

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.11 + phase: 0
(327 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
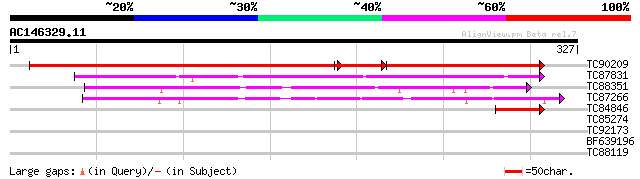
TC90209 similar to GP|13926307|gb|AAK49620.1 AT4g08790/T32A17_10... 350 e-158
TC87831 similar to GP|19715574|gb|AAL91613.1 AT5g12040/F14F18_21... 148 2e-36
TC88351 similar to GP|5262946|emb|CAB45873.1 beta-alanine syntha... 88 4e-18
TC87266 similar to PIR|T03739|T03739 nitrilase (EC 3.5.5.1) 4B -... 63 1e-10
TC84846 similar to GP|13926307|gb|AAK49620.1 AT4g08790/T32A17_10... 55 3e-08
TC85274 similar to PIR|T12282|T12282 L-ascorbate peroxidase (EC ... 33 0.15
TC92173 similar to SP|Q42965|NRL4_TOBAC Nitrilase 4 (EC 3.5.5.1)... 32 0.26
BF639196 similar to GP|20259512|gb| unknown protein {Arabidopsis... 28 4.9
TC88119 similar to GP|10178284|emb|CAC08342. rev interacting pro... 27 8.3
>TC90209 similar to GP|13926307|gb|AAK49620.1 AT4g08790/T32A17_100
{Arabidopsis thaliana}, partial (85%)
Length = 1095
Score = 350 bits (899), Expect(3) = e-158
Identities = 177/181 (97%), Positives = 178/181 (97%)
Frame = +3
Query: 12 IRINNTNGTSIRRRLRASLSVTTGESTMATNSVRVAAAQMTSITDLASNFSTCSRLVKEA 71
+ INNTNGTSIRRRLRASLSVTTGESTMATNSVRVAAAQMTSITDLASNFSTCSRLVKEA
Sbjct: 21 LAINNTNGTSIRRRLRASLSVTTGESTMATNSVRVAAAQMTSITDLASNFSTCSRLVKEA 200
Query: 72 ASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGS 131
ASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGS
Sbjct: 201 ASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGS 380
Query: 132 DPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRL 191
DPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIG
Sbjct: 381 DPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGPF 560
Query: 192 G 192
G
Sbjct: 561 G 563
Score = 183 bits (465), Expect(3) = e-158
Identities = 91/91 (100%), Positives = 91/91 (100%)
Frame = +2
Query: 218 LVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVG 277
LVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVG
Sbjct: 641 LVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVG 820
Query: 278 RLPDRLSTGIVVADIDLSLVDSVREKMPIAK 308
RLPDRLSTGIVVADIDLSLVDSVREKMPIAK
Sbjct: 821 RLPDRLSTGIVVADIDLSLVDSVREKMPIAK 913
Score = 64.3 bits (155), Expect(3) = e-158
Identities = 29/30 (96%), Positives = 30/30 (99%)
Frame = +1
Query: 188 IGRLGLSVCYDLRFPELYQLLRFQHGAQIL 217
+GRLGLSVCYDLRFPELYQLLRFQHGAQIL
Sbjct: 550 LGRLGLSVCYDLRFPELYQLLRFQHGAQIL 639
>TC87831 similar to GP|19715574|gb|AAL91613.1 AT5g12040/F14F18_210
{Arabidopsis thaliana}, partial (82%)
Length = 1378
Score = 148 bits (374), Expect = 2e-36
Identities = 87/276 (31%), Positives = 143/276 (51%), Gaps = 5/276 (1%)
Frame = +2
Query: 38 TMATNSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVS 97
T + ++ Q++ +D N + +++AA+ GAKL+ PE ++ + D V
Sbjct: 350 TPPLTNFKIGLCQLSVTSDKDKNIAHARTAIQDAAAKGAKLILLPEIWNSPYSNDSFPV- 526
Query: 98 IAQPLDG-----PIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTT 152
