

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
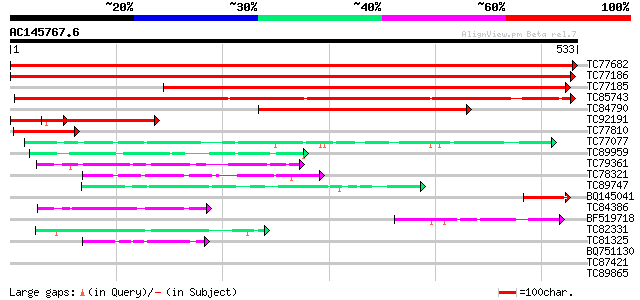
Query= AC145767.6 - phase: 0
(533 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC77682 phosphate transporter 1067 0.0
TC77186 phosphate transporter 1059 0.0
TC77185 similar to GP|12697486|emb|CAC28219. phosphate transport... 665 0.0
TC85743 phosphate transporter PT4 [Medicago truncatula] 651 0.0
TC84790 similar to GP|13676624|gb|AAK38197.1 phosphate transport... 341 5e-94
TC92191 homologue to GP|12697484|emb|CAC28218. phosphate transpo... 197 8e-51
TC77810 similar to GP|13676624|gb|AAK38197.1 phosphate transport... 94 1e-19
TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter p... 60 2e-09
TC89959 similar to GP|15795137|dbj|BAB02515. transporter-like pr... 59 5e-09
TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 58 8e-09
TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 58 1e-08
TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported]... 57 2e-08
BQ145041 similar to GP|13676624|gb| phosphate transporter 2 {Lup... 53 3e-07
TC84386 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 51 1e-06
BF519718 weakly similar to PIR|G84564|G84 probable sugar transpo... 49 6e-06
TC82331 weakly similar to EGAD|120604|128852 hypothetical protei... 46 3e-05
TC81325 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 44 2e-04
BQ751130 weakly similar to GP|15289796|dbj putative monosacchari... 41 0.001
TC87421 sugar transporter 40 0.002
TC89865 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 40 0.003
>TC77682 phosphate transporter
Length = 2245
Score = 1067 bits (2760), Expect = 0.0
Identities = 533/533 (100%), Positives = 533/533 (100%)
Frame = +2
Query: 1 MSGELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTR 60
MSGELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTR
Sbjct: 119 MSGELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTR 298
Query: 61 PGTLPPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGS 120
PGTLPPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGS
Sbjct: 299 PGTLPPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGS 478
Query: 121 SPKSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGI 180
SPKSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGI
Sbjct: 479 SPKSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGI 658
Query: 181 VALTVASIFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETA 240
VALTVASIFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETA
Sbjct: 659 VALTVASIFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETA 838
Query: 241 RYTALVAKNAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQFAARHGLALFG 300
RYTALVAKNAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQFAARHGLALFG
Sbjct: 839 RYTALVAKNAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQFAARHGLALFG 1018
Query: 301 TCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGY 360
TCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGY
Sbjct: 1019TCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGY 1198
Query: 361 WFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVVIYSLTFFFANFG 420
WFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVVIYSLTFFFANFG
Sbjct: 1199WFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVVIYSLTFFFANFG 1378
