

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
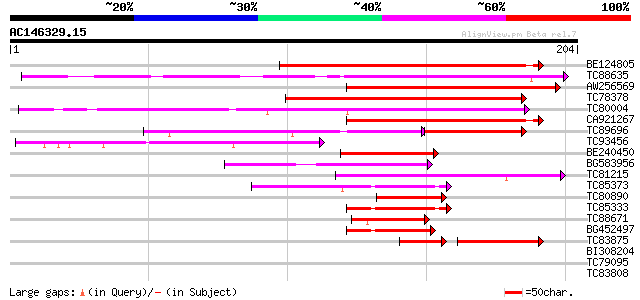
Query= AC146329.15 - phase: 0
(204 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE124805 similar to GP|10177426|db GATA-binding transcription fa... 127 4e-30
TC88635 similar to GP|12711287|emb|CAC28528. GATA-1 zinc finger ... 121 2e-28
AW256569 similar to GP|10177426|dbj GATA-binding transcription f... 119 6e-28
TC78378 similar to GP|10177426|dbj|BAB10711. GATA-binding transc... 118 2e-27
TC80004 similar to GP|8778844|gb|AAF79843.1| T6D22.9 {Arabidopsi... 117 2e-27
CA921267 homologue to GP|10177426|dbj GATA-binding transcription... 114 2e-26
TC89696 similar to GP|10177426|dbj|BAB10711. GATA-binding transc... 61 3e-22
TC93456 similar to GP|12711287|emb|CAC28528. GATA-1 zinc finger ... 76 8e-15
BE240450 similar to GP|21555178|gb| transcription factor-like pr... 61 3e-10
BG583956 homologue to GP|10177159|dbj gb|AAF63824.1~gene_id:K21P... 54 3e-08
TC81215 similar to GP|17064972|gb|AAL32640.1 Unknown protein {Ar... 54 5e-08
TC85373 similar to GP|15028099|gb|AAK76580.1 putative flowering ... 47 5e-06
TC80890 similar to PIR|T04270|T04270 hypothetical protein F20B18... 45 2e-05
TC85333 similar to GP|15028099|gb|AAK76580.1 putative flowering ... 44 6e-05
TC88671 similar to PIR|JC7336|JC7336 zinc-finger protein - Arabi... 44 6e-05
BG452497 similar to GP|15028099|gb| putative flowering protein C... 43 9e-05
TC83875 similar to GP|17380994|gb|AAL36309.1 putative GATA trans... 39 3e-04
BI308204 similar to PIR|JC7336|JC7 zinc-finger protein - Arabido... 30 0.64
TC79095 similar to GP|22655232|gb|AAM98206.1 unknown protein {Ar... 30 0.83
TC83808 similar to PIR|T04231|T04231 hypothetical protein F14M19... 29 1.1
>BE124805 similar to GP|10177426|db GATA-binding transcription factor-like
protein {Arabidopsis thaliana}, partial (36%)
Length = 715
Score = 127 bits (318), Expect = 4e-30
Identities = 59/95 (62%), Positives = 71/95 (74%)
Frame = +2
Query: 98 ESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGR 157
E K + KA + + D A G+ R+C+HC KTPQWRTGP GPKTLCNACGVRYKSGR
Sbjct: 350 ECSKPAEKKAKRMVSPDGEARGVPRRCSHCGVQKTPQWRTGPGGPKTLCNACGVRYKSGR 529
Query: 158 LCPEYRPAASSTFSPDLHSNSHKKILEMRVMRRKD 192
L PEYRPA S TFS +LHSN H+K++EMR R+K+
Sbjct: 530 LLPEYRPACSPTFSSELHSNHHRKVIEMR--RKKE 628
>TC88635 similar to GP|12711287|emb|CAC28528. GATA-1 zinc finger protein
{Nicotiana tabacum}, partial (21%)
Length = 1327
Score = 121 bits (303), Expect = 2e-28
Identities = 79/198 (39%), Positives = 104/198 (51%), Gaps = 1/198 (0%)
Frame = +1
