+ +F S SSS S+ SAS + R + N
Sbjct: 442 NWNISASAFLSLTILSSSSICDFFFCLSILSSSASLALSASSRSQASRAKRAFLNILI-F 266
Query: 63 SKARLSLLVVATSGTGFVL--GSGGAVDLSMLSYTCLGTMMVAASASTLN 110
S A S+ ++A+ T +L S GA++L + +++ L ++S T+N
Sbjct: 265 SSATNSIFLIASCTTSAILLHASCGAINLPLSTFSRLFGESRSSS*KTVN 116
>TC86619 similar to GP|16648824|gb|AAL25602.1 AT4g18950/F13C5_120
{Arabidopsis thaliana}, partial (77%)
Length = 1505
Score = 31.6 bits (70), Expect = 0.57
Identities = 26/78 (33%), Positives = 36/78 (45%)
Frame = -2
Query: 7 NCATLYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKAR 66
NC T R+ + +SSSL FT S +SSPS + S+ RH Q
Sbjct: 403 NCTTFGCRIF*NRASSSLKAFTFSSSLLTSSPSFFTA-----TSVPRHNAR---QKVPFV 248
Query: 67 LSLLVVATSGTGFVLGSG 84
+S L+V +S G + SG
Sbjct: 247 ISTLLVKSSSLGLISYSG 194
>CA920710 similar to PIR|S64314|S643 probable membrane protein YGR023w -
yeast (Saccharomyces cerevisiae), partial (7%)
Length = 774
Score = 30.4 bits (67), Expect = 1.3
Identities = 25/62 (40%), Positives = 34/62 (54%), Gaps = 3/62 (4%)
Frame = +1
Query: 17 NSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKAR---LSLLVVA 73
+SSSSSS +F+ S+FSSS S SAS S + + G S S A +SL ++A
Sbjct: 292 SSSSSSSSSSFSSSSSSFSSSSSSSASASASIYPDFQGVGTKLST*SMAENQ*ISLEILA 471
Query: 74 TS 75
S
Sbjct: 472 LS 477
>TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown protein
{Arabidopsis thaliana}, partial (19%)
Length = 679
Score = 30.0 bits (66), Expect = 1.7
Identities = 19/40 (47%), Positives = 25/40 (62%)
Frame = -3
Query: 17 NSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYG 56
+SSSSSS + + S+ SSS S S+SKS IS + YG
Sbjct: 332 SSSSSSSSSSSSSSSSSSSSSSSSSSSKSISSISSSESYG 213
>BF648665 weakly similar to GP|15294296|gb AT3g56820/T8M16_150 {Arabidopsis
thaliana}, partial (48%)
Length = 647
Score = 28.9 bits (63), Expect = 3.7
Identities = 24/66 (36%), Positives = 34/66 (51%), Gaps = 2/66 (3%)
Frame = +2
Query: 18 SSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSL--LVVATS 75
S+ SSLH+F + + SSSPS S+S L SL+ H LS + + L+ A S
Sbjct: 140 SAPPSSLHSFKDLTLSLSSSPSSSSS----LGSLSAHIALLIHSLSLPQTTTLKLIFADS 307
Query: 76 GTGFVL 81
F+L
Sbjct: 308 LPAFLL 325
>CB892643 similar to PIR|B96582|B96 hypothetical protein F15I1.21 [imported]
- Arabidopsis thaliana, partial (35%)
Length = 758
Score = 28.5 bits (62), Expect = 4.8
Identities = 13/25 (52%), Positives = 14/25 (56%)
Frame = -3
Query: 117 NDAIMRRTSQRPLPSGRITVPHAVG 141
NDAI+ TS P P G T P VG
Sbjct: 330 NDAILANTSGAPFPRGSNTTPATVG 256
>TC87549 similar to PIR|T01734|T01734 hypothetical protein A_IG002N01.30 -
Arabidopsis thaliana (fragment), partial (38%)
Length = 1766
Score = 28.1 bits (61), Expect = 6.3
Identities = 18/54 (33%), Positives = 28/54 (51%)
Frame = -3
Query: 7 NCATLYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYS 60
NC L++ V++SSSS + S+ SS S+S S+ LI + +C S
Sbjct: 1713 NCLFLFNSDVSASSSSKSGGESSSSSSIQSSSISSSSSSSSLIKSSASDHSCSS 1552
>TC89234 similar to GP|22945532|gb|AAF51469.2 CG2839-PA {Drosophila
melanogaster}, partial (2%)
Length = 717
Score = 28.1 bits (61), Expect = 6.3
Identities = 17/48 (35%), Positives = 26/48 (53%)
Frame = +1
Query: 18 SSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKA 65
SSSS LH F+ I S F+ S V ++ L ++ CYSQ++ +
Sbjct: 70 SSSSPILHQFSSILSPFAFSLRVGFG----IVFLVCYFDLCYSQVASS 201
>TC81178
Length = 476
Score = 28.1 bits (61), Expect = 6.3
Identities = 17/38 (44%), Positives = 21/38 (54%), Gaps = 2/38 (5%)
Frame = -2
Query: 12 YSRLVNSSSSSSLHNFTHIRSTFSSSPS--VSASKSND 47
Y R S +++ F HIR+T SPS VS S SND
Sbjct: 214 YIRNAGSLKTTTASRFLHIRTTVVLSPSSMVSESASND 101
>TC87088 similar to PIR|F86399|F86399 protein F17L21.22 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 2997
Score = 27.7 bits (60), Expect = 8.2
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = +1
Query: 77 TGFVLGSGGAVDLSMLSYTCLGTMMVAASAST 108
TGF+L G + LS+ + TCLG + S+ T
Sbjct: 2290 TGFLLQQKGLIMLSISALTCLGFCLPQTSSQT 2385
>TC77193 homologue to GP|21593495|gb|AAM65462.1 glycine rich protein-like
{Arabidopsis thaliana}, partial (85%)
Length = 989
Score = 27.7 bits (60), Expect = 8.2
Identities = 18/59 (30%), Positives = 31/59 (52%), Gaps = 1/59 (1%)
Frame = -2
Query: 24 LHNFTHIRSTFSSSPSVSA-SKSNDLISLARHYGNCYSQLSKARLSLLVVATSGTGFVL 81
+H H+ + S+SPS SA S+ + L S A G+ + + K S +A++ T +L
Sbjct: 586 VHTSYHLHRSTSTSPSTSASSRLSTLASPASASGSSSTTIIKTTPSAFCLASTSTSSIL 410
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,936,543
Number of Sequences: 36976
Number of extensions: 186272
Number of successful extensions: 1395
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1270
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1343
length of query: 402
length of database: 9,014,727
effective HSP length: 98
effective length of query: 304
effective length of database: 5,391,079
effective search space: 1638888016
effective search space used: 1638888016
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146329.8