Query: 421 PNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGI 480
PNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGI
Sbjct: 1379PNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGI 1558
Query: 481 GIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQEGSRV 533
GIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQEGSRV
Sbjct: 1559GIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQEGSRV 1717
>TC77186 phosphate transporter
Length = 2314
Score = 1059 bits (2739), Expect = 0.0
Identities = 528/532 (99%), Positives = 530/532 (99%)
Frame = +1
Query: 1 MSGELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTR 60
MSGELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTR
Sbjct: 397 MSGELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTR 576
Query: 61 PGTLPPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGS 120
PGTLPPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGS
Sbjct: 577 PGTLPPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGS 756
Query: 121 SPKSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGI 180
SPKSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGI
Sbjct: 757 SPKSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGI 936
Query: 181 VALTVASIFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETA 240
VAL VASIFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETA
Sbjct: 937 VALIVASIFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETA 1116
Query: 241 RYTALVAKNAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQFAARHGLALFG 300
RYTALVAKNAKQAAADMSKVLQVELEVEEEKV+KMTSDKRNSYGLFSKQFAARHGLALFG
Sbjct: 1117RYTALVAKNAKQAAADMSKVLQVELEVEEEKVQKMTSDKRNSYGLFSKQFAARHGLALFG 1296
Query: 301 TCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGY 360
TCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGY
Sbjct: 1297TCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGY 1476
Query: 361 WFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVVIYSLTFFFANFG 420
WFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVV+YSLTFFFANFG
Sbjct: 1477WFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVVMYSLTFFFANFG 1656
Query: 421 PNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGI 480
PNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGI
Sbjct: 1657PNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGI 1836
Query: 481 GIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQEGSR 532
GIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQEG R
Sbjct: 1837GIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQEGPR 1992
>TC77185 similar to GP|12697486|emb|CAC28219. phosphate transporter
{Sesbania rostrata}, partial (71%)
Length = 1467
Score = 665 bits (1715), Expect = 0.0
Identities = 322/383 (84%), Positives = 355/383 (92%)
Frame = +2
Query: 145 PLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHKYKVPTFEENPAA 204
PLSATIMSEYANKKTRGAFIAAVFAMQGFGI+ GG+ AL VA+ FDHKY+VPT+EEN A
Sbjct: 2 PLSATIMSEYANKKTRGAFIAAVFAMQGFGIMAGGVFALIVAAAFDHKYQVPTYEENAKA 181
Query: 205 SLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAKNAKQAAADMSKVLQVE 264
SL++P FDYVWR+ILMFGA+PAALTYYWRMKMPETARYTALVAKN KQAA+DMSKVLQVE
Sbjct: 182 SLVLPAFDYVWRIILMFGAVPAALTYYWRMKMPETARYTALVAKNGKQAASDMSKVLQVE 361
Query: 265 LEVEEEKVEKMTSDKRNSYGLFSKQFAARHGLALFGTCSTWFLLDIAFYSQNLFQKDIFS 324
+E EEEKV+ + ++ +GLF++QFA RHGL L GT +TWFLLDIAFYSQNLFQKDIF+
