

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
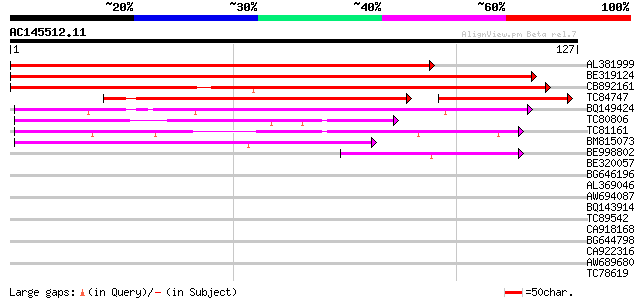
Query= AC145512.11 - phase: 0
(127 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AL381999 similar to GP|18033111|gb functional candidate resistan... 212 4e-56
BE319124 homologue to GP|18033111|gb functional candidate resist... 172 3e-44
CB892161 homologue to GP|18033111|gb functional candidate resist... 139 2e-34
TC84747 89 9e-30
BQ149424 similar to GP|12056928|g putative resistance protein {G... 62 6e-11
TC80806 similar to GP|18033111|gb|AAL56987.1 functional candidat... 40 2e-04
TC81161 similar to GP|23497565|gb|AAN37106.1 sortilin putative ... 39 5e-04
BM815073 weakly similar to GP|18033111|gb functional candidate r... 39 7e-04
BE998802 39 7e-04
BE320057 weakly similar to GP|18033111|gb functional candidate r... 35 0.010
BG646196 weakly similar to GP|12056928|gb putative resistance pr... 34 0.013
AL369046 weakly similar to GP|23497208|gb hypothetical protein {... 33 0.039
AW694087 weakly similar to GP|18033111|gb functional candidate r... 32 0.050
BQ143914 homologue to PIR|T39291|T392 hypothetical C2H2 zinc fin... 27 2.1
TC89542 homologue to PIR|F96805|F96805 unknown protein T5M16.20 ... 26 3.6
CA918168 similar to GP|12056928|gb putative resistance protein {... 26 3.6
BG644798 26 3.6
CA922316 similar to PIR|F96807|F968 unknown protein T32E8.10 [im... 26 3.6
AW689680 similar to PIR|G96770|G96 hypothetical protein F1O17.9 ... 26 4.7
TC78619 similar to GP|6552731|gb|AAF16530.1| T26F17.9 {Arabidops... 26 4.7
>AL381999 similar to GP|18033111|gb functional candidate resistance protein
KR1 {Glycine max}, partial (4%)
Length = 451
Score = 212 bits (539), Expect = 4e-56
Identities = 95/95 (100%), Positives = 95/95 (100%)
Frame = +3
Query: 1 MSVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLH 60
MSVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLH
Sbjct: 165 MSVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLH 344
Query: 61 LFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMF 95
LFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMF
Sbjct: 345 LFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMF 449
>BE319124 homologue to GP|18033111|gb functional candidate resistance protein
KR1 {Glycine max}, partial (1%)
Length = 522
Score = 172 bits (436), Expect = 3e-44
Identities = 82/118 (69%), Positives = 93/118 (78%)
Frame = +2
Query: 1 MSVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLH 60
+S SFWF NKFP+IAL VV + T + P RV+IN NTFFYTH KIDR SRPD YHLH
Sbjct: 41 LSFSFWFRNKFPAIALCVVCSSTLHDSQRPVRVVINGNTFFYTHDSKIDRSSRPDMYHLH 220
Query: 61 LFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNPEND 118
LFHMQ+E FN NMDKAL ENKWNHAE+DFG F+ SGIHVLKE+S+M DIRF NP+ D
Sbjct: 221 LFHMQMENFNENMDKALSENKWNHAELDFGLSFLESGIHVLKERSSMNDIRFINPKYD 394
>CB892161 homologue to GP|18033111|gb functional candidate resistance protein
KR1 {Glycine max}, partial (2%)
Length = 665
Score = 139 bits (351), Expect = 2e-34
Identities = 70/122 (57%), Positives = 87/122 (70%), Gaps = 1/122 (0%)
Frame = +3
Query: 1 MSVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSR-PDTYHL 59