Query: 5 EQHPSSSVNKEDFVLPKSNSSPTCEKTTVRRTRSKRPRLATFSSHHSTMQLISSTSSFVG 64
E SSV +F LP +R R KR RL++F S I +F
Sbjct: 547 ESSSYSSVESYNFELPV---------IPAKRPRGKRRRLSSFDRLFS----IPFIPAFQK 687
Query: 65 ENMQDSVISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQC 124
+ +K P +K + + + +TK L H ++ R+C
Sbjct: 688 QQSDKQRAGKSLCRAQKRP--------RKDDTSQLSDSIETKRSSL----HESIA-PRKC 828
Query: 125 THCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILE 184
THCE T+TPQWR GP+GPKTLCNACGVRY+SGRL PEYRPAAS TF +HSNSHKK+LE
Sbjct: 829 THCEVTETPQWREGPKGPKTLCNACGVRYRSGRLFPEYRPAASPTFEASVHSNSHKKVLE 1008
Query: 185 MR-VMRRKDNKNSGILAL 201
+R + ++ K S +L L
Sbjct: 1009IRKKVNQETVKGSSMLYL 1062
>AW256569 similar to GP|10177426|dbj GATA-binding transcription factor-like
protein {Arabidopsis thaliana}, partial (20%)
Length = 613
Score = 119 bits (299), Expect = 6e-28
Identities = 52/77 (67%), Positives = 61/77 (78%)
Frame = -3
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C+HC KTPQWR GP GPKTLCNACGVR+KSGRL PEYRPA S TFS ++HSNSH+K
Sbjct: 509 RRCSHCHVQKTPQWRAGPLGPKTLCNACGVRFKSGRLFPEYRPACSPTFSGEIHSNSHRK 330
Query: 182 ILEMRVMRRKDNKNSGI 198
+LEMR + D SG+
Sbjct: 329 VLEMRRRKEVDEPVSGL 279
>TC78378 similar to GP|10177426|dbj|BAB10711. GATA-binding transcription
factor-like protein {Arabidopsis thaliana}, partial
(23%)
Length = 1284
Score = 118 bits (295), Expect = 2e-27
Identities = 54/87 (62%), Positives = 65/87 (74%)
Frame = +3
Query: 100 KKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLC 159
K+ KK +A ++ L R+C+HC+ KTPQWRTGP G KTLCNACGVRYKSGRL
Sbjct: 747 KQKKKPEAQVVGHEAQEEGQLQRRCSHCQVQKTPQWRTGPMGAKTLCNACGVRYKSGRLF 926
Query: 160 PEYRPAASSTFSPDLHSNSHKKILEMR 186
EYRPA S TFS ++HSNSH+K+LEMR
Sbjct: 927 SEYRPACSPTFSSEIHSNSHRKVLEMR 1007
>TC80004 similar to GP|8778844|gb|AAF79843.1| T6D22.9 {Arabidopsis
thaliana}, partial (8%)
Length = 1058
Score = 117 bits (294), Expect = 2e-27
Identities = 74/186 (39%), Positives = 104/186 (55%), Gaps = 2/186 (1%)
Frame = +3
Query: 4 VEQHPSSSVNKEDFVLPKSNSSPTCEKTTVRRTRSKRPRLATFSSHHSTMQLISSTSSFV 63
V PS++V K + N SP + RSKR RL+ + ++S+T +
Sbjct: 327 VSSKPSNTVKKPK---QEQNESPFA---VPGKARSKRKRLSAPRRPKDPLSILSNTLNPQ 488
Query: 64 GENMQDSVISNKGASTEKFPDSQIAAKK-QKLSSGESKKNKKTKAPLLAALDHNALGL-V 121
E++ K A DS++ K +K + + + +K + +++ +
Sbjct: 489 NESLCSDPPLLKQAYW--LADSELMVPKGEKEVTKDCEVVEKERFDFEGFVNNGQNPIPT 662
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+CTHC + +TPQWR GP GPKTLCNACGVRYKSGRL PEYRPA S TF LHSNSHKK
Sbjct: 663 RRCTHCLSQRTPQWRAGPLGPKTLCNACGVRYKSGRLLPEYRPAKSPTFVSFLHSNSHKK 842
Query: 182 ILEMRV 187
++EMR+
Sbjct: 843 VMEMRM 860
