

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.5 + phase: 0
(639 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
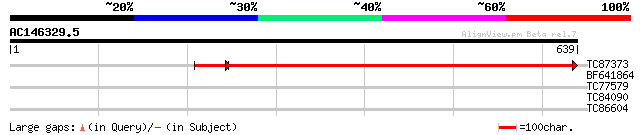
Sequences producing significant alignments: (bits) Value
TC87373 homologue to PIR|A43350|A43350 formate--tetrahydrofolate... 766 0.0
BF641864 30 2.9
TC77579 similar to SP|Q43314|DHE1_ARATH Glutamate dehydrogenase ... 29 4.9
TC84090 similar to PIR|D84555|D84555 probable protein kinase [im... 29 4.9
TC86604 similar to SP|Q9S7A0|DHE3_ARATH Probable glutamate dehyd... 28 8.3
>TC87373 homologue to PIR|A43350|A43350 formate--tetrahydrofolate ligase (EC
6.3.4.3) - spinach, partial (67%)
Length = 1591
Score = 766 bits (1978), Expect(2) = 0.0
Identities = 388/393 (98%), Positives = 389/393 (98%)
Frame = +3
Query: 247 KITVGQGPDEKGMVRETAFDISVASEIMAVLALTTSLTDMRERLGKMVIGNSKSGDPVTA 306
K+ VG GPDEKGMV ETAFDISVASEIMAVLALTTSLTDMRERLGKM IGNSKSGDPVTA
Sbjct: 117 KLLVGHGPDEKGMVPETAFDISVASEIMAVLALTTSLTDMRERLGKMGIGNSKSGDPVTA 296
Query: 307 DDLGIGGALTVLMKDAIHPTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVGPG 366
DDLGIGGALTVLMKDAIHPTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVGPG
Sbjct: 297 DDLGIGGALTVLMKDAIHPTLMQTLEGTPVLVHAGPFANIAHGNSSIVADKIALKLVGPG 476
Query: 367 GFVVTEAGFGADIGTEKFMNIKCRYSGLTPQCAIIVATIRALKMHGGGPAVVAGKPLDHA 426
GFVVTEAGFGADIGTEKFMNIKCRYSGLTPQCAIIVATIRALKMHGGGPAVVAGKPLDHA
Sbjct: 477 GFVVTEAGFGADIGTEKFMNIKCRYSGLTPQCAIIVATIRALKMHGGGPAVVAGKPLDHA 656
Query: 427 YLTENVALVEAGCANLARHILNSKAYGANVIVAINKFSTDTDAELNAVKKAALDAGAYDA 486
YLTENVALVEAGCANLARHILNSKAYGANVIVAINKFSTDTDAELNAVKKAALDAGAYDA
Sbjct: 657 YLTENVALVEAGCANLARHILNSKAYGANVIVAINKFSTDTDAELNAVKKAALDAGAYDA 836
Query: 487 VICSHHAHGGRGAVDLGIAVQKACENATQPLKFLYPLDIGIKEKIEAIAKSYGASGVEYS 546
VICSHHAHGGRGAVDLGIAVQKACENATQPLKFLYPLDIGIKEKIEAIAKSYGASGVEYS
Sbjct: 837 VICSHHAHGGRGAVDLGIAVQKACENATQPLKFLYPLDIGIKEKIEAIAKSYGASGVEYS 1016
Query: 547 EQAEKKIELYTKQGFSGLPICMAKTQYSFSDNAAAKGAPTGFILPIRDVRASIGAGFIYP 606
EQAEKKIELYTKQGFSGLPICMAKTQYSFSDNAAAKGAPTGFILPIRDVRASIGAGFIYP
Sbjct: 1017EQAEKKIELYTKQGFSGLPICMAKTQYSFSDNAAAKGAPTGFILPIRDVRASIGAGFIYP 1196
Query: 607 LVGTMSTMPGLPTRPCFYDIDLDTTTGQVIGLS 639
LVGTMSTMPGLPTRPCFYDIDLDTTTGQVIGLS
Sbjct: 1197LVGTMSTMPGLPTRPCFYDIDLDTTTGQVIGLS 1295
Score = 88.6 bits (218), Expect(2) = 0.0
Identities = 41/41 (100%), Positives = 41/41 (100%)
Frame = +2
Query: 209 TNPDDLTPEEVNKFARLDIDPDSITWRRVMDINDRFLRKIT 249
TNPDDLTPEEVNKFARLDIDPDSITWRRVMDINDRFLRKIT
Sbjct: 2 TNPDDLTPEEVNKFARLDIDPDSITWRRVMDINDRFLRKIT 124
>BF641864
Length = 656
Score = 30.0 bits (66), Expect = 2.9
Identities = 19/55 (34%), Positives = 30/55 (54%), Gaps = 4/55 (7%)
Frame = +1
Query: 57 KVLLSALDELQESKDGYYVVVGGI---TPTPLGEGKSTTTVGL-CQALGAFLDKK 107
K + LD++ ESK+ + + +GG P + EG+ T + L +ALG DKK
Sbjct: 421 KEITRRLDDIAESKNKFSLQMGGTLREIPXQVAEGRQTXSTPLESKALGRDXDKK 585
>TC77579 similar to SP|Q43314|DHE1_ARATH Glutamate dehydrogenase 1 (EC
1.4.1.3) (GDH 1). [Mouse-ear cress] {Arabidopsis
thaliana}, complete
Length = 1530
Score = 29.3 bits (64), Expect = 4.9
Identities = 18/48 (37%), Positives = 22/48 (45%)
Frame = +2
Query: 409 KMHGGGPAVVAGKPLDHAYLTENVALVEAGCANLARHILNSKAYGANV 456
K HG PAVV GKP+D A G A +LN YG ++
Sbjct: 584 KFHGYSPAVVTGKPIDLGGSLGRDAATGRGVMFAAEALLNE--YGKSI 721
>TC84090 similar to PIR|D84555|D84555 probable protein kinase [imported] -
Arabidopsis thaliana, partial (19%)
Length = 794
Score = 29.3 bits (64), Expect = 4.9
Identities = 12/39 (30%), Positives = 23/39 (58%)
Frame = -1
Query: 215 TPEEVNKFARLDIDPDSITWRRVMDINDRFLRKITVGQG 253
TP + N F+ ++I SITWR+ +++ + K+ + G
Sbjct: 305 TPMK*NIFSSINIHIKSITWRQTIEVRIKLFTKLRIRYG 189
>TC86604 similar to SP|Q9S7A0|DHE3_ARATH Probable glutamate dehydrogenase 3
(EC 1.4.1.3) (GDH 3). [Mouse-ear cress] {Arabidopsis
thaliana}, complete
Length = 1566
Score = 28.5 bits (62), Expect = 8.3
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = +1
Query: 409 KMHGGGPAVVAGKPLD 424
K HG PAVV GKP+D
Sbjct: 598 KFHGHSPAVVTGKPID 645
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,392,952
Number of Sequences: 36976
Number of extensions: 203342
Number of successful extensions: 931
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 928
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 931
length of query: 639
length of database: 9,014,727
effective HSP length: 102
effective length of query: 537
effective length of database: 5,243,175
effective search space: 2815584975
effective search space used: 2815584975
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146329.5