Sbjct: 362 IEAEEEKVQNLAENQNQKFGLFTRQFAKRHGLHLLGTTTTWFLLDIAFYSQNLFQKDIFT 541
Query: 325 AIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVAFIDHMGRFAIQMMGFFFMT 384
AIGWIPPAKEMNAI+E+Y+IARAQTLIALCSTVPGYWFTVAFID+MGRFAIQ+MGFFFMT
Sbjct: 542 AIGWIPPAKEMNAINELYRIARAQTLIALCSTVPGYWFTVAFIDYMGRFAIQLMGFFFMT 721
Query: 385 VFMFALAIPYDHWSKEENRIGFVVIYSLTFFFANFGPNATTFVVPAEIFPARLRSTCHGI 444
VFMFALAIPYDHW+K+ENRIGFVV+YSLTFFFANFGPNATTFVVPAEIFPARLRSTCHGI
Sbjct: 722 VFMFALAIPYDHWTKKENRIGFVVMYSLTFFFANFGPNATTFVVPAEIFPARLRSTCHGI 901
Query: 445 SAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLLVP 504
SAAAGKAGAIVGAFGFLYAAQSKDPTKTD GYPTGIGIKNSLI+LGV+NF GM+ T LVP
Sbjct: 902 SAAAGKAGAIVGAFGFLYAAQSKDPTKTDHGYPTGIGIKNSLIVLGVVNFFGMVFTFLVP 1081
Query: 505 ESKGKSLEELSGENEGEGAEATE 527
E GKSLEE+SGENE + AEA E
Sbjct: 1082EPNGKSLEEMSGENEDDDAEAIE 1150
>TC85743 phosphate transporter PT4 [Medicago truncatula]
Length = 1850
Score = 651 bits (1679), Expect = 0.0
Identities = 327/528 (61%), Positives = 400/528 (74%)
Frame = +2
Query: 5 LGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPGTL 64
L VL ALD A+TQ YH T IVIAGMGFFTDAYDLFCIS V+KLLGR+YY +P+ +PG L
Sbjct: 35 LEVLEALDSARTQWYHVTAIVIAGMGFFTDAYDLFCISTVSKLLGRLYYFDPSTNKPGKL 214
Query: 65 PPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKS 124
PPS + VTGVALVGTL+GQL FGWLGDKLGRKKVYG+TLI+MV C++ SGLSFGSS KS
Sbjct: 215 PPSVNNVVTGVALVGTLSGQLVFGWLGDKLGRKKVYGVTLIIMVACAICSGLSFGSSAKS 394
Query: 125 VMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALT 184
VM TLCFFRFWLGFGIGGDYPLSATIMSEYANK+TRGAFIAAVFAMQG GI+ G+V++
Sbjct: 395 VMITLCFFRFWLGFGIGGDYPLSATIMSEYANKRTRGAFIAAVFAMQGVGIIFAGLVSMV 574
Query: 185 VASIFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTA 244
+ IF Y+ P F E+P S P+ D +WRLILM GA+PAA+TYYWRMKMPET RYTA
Sbjct: 575 FSGIFKAYYQAPRFNEDPILS-TQPEGDLLWRLILMIGAVPAAMTYYWRMKMPETGRYTA 751
Query: 245 LVAKNAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQFAARHGLALFGTCST 304
+V NAKQAAADM++VL +E+ E++K+ + + N Y L+S +F RHG L GT S
Sbjct: 752 IVEGNAKQAAADMARVLDIEIIAEQDKLAEFKA--ANDYPLWSSEFFNRHGRHLIGTMSC 925
Query: 305 WFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTV 364
WFLLDIAFYSQNL QKDI+ A+G I KEMNAI EV++ +RA ++AL T PGYWFTV
Sbjct: 926 WFLLDIAFYSQNLTQKDIYPAMGLIRQDKEMNAIDEVFQTSRAMFVVALFGTFPGYWFTV 1105
Query: 365 AFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVVIYSLTFFFANFGPNAT 424
FI+ +GRF IQ++GFF M+ FMF + + Y+ + K+EN+ F ++Y LTFFFANFGPN+T
Sbjct: 1106FFIEKLGRFKIQLVGFFMMSFFMFVIGVKYE-YLKDENKNLFALLYGLTFFFANFGPNST 1282
Query: 425 TFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKN 484
TFV+PAE+FP R+RSTCH SAA+GKAGA+VGAFG Y P K I+
Sbjct: 1283TFVLPAELFPTRVRSTCHAFSAASGKAGAMVGAFGIQYYTLDGTPRK----------IRR 1432
Query: 485 SLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQEGSR 532
++++L N +G CT LV E+KG+SLEE+SGE +G +E T R
Sbjct: 1433AMMILAFTNLIGFFCTFLVTETKGRSLEEISGE-DGRESELTATPNDR 1573
>TC84790 similar to GP|13676624|gb|AAK38197.1 phosphate transporter 2
{Lupinus albus}, partial (37%)
Length = 639
Score = 341 bits (874), Expect = 5e-94
Identities = 160/200 (80%), Positives = 183/200 (91%)
Frame = +2
Query: 235 KMPETARYTALVAKNAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQFAARH 294
KMPETARYTALVAKNAKQAA DMS VLQVE+E E+EKV+K+ +NS+GLFSK+F RH
Sbjct: 2 KMPETARYTALVAKNAKQAAQDMSSVLQVEIEAEQEKVDKIGVQDKNSFGLFSKEFLRRH 181