>CA921267 homologue to GP|10177426|dbj GATA-binding transcription factor-like
protein {Arabidopsis thaliana}, partial (20%)
Length = 694
Score = 114 bits (286), Expect = 2e-26
Identities = 51/71 (71%), Positives = 60/71 (83%)
Frame = -1
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C+HC TKTPQWR+GP G KTLCNACGVR+KSGRL PEYRPA S TFS +LHSN H+K
Sbjct: 538 RRCSHCGVTKTPQWRSGPLGAKTLCNACGVRFKSGRLLPEYRPACSPTFSSELHSNHHRK 359
Query: 182 ILEMRVMRRKD 192
+LEMR R+K+
Sbjct: 358 VLEMR--RKKE 332
>TC89696 similar to GP|10177426|dbj|BAB10711. GATA-binding transcription
factor-like protein {Arabidopsis thaliana}, partial
(25%)
Length = 806
Score = 61.2 bits (147), Expect(2) = 3e-22
Identities = 39/111 (35%), Positives = 50/111 (44%), Gaps = 9/111 (8%)
Frame = +2
Query: 49 HHSTMQLI-SSTSSFVGENMQDSVISNKGASTEKFPDSQIAAKKQKLSSGESK------- 100
H T ++ +STSS N +S+ G + KFP + + K G +
Sbjct: 332 HQMTSPMLENSTSSSNSNNSSNSITLLSGYNHMKFPVRARSKSRSKPRLGLADASNLQFP 511
Query: 101 -KNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACG 150
K TK +G R+C HC TPQWR GP GPKTLCNACG
Sbjct: 512 WKQPSTKTSKEKVKQTPTIG--RKCHHCGVDDTPQWRAGPNGPKTLCNACG 658
Score = 60.1 bits (144), Expect(2) = 3e-22
Identities = 26/37 (70%), Positives = 30/37 (80%)
Frame = +3
Query: 150 GVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
GVRYKSGRL PEYRPA TF D+HSN H+K++EMR
Sbjct: 657 GVRYKSGRLVPEYRPANRPTFRSDVHSNXHRKVVEMR 767
>TC93456 similar to GP|12711287|emb|CAC28528. GATA-1 zinc finger protein
{Nicotiana tabacum}, partial (6%)
Length = 663
Score = 76.3 bits (186), Expect = 8e-15
Identities = 61/134 (45%), Positives = 76/134 (56%), Gaps = 23/134 (17%)
Frame = +3
Query: 3 KVEQHPSSS-VNKED-----FVLP------KSNSSPTCEKTT---------VRRTRSKRP 41
KVEQ PSSS V+KED F P +S+SS + KTT R R+KRP
Sbjct: 219 KVEQQPSSSAVSKEDSGHYQFQTPSPISVLESSSSCSGGKTTGIYVPIPVPCGRARTKRP 398
Query: 42 RLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGAST--EKFPDSQIAAKKQKLSSGES 99
R F+ S MQLIS TSS V ENMQ +VIS K S+ E F +S+I KK KLSSGE
Sbjct: 399 RPTAFNPR-SAMQLISPTSSSVEENMQPNVISTKAMSSDFENFAESRIIXKKPKLSSGEP 575
Query: 100 KKNKKTKAPLLAAL 113
++ +K++ L L
Sbjct: 576 RRKRKSRHHFLLLL 617
>BE240450 similar to GP|21555178|gb| transcription factor-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 529
Score = 61.2 bits (147), Expect = 3e-10
Identities = 22/35 (62%), Positives = 28/35 (79%)
Frame = +1
Query: 120 LVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
L R+C C++T TP WR GP GP++LCNACG+RYK
Sbjct: 22 LARRCASCDSTSTPLWRNGPRGPESLCNACGIRYK 126
>BG583956 homologue to GP|10177159|dbj gb|AAF63824.1~gene_id:K21P3.18~similar