A+ +D P L+ I + G E+ D L+NT V GK++
Sbjct: 527 YAEDIDAGGDASPSTAMLSELSSLLKITIVGGSIPERSGD--RLYNTCCVFGTDGKLKAK 700
Query: 153 YRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQH 212
+RKIHLFD+D+PG + ES +G VD+ +GR+G+ +CYD+RFPEL ++
Sbjct: 701 HRKIHLFDIDIPGKITFIESLTLTAGDTPTIVDTEVGRIGIGICYDIRFPEL-AMIYAAR 877
Query: 213 GAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPW 272
GA +L P AF TG HWE+L RARA +NQ YV + A ++G + ++ P+
Sbjct: 878 GAHLLCYPGAFNMTTGPLHWELLQRARATDNQLYVATCSPARDTTGGYVAWGHSTLVGPF 1057
Query: 273 GTVVGRLPDRLSTGIVVADIDLSLVDSVREKMPIAK 308
G V+ +T ++A+ID S+++ R +P+ K
Sbjct: 1058GEVLATTEHEETT--IIAEIDYSILEQRRTNLPVTK 1159
>TC88351 similar to GP|5262946|emb|CAB45873.1 beta-alanine synthase
{Lycopersicon esculentum}, partial (98%)
Length = 1255
Score = 88.2 bits (217), Expect = 4e-18
Identities = 73/276 (26%), Positives = 130/276 (46%), Gaps = 18/276 (6%)
Frame = +3
Query: 44 VRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFS---FVGAKDGDSVSIAQ 100
V V+A Q D+++N +T RLV+ A GA ++ E F F A+ D + A+
Sbjct: 105 VVVSALQFACTDDVSTNVTTAERLVRAAHKQGANIVLIQELFEGYYFCQAQREDFIQRAK 284
Query: 101 PL-DGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLF 159
P D P + + LA+E + + + F+E + +N+ ++D G YRK H
Sbjct: 285 PYKDHPTIMRLQKLAKELGVVIPVSFFEEANNAH---YNSIAIIDADGTDLGIYRKSH-- 449
Query: 160 DVDVPGGRVYKESNFTESGKDIVAV-DSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILL 218
+P G Y+E + G V + ++G+++C+D FPE + + Q GA+IL
Sbjct: 450 ---IPDGPGYEEKFYFNPGDTGFKVFQTKYAKIGVAICWDQWFPEAARAMALQ-GAEILF 617
Query: 219 VPAAF------TKVTGEAHWEILLRARAIENQCYVIAAAQAG-----THNDKRE--SYGD 265
P A + HW+ +++ A N ++A+ + G T + K E YG+
Sbjct: 618 YPTAIGSEPHDQSIDSRDHWKRVMQGHAGANLVPLVASNRIGNEIIETEHGKSEIKFYGN 797
Query: 266 TLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVR 301
+ I P G +V + D +++A+ +L + S+R
Sbjct: 798 SFIAGPTGEIVS-IADDKEEAVLIAEFNLDKIKSMR 902
>TC87266 similar to PIR|T03739|T03739 nitrilase (EC 3.5.5.1) 4B - common
tobacco, partial (97%)
Length = 1382
Score = 63.2 bits (152), Expect = 1e-10
Identities = 76/322 (23%), Positives = 131/322 (40%), Gaps = 44/322 (13%)
Frame = +3
Query: 43 SVRVAAAQMTSIT-DLASNFSTCSRLVKEAASAGAKLLCFPEAF-------SFVGAKDGD 94
+VR Q ++I D + RL+ EAAS G++L+ FPEAF S G G+
Sbjct: 123 TVRATVVQASTIFYDTPATLDKAERLLAEAASYGSQLVVFPEAFIGGYPRGSGFGVSIGN 302
Query: 95 SV-----------SIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVV 143
S A + GP +D+ +A + + L +G + G L+ T +
Sbjct: 303 RTAKGREDFRKYHSAAIDVPGPEVDRLAVMAGKYKVHLVMGVIERDGYT---LYCTVLFF 473
Query: 144 DDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPE 203
D G +RKI +P F + G I ++ IG++G ++C++ + P
Sbjct: 474 DSQGHYLGKHRKI------MPTALERIIWGFGD-GSTIPVFETQIGKIGAAICWENKMP- 629
Query: 204 LYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRES- 262
L + + G +I P A ++ W+ + A+E C+V++A Q D +
Sbjct: 630 LLRTAMYAKGVEIYCAPTADSREL----WQASMTHIALEGGCFVLSANQFCRRKDYPPAP 797
Query: 263 ------------------YGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVREKM 304