Query: 295 GLALFGTCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALC 354
GL L GT STWFLLDIA+YS NLFQKDI+S+IGW+PPA++MNAIHEV+K+ARA TLIALC
Sbjct: 182 GLHLLGTTSTWFLLDIAYYSSNLFQKDIYSSIGWLPPAQDMNAIHEVFKVARATTLIALC 361
Query: 355 STVPGYWFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVVIYSLTF 414
TVPGYWFTVAFID +GRFAIQ+MGFFFMTVFMFALAIPYDHW+K ENRIGF+V+Y++TF
Sbjct: 362 GTVPGYWFTVAFIDVIGRFAIQLMGFFFMTVFMFALAIPYDHWTKRENRIGFLVMYAMTF 541
Query: 415 FFANFGPNATTFVVPAEIFP 434
FFANFGPN+TTFVVPAEIFP
Sbjct: 542 FFANFGPNSTTFVVPAEIFP 601
>TC92191 homologue to GP|12697484|emb|CAC28218. phosphate transporter
{Sesbania rostrata}, partial (26%)
Length = 531
Score = 197 bits (501), Expect = 8e-51
Identities = 93/111 (83%), Positives = 101/111 (90%)
Frame = +2
Query: 31 FFTDAYDLFCISLVTKLLGRIYYTEPNPTRPGTLPPSAQSAVTGVALVGTLAGQLFFGWL 90
F DAYDLFCISLVTKLLGRIYYT+P +PG LPP Q+AVTGVALVGTL+GQLFFGWL
Sbjct: 197 FSLDAYDLFCISLVTKLLGRIYYTDPTQPKPGVLPPGVQAAVTGVALVGTLSGQLFFGWL 376
Query: 91 GDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRFWLGFGIG 141
GDKLGRKKVYG+TL+LMV CS+ASGLSFG+SPK VMATLCFFRFWLGFGIG
Sbjct: 377 GDKLGRKKVYGITLMLMVGCSLASGLSFGNSPKGVMATLCFFRFWLGFGIG 529
Score = 64.3 bits (155), Expect = 1e-10
Identities = 37/60 (61%), Positives = 42/60 (69%), Gaps = 5/60 (8%)
Frame = +1
Query: 1 MSGELGVLNALDVAKTQLYHFTTIVIAGMGFFT----DAYDLFCISLV-TKLLGRIYYTE 55
M+G+LGVLNALDVAKTQ+YHFT IVIAGMGFFT C +V T LL R Y T+
Sbjct: 106 MAGQLGVLNALDVAKTQMYHFTAIVIAGMGFFTRCL*SLLHFPCHKIVRTNLLHRSYPTK 285
>TC77810 similar to GP|13676624|gb|AAK38197.1 phosphate transporter 2
{Lupinus albus}, partial (11%)
Length = 1035
Score = 94.0 bits (232), Expect = 1e-19
Identities = 45/62 (72%), Positives = 50/62 (80%)
Frame = +2
Query: 4 ELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPGT 63
+LGVL +L AKTQ Y FT I+IAGMGF TDAYDLFCI VTKLLGRIYYT P+ +PGT
Sbjct: 344 QLGVLTSLHAAKTQWYLFTAIIIAGMGFCTDAYDLFCIPNVTKLLGRIYYTHPDALKPGT 523
Query: 64 LP 65
LP
Sbjct: 524 LP 529
>TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter protein
{Arabidopsis thaliana}, partial (85%)
Length = 1752
Score = 60.5 bits (145), Expect = 2e-09
Identities = 115/536 (21%), Positives = 196/536 (36%), Gaps = 36/536 (6%)
Frame = +1
Query: 15 KTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPGTLPPSAQSAVTG 74
K Y F ++A M YD+ +S IY G + + G
Sbjct: 94 KRNKYAFACAMLASMTSILLGYDIGVMSGAA-----IYIKRDLKVSDGKI-----EVLLG 243
Query: 75 VALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRF 134
+ + +L G G D +GR+ T++ L G SP L F RF
Sbjct: 244 IINIYSLIGSCLAGRTSDWIGRR----YTIVFAGAIFFVGALLMGFSPNYNF--LMFGRF 405
Query: 135 WLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHKYK 194
G GIG ++ +E + +RG + GIL G I
Sbjct: 406 VAGVGIGYALMIAPVYTAEVSPASSRGFLTSFPEVFINSGILLGYI-------------- 543
Query: 195 VPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAK------ 248
N A S L + WR++L GA+P+ + + MPE+ R+ + +
Sbjct: 544 -----SNYAFSKLSLKLG--WRMMLGVGAIPSVILAVGVLAMPESPRWLVMRGRLGDAIK 702
Query: 249 ----------NAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQF-----AAR 293
A+ A++ + + + ++ VE K G++ + F A R
Sbjct: 703 VLNKTSDSKEEAQLRLAEIKQAAGIPEDCNDDVVEVKV--KNTGEGVWKELFLYPTPAVR 876
Query: 294 H------GLALFGTCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARA 347
H G+ F S + + YS +F+K + + + + +