to unknown protein {Arabidopsis thaliana}, partial (43%)
Length = 818
Score = 54.3 bits (129), Expect = 3e-08
Identities = 27/75 (36%), Positives = 35/75 (46%)
Frame = +2
Query: 78 STEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRT 137
S K + K+S GE AA+ + + C C +KTP WR
Sbjct: 152 SESKSESKMVVDHSDKVSEGEDSNPN-------AAVSSDNSNPKKTCADCGTSKTPLWRG 310
Query: 138 GPEGPKTLCNACGVR 152
GP GPK+LCNACG+R
Sbjct: 311 GPAGPKSLCNACGIR 355
>TC81215 similar to GP|17064972|gb|AAL32640.1 Unknown protein {Arabidopsis
thaliana}, partial (14%)
Length = 938
Score = 53.5 bits (127), Expect = 5e-08
Identities = 29/91 (31%), Positives = 41/91 (44%), Gaps = 8/91 (8%)
Frame = +3
Query: 118 LGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSN 177
+G C HC T TP WR GP LCNACG R+++ Y P + T D
Sbjct: 429 MGKQGPCFHCGVTSTPLWRNGPPEKPILCNACGSRWRTKGTLANYTPLHARTDPVDFEDQ 608
Query: 178 --------SHKKILEMRVMRRKDNKNSGILA 200
S K E+++++RK N ++ A
Sbjct: 609 RVSRLKIVSLNKNKEVKLVKRKQNYDNAYAA 701
>TC85373 similar to GP|15028099|gb|AAK76580.1 putative flowering protein
CONSTANS {Arabidopsis thaliana}, partial (22%)
Length = 683
Score = 47.0 bits (110), Expect = 5e-06
Identities = 28/74 (37%), Positives = 41/74 (54%), Gaps = 2/74 (2%)
Frame = +1
Query: 88 AAKKQKLSSGESKKNKKTKAPLLAALDHNAL--GLVRQCTHCEATKTPQWRTGPEGPKTL 145
+AK + S ++ N T L+A D + + RQC E TP R GPEGP+TL
Sbjct: 145 SAKSKNDESASAEMNCGTSEGLMADNDGSQQQDNVCRQCNISEKC-TPMMRRGPEGPRTL 321
Query: 146 CNACGVRYKSGRLC 159
CNACG+ + + ++C
Sbjct: 322 CNACGLMW-ANKIC 360
>TC80890 similar to PIR|T04270|T04270 hypothetical protein F20B18.260 -
Arabidopsis thaliana, partial (14%)
Length = 710
Score = 45.1 bits (105), Expect = 2e-05
Identities = 16/25 (64%), Positives = 20/25 (80%)
Frame = +1
Query: 133 PQWRTGPEGPKTLCNACGVRYKSGR 157
P WR+GP GPK+LCNACG+R + R
Sbjct: 7 PLWRSGPTGPKSLCNACGIRQRKAR 81
>TC85333 similar to GP|15028099|gb|AAK76580.1 putative flowering protein
CONSTANS {Arabidopsis thaliana}, partial (22%)
Length = 1421
Score = 43.5 bits (101), Expect = 6e-05
Identities = 20/38 (52%), Positives = 26/38 (67%)
Frame = +1
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLC 159
RQC E TP R GPEGP+TLCNACG+ + + ++C
Sbjct: 910 RQCNISEKC-TPMMRRGPEGPRTLCNACGLMW-ANKIC 1017
>TC88671 similar to PIR|JC7336|JC7336 zinc-finger protein - Arabidopsis
thaliana, partial (44%)
Length = 1272
Score = 43.5 bits (101), Expect = 6e-05
Identities = 17/30 (56%), Positives = 21/30 (69%), Gaps = 2/30 (6%)
Frame = +3
Query: 124 CTHC--EATKTPQWRTGPEGPKTLCNACGV 151
CTHC + TP R GP GP++LCNACG+
Sbjct: 726 CTHCGTSSKSTPMMRRGPSGPRSLCNACGL 815