G ++II P G V+ P+ ++ AD+DL + +
Sbjct: 798 DYVFEGSEENLTPDSVVCAGGSVIISPSGAVLAG-PNYEGEALISADLDLGEIARAKFDF 974
Query: 305 PIA------KVIVCNVYIHPTN 320
+ +V+ V HPTN
Sbjct: 975 DVVGHYARPEVLSLIVKDHPTN 1040
>TC84846 similar to GP|13926307|gb|AAK49620.1 AT4g08790/T32A17_100
{Arabidopsis thaliana}, partial (9%)
Length = 510
Score = 55.5 bits (132), Expect = 3e-08
Identities = 28/28 (100%), Positives = 28/28 (100%)
Frame = +3
Query: 281 DRLSTGIVVADIDLSLVDSVREKMPIAK 308
DRLSTGIVVADIDLSLVDSVREKMPIAK
Sbjct: 204 DRLSTGIVVADIDLSLVDSVREKMPIAK 287
>TC85274 similar to PIR|T12282|T12282 L-ascorbate peroxidase (EC 1.11.1.11)
precursor - common ice plant, partial (86%)
Length = 2018
Score = 33.1 bits (74), Expect = 0.15
Identities = 35/137 (25%), Positives = 55/137 (39%), Gaps = 3/137 (2%)
Frame = +1
Query: 3 LSSCWVNNNIRINNTNGTSIRRRLRASLSVTTGESTMATNSVRVAAAQMTSITDLASNFS 62
L+SC + + + ++ + S+ L SLS++ S + NS R +++ T T
Sbjct: 328 LASCRIRHEVSLSLSLSLSLSLSLSLSLSLSLSLSLLVPNSARGSSSPTTMATITG---G 498
Query: 63 TCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQPL-DGPIMDQYCSLARESSIWL 121
+R++ A A L FSF A SVS L P + R + +
Sbjct: 499 AAARMIPSATRATVSLSTSRSFFSFSLASSSRSVSSLNCLRSSPRISHIFLNQRRGEVRV 678
Query: 122 SLGGFQEK--GSDPRHL 136
S G F SDP L
Sbjct: 679 SSGRFGTVAFASDPDQL 729
>TC92173 similar to SP|Q42965|NRL4_TOBAC Nitrilase 4 (EC 3.5.5.1). [Common
tobacco] {Nicotiana tabacum}, partial (16%)
Length = 491
Score = 32.3 bits (72), Expect = 0.26
Identities = 18/44 (40%), Positives = 25/44 (55%), Gaps = 1/44 (2%)
Frame = +1
Query: 43 SVRVAAAQMTSIT-DLASNFSTCSRLVKEAASAGAKLLCFPEAF 85
+VR Q ++I D + RLV EAA G++L+ FPEAF
Sbjct: 79 TVRATVVQASTIYYDTPATLVKAERLVAEAAGNGSQLVAFPEAF 210
>BF639196 similar to GP|20259512|gb| unknown protein {Arabidopsis thaliana},
partial (9%)
Length = 460
Score = 28.1 bits (61), Expect = 4.9
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = -3
Query: 104 GPIMDQYCSLARESSIWLSLG 124
GP + +C L+RE S W+S G
Sbjct: 65 GPYLQVFCQLSREKSHWVSAG 3
>TC88119 similar to GP|10178284|emb|CAC08342. rev interacting protein
mis3-like {Arabidopsis thaliana}, partial (79%)
Length = 1369
Score = 27.3 bits (59), Expect = 8.3
Identities = 17/41 (41%), Positives = 21/41 (50%)
Frame = -3
Query: 59 SNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIA 99
SNFST + ++S G L CFP F F + D SIA
Sbjct: 260 SNFSTFQWSMLGSSSHGFGLSCFPFFFPFTTSFDTSFSSIA 138
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,978,312
Number of Sequences: 36976
Number of extensions: 124075
Number of successful extensions: 525
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 519
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 523
length of query: 327
length of database: 9,014,727
effective HSP length: 96
effective length of query: 231
effective length of database: 5,465,031
effective search space: 1262422161
effective search space used: 1262422161
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146329.11