Sbjct: 877 HIVIAALGIHFFQQASG--VDAVVLYSPTIFKKAGING--------DTHLLIATIAVGFV 1026
Query: 348 QTLIALCSTVPGYWFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPY---DHWSKEEN-- 402
+TL L +T +D GR + + M V + LA+ DH + + N
Sbjct: 1027KTLFILVATF--------MLDRYGRRPLLLTSVGGMVVSLLTLAVSLTIIDHSNTKLNWA 1182
Query: 403 ---RIGFVVIYSLTFFFANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFG 459
I V+ Y TF + G T+V +EIFP RLR+ + + V +
Sbjct: 1183IGLSIATVLSYVATF---SIGAGPITWVYSSEIFPLRLRAQGAACGVVVNRVTSGVISMT 1353
Query: 460 FLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVG-MLCTLLVPESKGKSLEEL 514
FL ++ GI I + + G I +G + +++PE++GK+LEE+
Sbjct: 1354FLSLSK-------------GITIGGAFFLFGGIATIGWIFFYIMLPETQGKTLEEM 1482
>TC89959 similar to GP|15795137|dbj|BAB02515. transporter-like protein
{Arabidopsis thaliana}, partial (87%)
Length = 1681
Score = 58.9 bits (141), Expect = 5e-09
Identities = 59/265 (22%), Positives = 105/265 (39%), Gaps = 2/265 (0%)
Frame = +3
Query: 19 YHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPGTLPPSAQSAVTGVALV 78
+ ++ AG+G+ ++A ++ +S V P L +S +T V
Sbjct: 186 FQILVLIYAGIGWISEAMEMMLLSFVG----------PAVQSAWNLSSHQESFITSVVFA 335
Query: 79 GTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRFWLGF 138
G L G +G + D+ GR+K + ++ +V + F SS +L FR +G
Sbjct: 336 GMLIGAYSWGIVSDQHGRRKGF------LITATVTATAGFFSSLSPNYVSLLVFRSLVGL 497
Query: 139 GIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHKYKVPTF 198
G+GG LS+ + E+ RG ++ +F + V T
Sbjct: 498 GLGGGPVLSSWFL-EFVPAPNRGTWMV----------------------VFSGFWTVGTI 608
Query: 199 EENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAKNAKQAAADMS 258
E A +++P+ WR +L +LP + + PE+ RY L K +
Sbjct: 609 FEASLAWIVMPRLG--WRWLLALSSLPTSFLLLFYKMTPESPRYLCL-----KGRTTEAI 767
Query: 259 KVLQVELEVEEEKVEK--MTSDKRN 281
VL+ + +K+ + SDK N
Sbjct: 768 DVLETISRLNGKKLPSGVLVSDKSN 842
Score = 36.2 bits (82), Expect = 0.032
Identities = 31/168 (18%), Positives = 71/168 (41%), Gaps = 5/168 (2%)
Frame = +1
Query: 351 IALCSTVPGYWFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFV--- 407
IA + +PG + +D +GR + FF +F+ +P + E+ G +
Sbjct: 1141 IASFAELPGLLLSAVAVDKLGRKLSMSIMFFMCCIFL----LPVTFYLPEDLTTGLLFGA 1308
Query: 408 -VIYSLTFFFANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQS 466
+ ++TF ++ EI+P +R+T G++++ G+ G ++
Sbjct: 1309 RICITVTF--------TIVYIYAPEIYPTSVRTTGVGVASSVGRIGGMLCPL-------- 1440
Query: 467 KDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLLVP-ESKGKSLEE 513
G G ++++ ++ V +C + +P E+ G+ L++
Sbjct: 1441 -----VAVGLVHGCHQTAAVLLFIIVALVSGICVVFIPFETMGQELQD 1569
>TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (55%)
Length = 949
Score = 58.2 bits (139), Expect = 8e-09
Identities = 62/256 (24%), Positives = 109/256 (42%), Gaps = 4/256 (1%)
Frame = +1
Query: 26 IAGMGFFTDAYDLFCISLVTKLLGRIYYTE---PNPTRPGTLPPSAQSAVTGVALVGTLA 82
+AG+G YD IS G + Y + P L Q + +A+ G +
Sbjct: 187 VAGIGGLLFGYDTGVIS------GALLYIKDDFPQVRNSNFL----QETIVSMAIAGAIV 336
Query: 83 GQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRFWLGFGIGG 142
G F GWL D GRKK L ++ ++ ++ ++ P ++A R +G G+G
Sbjct: 337 GAAFGGWLNDAYGRKKATLLADVIFILGAIL--MAAAPDPYVLIAG----RLLVGLGVGI 498
Query: 143 DYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHKYKVPTFEENP 202
+ ++E A + RG+ ++ M I GG V+ V
Sbjct: 499 ASVTAPVYIAEVAPSEIRGSLVSTNVLM----ITGGQFVSYLV----------------- 615