>BG452497 similar to GP|15028099|gb| putative flowering protein CONSTANS
{Arabidopsis thaliana}, partial (12%)
Length = 648
Score = 42.7 bits (99), Expect = 9e-05
Identities = 19/32 (59%), Positives = 22/32 (68%)
Frame = +1
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRY 153
RQC E TP R GPEGP+TLCNACG+ +
Sbjct: 199 RQCNISEKC-TPMMRRGPEGPRTLCNACGLMW 291
>TC83875 similar to GP|17380994|gb|AAL36309.1 putative GATA transcription
factor 3 {Arabidopsis thaliana}, partial (8%)
Length = 828
Score = 38.9 bits (89), Expect(2) = 3e-04
Identities = 18/31 (58%), Positives = 22/31 (70%)
Frame = +1
Query: 162 YRPAASSTFSPDLHSNSHKKILEMRVMRRKD 192
YRPA S T +HSNSH+K+LEMR + KD
Sbjct: 67 YRPANSPTLVASVHSNSHRKVLEMRGVVIKD 159
Score = 21.6 bits (44), Expect(2) = 3e-04
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = +3
Query: 141 GPKTLCNACGVRYKSGR 157
GP + AC V Y+SGR
Sbjct: 6 GPTSQTFACCVSYRSGR 56
>BI308204 similar to PIR|JC7336|JC7 zinc-finger protein - Arabidopsis
thaliana, partial (46%)
Length = 811
Score = 30.0 bits (66), Expect = 0.64
Identities = 13/21 (61%), Positives = 15/21 (70%)
Frame = +3
Query: 138 GPEGPKTLCNACGVRYKSGRL 158
G GPKTLCNACG+ + G L
Sbjct: 729 GRLGPKTLCNACGLFWAIGVL 791
>TC79095 similar to GP|22655232|gb|AAM98206.1 unknown protein {Arabidopsis
thaliana}, partial (28%)
Length = 1592
Score = 29.6 bits (65), Expect = 0.83
Identities = 14/38 (36%), Positives = 21/38 (54%)
Frame = -3
Query: 34 RRTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSV 71
R R P+LA +S HH +Q S + F+G + Q S+
Sbjct: 1128 RTERFACPKLANWSPHHCFLQESSPSDPFLGHHWQTSI 1015
>TC83808 similar to PIR|T04231|T04231 hypothetical protein F14M19.50 -
Arabidopsis thaliana, partial (24%)
Length = 685
Score = 29.3 bits (64), Expect = 1.1
Identities = 15/51 (29%), Positives = 23/51 (44%), Gaps = 4/51 (7%)
Frame = -3
Query: 83 PDSQIAAKKQKLSSGESKKNKKTKAPLLAA----LDHNALGLVRQCTHCEA 129
P ++ A K +SG TK PL A H+ +++ C HC+A
Sbjct: 521 PPNECATKDILCTSGHLLTTDSTKVPLQLASHPLCQHHQTSILKNCIHCDA 369
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.124 0.353
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,317,115
Number of Sequences: 36976
Number of extensions: 84680
Number of successful extensions: 428
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 425
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 427
length of query: 204
length of database: 9,014,727
effective HSP length: 92
effective length of query: 112
effective length of database: 5,612,935
effective search space: 628648720
effective search space used: 628648720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146329.15