+S+SFWF NKFP+I L VVS T + +V IN TFFY R ++ P ++HL
Sbjct: 24 LSISFWFRNKFPAIVLCVVSPLTRDNYQPNVKVFINGKTFFY---RDVEADYEWPISFHL 194
Query: 60 HLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNPENDA 119
H+FHMQ+E FN ++D ALLEN+WNH VDFGF F SGIHVLKEKS+M DI+FTNPEND
Sbjct: 195 HIFHMQIEKFNDDVDAALLENEWNHVVVDFGFEFHKSGIHVLKEKSSMMDIQFTNPENDV 374
Query: 120 NI 121
N+
Sbjct: 375 NM 380
>TC84747
Length = 920
Score = 88.6 bits (218), Expect(2) = 9e-30
Identities = 43/69 (62%), Positives = 48/69 (69%)
Frame = +2
Query: 22 YTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENK 81
+ +GY H RV IN N+F Y +I RPD YHLHLFHMQ E FN NMDKALLENK
Sbjct: 5 FWFGY--HGIRVSINGNSFIYKSNSEICWRQRPDMYHLHLFHMQTENFNDNMDKALLENK 178
Query: 82 WNHAEVDFG 90
WNHAE+ FG
Sbjct: 179WNHAEIYFG 205
Score = 57.0 bits (136), Expect(2) = 9e-30
Identities = 25/30 (83%), Positives = 28/30 (93%)
Frame = +3
Query: 97 GIHVLKEKSNMKDIRFTNPENDANIVLHSG 126
G+HVLKEK+ M+DIRFTNP NDANIVLHSG
Sbjct: 228 GVHVLKEKNXMEDIRFTNPANDANIVLHSG 317
>BQ149424 similar to GP|12056928|g putative resistance protein {Glycine max},
partial (14%)
Length = 625
Score = 62.0 bits (149), Expect = 6e-11
Identities = 45/130 (34%), Positives = 65/130 (49%), Gaps = 14/130 (10%)
Frame = +2
Query: 2 SVSFWFCNKFPSIAL---GVVSAYTWGYLEHPARVIINDNTF--FYTHGRKIDRCSRPDT 56
S+SFWF KFP+++L G++ G+ P VIIN N + D T
Sbjct: 191 SISFWFRGKFPALSLCFVGLMHKIPTGF--RPI-VIINGNIMKTMLPAEKWFDFEFPVLT 361
Query: 57 YHLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFS---------GIHVLKEKSNM 107
H+ + + F NMD+ L EN+WNHA + F +S G+HV+K+K +M
Sbjct: 362 DHIFIVGERHIKFEDNMDEVLSENEWNHAVISIDIDFKWSSSGLFAAWIGLHVIKQKCSM 541
Query: 108 KDIRFTNPEN 117
I+FTNP N
Sbjct: 542 DRIQFTNPCN 571
>TC80806 similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (3%)
Length = 431
Score = 40.0 bits (92), Expect = 2e-04
Identities = 29/93 (31%), Positives = 43/93 (46%), Gaps = 7/93 (7%)
Frame = +1
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTY---H 58
S+SFWFC K PSI ++ YL N F H + + Y H
Sbjct: 172 SISFWFCKKIPSITFIIILPNVEYYL--------GFNLFVKGHECTVQKFEFRLPYFSGH 327
Query: 59 LHLFHM----QVEYFNGNMDKALLENKWNHAEV 87
+LF M +E++ +DKALL+N+W + E+
Sbjct: 328 TYLFDMNLEENIEHYK-ILDKALLKNEWIYVEL 423
>TC81161 similar to GP|23497565|gb|AAN37106.1 sortilin putative {Plasmodium
falciparum 3D7}, partial (1%)
Length = 585
Score = 38.9 bits (89), Expect = 5e-04
Identities = 37/144 (25%), Positives = 59/144 (40%), Gaps = 30/144 (20%)
Frame = +2
Query: 2 SVSFWFCNKFPSIALG-----------VVSAYTWGYLEHPA-------RVIINDNTFFYT 43
++SFWF + PSI+ + + GY + + V+++++TF
Sbjct: 137 TISFWFDKELPSISFSFILICQLYIYPTIKLFVDGYEKEISCKFTGEFSVLVDNDTFL-- 310
Query: 44 HGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDFG-----------FP 92
H L H+++E N + LL+N+W H E G F
Sbjct: 311 ------------NDHTTLLHIKLEEDN-ELGVRLLKNEWIHVEFKLGCCYSGFDEEKIFE 451
Query: 93 FMFSGIHVLKEKSNMK-DIRFTNP 115
GIHV KEKSN + +RF +P
Sbjct: 452 NTQIGIHVWKEKSNTEGGVRFIDP 523
>BM815073 weakly similar to GP|18033111|gb functional candidate resistance
protein KR1 {Glycine max}, partial (4%)
Length = 516
Score = 38.5 bits (88), Expect = 7e-04