Query: 203 AASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAKNAKQAAAD-MSKVL 261
+L+ Q WR +L +PA + + + +PE+ R+ L KN K A D +SK+
Sbjct: 616 --NLVFTQVPGTWRWMLGVSGVPALIQFICMLFLPESPRW--LFIKNRKNEAVDVISKI- 780
Query: 262 QVELEVEEEKVEKMTS 277
+L E++++ +T+
Sbjct: 781 -YDLSRLEDEIDFLTA 825
>TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 2093
Score = 57.8 bits (138), Expect = 1e-08
Identities = 55/234 (23%), Positives = 99/234 (41%), Gaps = 6/234 (2%)
Frame = +2
Query: 69 QSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMAT 128
Q A+ A+ G + G GW+ D+ GRK +++I+ + + ++P AT
Sbjct: 347 QEAIVSTAIAGAIIGAAIGGWINDRFGRK----VSIIVADTLFLLGSIILAAAPNP--AT 508
Query: 129 LCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASI 188
L R ++G G+G S +SE + + RGA + ++ F I GG ++ +
Sbjct: 509 LIVGRVFVGLGVGMASMASPLYISEASPTRVRGALV----SLNSFLITGGQFLSYLINLA 676
Query: 189 FDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAK 248
F K P WR +L A PA + + +PE+ R+ K
Sbjct: 677 FT---KAPG----------------TWRWMLGVAAAPAVIQIVLMLSLPESPRWLYRKGK 799
Query: 249 NAKQAAADMSKVLQV-----ELEVEEEKVE-KMTSDKRNSYGLFSKQFAARHGL 296
++A + K+ +V E++ +E VE ++ ++ S K + R GL
Sbjct: 800 E-EEAKVILKKIYEVEDYDNEIQALKESVEMELKETEKISIMQLVKTTSVRRGL 958
Score = 40.4 bits (93), Expect = 0.002
Identities = 32/115 (27%), Positives = 53/115 (45%), Gaps = 2/115 (1%)
Frame = +2
Query: 402 NRIGFVVIYSLTFFFANFGPNATT--FVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFG 459
+ G++ I +L + F P T +VV +EI+P R R C GI++ +V +
Sbjct: 1475 SNFGWIAILALALYIIFFSPGMGTVPWVVNSEIYPLRYRGICGGIASTTVWVSNLVVSQS 1654
Query: 460 FLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEEL 514
FL + P T + ++I + I FV + VPE+KG +EE+
Sbjct: 1655 FLSLTVAIGPAWTFMIF--------AIIAIVAIFFV----IIFVPETKGVPMEEV 1783
>TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1372
Score = 56.6 bits (135), Expect = 2e-08
Identities = 73/331 (22%), Positives = 130/331 (39%), Gaps = 7/331 (2%)
Frame = +3
Query: 68 AQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMA 127
A+ V + L G L G L GW+ D +GR++ + L + M++ G + ++ ++
Sbjct: 492 AEGLVVSICLGGALFGCLLSGWIADAVGRRRAFQLCALPMII-----GAAMSAATNNLFG 656
Query: 128 TLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVAS 187
L R ++G G+G P++A ++E + RG + A + FGILG + + V
Sbjct: 657 ML-VGRLFVGTGLGLGPPVAALYVTEVSPAFVRGTYGALIQIATCFGILGSLFIGIPVKE 833
Query: 188 IFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVA 247
I ++V + A++L D+ A + +W K TA
Sbjct: 834 I-SGWWRVCFWVSTIPAAILALAMDF------------CAESPHWLYKQGRTA------- 953
Query: 248 KNAKQAAADMSKVLQV-ELEVEEEKVEKMTSDKRNSYGLFSKQFAARHGLALFGTCSTWF 306
+A A+ ++L V E + ++ K+ + FS+ H +F ST F
Sbjct: 954 ----EAEAEFERLLGVSEAKFAMSQLSKVDRGEDTDTVKFSELLHGHHSKVVF-IGSTLF 1118
Query: 307 LL------DIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGY 360
L + FY F +F + G P+ N V + + G
Sbjct: 1119ALQQLSGINAVFY----FSSTVFKSAG--VPSDFANVCIGV-------------ANLTGS 1241
Query: 361 WFTVAFIDHMGRFAIQMMGFFFMTVFMFALA 391
+ +D +GR + FF M + M A
Sbjct: 1242IISTGLMDKLGRKVLLFWSFFGMAISMIIQA 1334
>BQ145041 similar to GP|13676624|gb| phosphate transporter 2 {Lupinus albus},
partial (7%)
Length = 522
Score = 52.8 bits (125), Expect = 3e-07
Identities = 27/44 (61%), Positives = 34/44 (76%)
Frame = +1
Query: 484 NSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATE 527
N+LI+L V+N +G+ T LVPE+ GKSLEE+SGENE E E TE
