

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.10 - phase: 0
(68 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
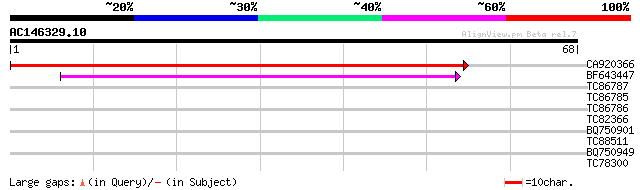
CA920366 weakly similar to GP|20161089|dbj putative ubiquitin-co... 69 5e-13
BF643447 similar to GP|16323334|gb AT3g15355/MJK13_1 {Arabidopsi... 49 3e-07
TC86787 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidop... 26 3.4
TC86785 similar to GP|13876504|gb|AAK43480.1 hypothetical protei... 26 3.4
TC86786 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidop... 26 3.4
TC82366 similar to GP|13160609|dbj|BAB32954. Ara4-interacting pr... 25 7.5
BQ750901 similar to SP|Q01115|FRQ_ Frequency clock protein. {Lep... 25 7.5
TC88511 weakly similar to GP|20259567|gb|AAM14126.1 unknown prot... 25 7.5
BQ750949 similar to PIR|S69206|S69 regulator protein white colla... 24 9.9
TC78300 similar to GP|17978899|gb|AAL47419.1 AT4g26860/F10M23_20... 24 9.9
>CA920366 weakly similar to GP|20161089|dbj putative ubiquitin-conjugating
protein {Oryza sativa (japonica cultivar-group)},
partial (10%)
Length = 696
Score = 68.6 bits (166), Expect = 5e-13
Identities = 30/55 (54%), Positives = 42/55 (75%)
Frame = -3
Query: 1 MVAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIGS 55
+VA AY++GAQVGCLVKGGVQD+D GDKS S++F SL ++++ FT +G+
Sbjct: 484 LVACKAYMDGAQVGCLVKGGVQDVDQGDKSCSKEFKQSLPAYVDMLVKEFTQVGA 320
>BF643447 similar to GP|16323334|gb AT3g15355/MJK13_1 {Arabidopsis thaliana},
partial (21%)
Length = 671
Score = 49.3 bits (116), Expect = 3e-07
Identities = 23/48 (47%), Positives = 29/48 (59%)
Frame = +1
Query: 7 YIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIG 54
Y+EG QVGCL KGGVQD+D+G+ S +F L +N F IG
Sbjct: 418 YMEGVQVGCLAKGGVQDVDEGEGKCSDKFKADLVGLVNNLVKEFERIG 561
>TC86787 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidopsis
thaliana}, partial (23%)
Length = 841
Score = 25.8 bits (55), Expect = 3.4
Identities = 10/23 (43%), Positives = 17/23 (73%)
Frame = +1
Query: 29 KSSSRQFNNSLARHINVAAPNFT 51
KSS + NNS+++H +A+ +FT
Sbjct: 283 KSSYKYSNNSISQHSTIASSHFT 351
>TC86785 similar to GP|13876504|gb|AAK43480.1 hypothetical protein
{Arabidopsis thaliana}, partial (8%)
Length = 663
Score = 25.8 bits (55), Expect = 3.4
Identities = 10/23 (43%), Positives = 17/23 (73%)
Frame = +2
Query: 29 KSSSRQFNNSLARHINVAAPNFT 51
KSS + NNS+++H +A+ +FT
Sbjct: 296 KSSYKYSNNSISQHSTIASSHFT 364
>TC86786 similar to GP|8778315|gb|AAF79324.1| F14J16.29 {Arabidopsis
thaliana}, partial (32%)
Length = 707
Score = 25.8 bits (55), Expect = 3.4
Identities = 10/23 (43%), Positives = 17/23 (73%)
Frame = +3
Query: 29 KSSSRQFNNSLARHINVAAPNFT 51
KSS + NNS+++H +A+ +FT
Sbjct: 324 KSSYKYSNNSISQHSTIASSHFT 392
>TC82366 similar to GP|13160609|dbj|BAB32954. Ara4-interacting protein
{Arabidopsis thaliana}, partial (16%)
Length = 772
Score = 24.6 bits (52), Expect = 7.5
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = +3
Query: 30 SSSRQFNNSLARHINVAAPN 49
SSS+Q NS+A I++A PN
Sbjct: 102 SSSKQTKNSIAISISMATPN 161
>BQ750901 similar to SP|Q01115|FRQ_ Frequency clock protein. {Leptosphaeria
australiensis}, partial (15%)
Length = 743
Score = 24.6 bits (52), Expect = 7.5
Identities = 11/37 (29%), Positives = 19/37 (50%)
Frame = +2
Query: 26 DGDKSSSRQFNNSLARHINVAAPNFTGIGSMLVQAPD 62
D D+ + N RH+ +A P+ TG+ + +PD
Sbjct: 200 DPDRLQNPTENMDYIRHLGLAPPHLTGLATYDDVSPD 310
>TC88511 weakly similar to GP|20259567|gb|AAM14126.1 unknown protein
{Arabidopsis thaliana}, partial (22%)
Length = 1060
Score = 24.6 bits (52), Expect = 7.5
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = -1
Query: 31 SSRQFNNSLARHINVAAPNFTGIGSM 56
+SR+ S +R + +A P F+G+G M
Sbjct: 370 NSRKMLESSSRSVALATPGFSGLGVM 293
>BQ750949 similar to PIR|S69206|S69 regulator protein white collar 1 -
Neurospora crassa, partial (7%)
Length = 276
Score = 24.3 bits (51), Expect = 9.9
Identities = 13/37 (35%), Positives = 19/37 (51%)
Frame = -1
Query: 13 VGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPN 49
V C V V+D+DDG ++ ARH++ A N
Sbjct: 165 VSCQVLKVVRDVDDGAVVERDVADDKSARHVHSAEMN 55
>TC78300 similar to GP|17978899|gb|AAL47419.1 AT4g26860/F10M23_200
{Arabidopsis thaliana}, partial (94%)
Length = 1051
Score = 24.3 bits (51), Expect = 9.9
Identities = 9/24 (37%), Positives = 14/24 (57%)
Frame = +1
Query: 39 LARHINVAAPNFTGIGSMLVQAPD 62
LA+H+ ++ PN G M + PD
Sbjct: 517 LAKHVKLSCPNLEFSGLMTIGMPD 588
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.135 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,542,702
Number of Sequences: 36976
Number of extensions: 13380
Number of successful extensions: 72
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 72
length of query: 68
length of database: 9,014,727
effective HSP length: 44
effective length of query: 24
effective length of database: 7,387,783
effective search space: 177306792
effective search space used: 177306792
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC146329.10