Sbjct: 1 NTLILLAVVNCLGIFFTFLVPEANGKSLEEMSGENEEE--EETE 126
>TC84386 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (30%)
Length = 664
Score = 50.8 bits (120), Expect = 1e-06
Identities = 45/163 (27%), Positives = 67/163 (40%)
Frame = +2
Query: 27 AGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPGTLPPSAQSAVTGVALVGTLAGQLF 86
AG+G F YD IS G + Y + + Q A+ AL G + G
Sbjct: 206 AGIGGFLFGYDTGVIS------GALLYIRDD-FKAVDRQTWLQEAIVSTALAGAIIGASV 364
Query: 87 FGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRFWLGFGIGGDYPL 146
GW+ D+ GRKK L L + SV + A L R ++G G+G
Sbjct: 365 GGWINDRFGRKKAIILADALFFIGSVIMAAAINP------AILIVGRVFVGLGVGMASMA 526
Query: 147 SATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIF 189
S +SE + + RGA + ++ GF I GG ++ + F
Sbjct: 527 SPLYISEASPTRVRGALV----SLNGFLITGGQXLSYVINLAF 643
>BF519718 weakly similar to PIR|G84564|G84 probable sugar transporter
[imported] - Arabidopsis thaliana, partial (20%)
Length = 659
Score = 48.5 bits (114), Expect = 6e-06
Identities = 47/167 (28%), Positives = 73/167 (43%), Gaps = 7/167 (4%)
Frame = +3
Query: 362 FTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYD--HWSKEENRIGFV-----VIYSLTF 414
F+ +D GR + ++G M V +F L + H S E+ V +++F
Sbjct: 39 FSALVLDRFGRRPMLLLGSSGMAVSLFGLGMGCTLLHNSDEKPMWAIALCVVAVCAAVSF 218
Query: 415 FFANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDK 474
F GP TT+V +EIFP RLR+ G S A I G + + S++ T
Sbjct: 219 FSIGLGP--TTWVYSSEIFPMRLRA--QGTSLAISVNRLISGVVSMSFLSISEEITFGGM 386
Query: 475 GYPTGIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGE 521
+ ++ GV+ + +PE+KGKSLEE+ E E
Sbjct: 387 FF----------VLAGVMVLATLFFYYFLPETKGKSLEEIEALFEEE 497
>TC82331 weakly similar to EGAD|120604|128852 hypothetical protein M01F1.5
{Caenorhabditis elegans}, partial (10%)
Length = 848
Score = 46.2 bits (108), Expect = 3e-05
Identities = 54/226 (23%), Positives = 88/226 (38%), Gaps = 6/226 (2%)
Frame = +3
Query: 25 VIAGMGFFTDAYDLFCIS---LVTKLLGRIYYTEPNPTRPGTLPPSAQSAVTGVALVGTL 81
V+ +G F YD IS L T R + T QS + + +GTL
Sbjct: 255 VVTSIGGFLFGYDTGQISSMLLFTDFRQRFATGPEDANGVKTWVSIIQSLMVSLMSIGTL 434
Query: 82 AGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRFWLGFGIG 141
G L + D GR+K +++ V+ ++ + + +M RF G G+G
Sbjct: 435 IGALSGAYTADWWGRRKSLSFGVVVFVIGNIVQITAMDTWVHMMMG-----RFIAGLGVG 599
Query: 142 GDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHKYKVPTFEEN 201
SE + K+ RGA +A+ M FGIL ++ V +I
Sbjct: 600 NLSVGVPMFQSECSPKEIRGAVVASYQLMITFGILISNLINFGVRNI------------- 740
Query: 202 PAASLLVPQFDYVWRLILMFG---ALPAALTYYWRMKMPETARYTA 244
+ D WR+++ G A+P L + +PE+ R+ A
Sbjct: 741 -------QESDASWRIVIGLGIAFAMPLGLGI---LVVPESPRWLA 848
>TC81325 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (32%)
Length = 636
Score = 43.9 bits (102), Expect = 2e-04
Identities = 34/120 (28%), Positives = 57/120 (47%)
Frame = +3
Query: 69 QSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMAT 128
Q + +ALVG + G GW+ D GRKK TL VV ++ S + S+P + +
Sbjct: 294 QETIVSMALVGAIIGAATGGWINDAFGRKKA---TLSADVVFTLGS-VVMASAPDAYV-- 455
Query: 129 LCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASI 188
L R +G G+G + ++E + + RG+ ++ M I GG + LT ++
Sbjct: 456 LILGRLLVGIGVGVASVTAPVYIAESSPSEIRGSLVSTNVLM----ITGGQVFFLTFVNL 623
>BQ751130 weakly similar to GP|15289796|dbj putative monosaccharide
transporter 3 {Oryza sativa (japonica cultivar-group)},
partial (7%)
Length = 795
Score = 40.8 bits (94), Expect = 0.001
Identities = 55/212 (25%), Positives = 84/212 (38%), Gaps = 4/212 (1%)
Frame = +3
Query: 67 SAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTL-ILMVVCSVASGLSFGSSPKSV 125
S QS + + GT G L G + D +GRK + I M+ + G+ ++
Sbjct: 174 SHQSLIVSILSCGTFFGALIAGDVSDWIGRKWTVIIGCGIYMIGVLIQMFTGMGAPLGAI 353
Query: 126 MATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTV 185
+A R G G+G + + MSE KK RGA +A G+L V
Sbjct: 354 VAG----RLIAGIGVGFESAIVILYMSEICPKKVRGALVAGYQFCITIGLLLAACVVYAT 521
Query: 186 ASIFD-HKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTA 244
D Y++P + P W LIL G L + + +K A TA
Sbjct: 522 KDRGDTGVYRIPIAIQFP------------WALILGGGLLLLPESPRFFVKKGRIADATA 665
Query: 245 LVAKNAKQAAADMSKVLQVELE--VEEEKVEK 274
+++ Q S+ +QVEL V E+ E+
Sbjct: 666 ALSRLRGQPKE--SEYIQVELAEIVANEEYER 755
>TC87421 sugar transporter
Length = 1753
Score = 40.4 bits (93), Expect = 0.002
Identities = 75/348 (21%), Positives = 132/348 (37%), Gaps = 19/348 (5%)
Frame = +2
Query: 133 RFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHK 192
R LGFGIG +SE A K RGA GIL ++ A I
Sbjct: 446 RILLGFGIGFANQPVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKI---- 613
Query: 193 YKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAKNAKQ 252
+ + WRL L +PA + + +P+T +++ + +
Sbjct: 614 -----------------KGGWGWRLSLGGAMVPALIITIGSLVLPDTPN--SMIERGDRD 736
Query: 253 AAADMSKVLQVELEVEEEKVEKMTSDKRNSY------GLFSKQFAARHGLALFGTCSTWF 306
A K ++ +V+EE + + + + + L +++ + +A+ F
Sbjct: 737 GAKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQF 916
Query: 307 --LLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTV 364
+ I FY+ LF ++IG+ A M+A+ I ++A C ++ G
Sbjct: 917 TGINVIMFYAPVLF-----NSIGFKDDASLMSAV-----ITGVVNVVATCVSIYG----- 1051
Query: 365 AFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIG-------FVVIYSLTFFFA 417
+D GR A+ + G M + A+A G VV+ + + A
Sbjct: 1052--VDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVA 1225
Query: 418 NF----GPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFL 461
F GP ++VP+EIFP +RS ++ + + A FL
Sbjct: 1226GFAWSWGPLG--WLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFL 1363
>TC89865 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (33%)
Length = 617
Score = 39.7 bits (91), Expect = 0.003
Identities = 28/117 (23%), Positives = 50/117 (41%)
Frame = +1
Query: 69 QSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMAT 128
Q + +A+ G + G GW+ D+ GRKK I+ V + + ++P +
Sbjct: 217 QETIVSMAVAGAIVGAAVGGWMNDRYGRKK----ATIIADVIFILGAIVMAAAPDPYI-- 378
Query: 129 LCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTV 185
L R +G G+G + ++E + + RG +A M I GG ++ V
Sbjct: 379 LILGRVLVGLGVGIASVTAPVYIAELSPSEIRGGLVATNVLM----ITGGQFISYLV 537
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,512,244
Number of Sequences: 36976
Number of extensions: 207061
Number of successful extensions: 1569
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 1529
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1549
length of query: 533
length of database: 9,014,727
effective HSP length: 101
effective length of query: 432
effective length of database: 5,280,151
effective search space: 2281025232
effective search space used: 2281025232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC145767.6