Identities = 27/88 (30%), Positives = 42/88 (47%), Gaps = 7/88 (7%)
Frame = +2
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCS-------RP 54
S+SFWF +FPSIAL V S T + + + + YT G + D S +P
Sbjct: 251 SISFWFRGRFPSIALFVASLSTDNRHPNSDFLSLTAHLRTYTDGIEFDINSVDLNLVIQP 430
Query: 55 DTYHLHLFHMQVEYFNGNMDKALLENKW 82
D +L+ QV +++K L ++W
Sbjct: 431 DHTYLYDLRQQVMELESDLEKTDLIDEW 514
>BE998802
Length = 608
Score = 38.5 bits (88), Expect = 7e-04
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 10/51 (19%)
Frame = +3
Query: 75 KALLENKWNHAEVDFGFPF----------MFSGIHVLKEKSNMKDIRFTNP 115
K +L+ +W HAE+ F F GIH+ K+ +NM+DI+FT+P
Sbjct: 276 KDILKTEWIHAEIRFEFKSNENEVMKSLSTLVGIHMFKKDNNMEDIKFTDP 428
>BE320057 weakly similar to GP|18033111|gb functional candidate resistance
protein KR1 {Glycine max}, partial (8%)
Length = 577
Score = 34.7 bits (78), Expect = 0.010
Identities = 25/81 (30%), Positives = 38/81 (46%), Gaps = 7/81 (8%)
Frame = +2
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCS-------RP 54
S+SFWF +FPSIAL V S T + + + + YT G + D S +P
Sbjct: 269 SISFWFRGRFPSIALFVASLSTDNRHPNSDFLSLTAHLRTYTDGIEFDINSVDLNLVIQP 448
Query: 55 DTYHLHLFHMQVEYFNGNMDK 75
D +L+ QV +++K
Sbjct: 449 DHTYLYDLRQQVMELESDLEK 511
>BG646196 weakly similar to GP|12056928|gb putative resistance protein
{Glycine max}, partial (3%)
Length = 447
Score = 34.3 bits (77), Expect = 0.013
Identities = 13/18 (72%), Positives = 17/18 (94%)
Frame = +2
Query: 2 SVSFWFCNKFPSIALGVV 19
S+SFWFCNKFP+I+L +V
Sbjct: 311 SMSFWFCNKFPAISLCLV 364
>AL369046 weakly similar to GP|23497208|gb hypothetical protein {Plasmodium
falciparum 3D7}, partial (1%)
Length = 486
Score = 32.7 bits (73), Expect = 0.039
Identities = 13/19 (68%), Positives = 16/19 (83%)
Frame = -2
Query: 97 GIHVLKEKSNMKDIRFTNP 115
GIH KE++NM DI+FTNP
Sbjct: 473 GIHFYKEQNNMDDIQFTNP 417
>AW694087 weakly similar to GP|18033111|gb functional candidate resistance
protein KR1 {Glycine max}, partial (4%)
Length = 655
Score = 32.3 bits (72), Expect = 0.050
Identities = 26/97 (26%), Positives = 39/97 (39%), Gaps = 11/97 (11%)
Frame = -2
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIIN-------DNTFFYT-HGRKIDRCSR 53
++SFWF K PSI ++ Y+ H + +N D F+Y H +
Sbjct: 396 TISFWFRKKIPSITCIIIVP---DYVVHEKFLFLNGKEITLTDRLFYYVDHDDDVVIWGH 226
Query: 54 PDTYHLHLFHMQVEYFNGNMD---KALLENKWNHAEV 87
+ L L E F D +A N+WNH E+
Sbjct: 225 AFLFDLKLEQRINESFANETDELYEAFKNNEWNHVEL 115
>BQ143914 homologue to PIR|T39291|T392 hypothetical C2H2 zinc finger protein
- fission yeast (Schizosaccharomyces pombe), partial
(1%)
Length = 849
Score = 26.9 bits (58), Expect = 2.1
Identities = 11/40 (27%), Positives = 23/40 (57%)
Frame = +2
Query: 3 VSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFY 42
+SF FPS+A ++ +++ YL +P + + + FF+
Sbjct: 725 LSFHLSLFFPSLAAHIIHSFSSSYLHYPFLALPSPSHFFF 844
>TC89542 homologue to PIR|F96805|F96805 unknown protein T5M16.20 [imported]
- Arabidopsis thaliana, partial (58%)
Length = 854
Score = 26.2 bits (56), Expect = 3.6
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = +2
Query: 77 LLENKWNHAEVDFGFPFMFSGIHVL 101
++ NKW ++DF FP S IH +
Sbjct: 308 IIMNKWIFQKLDFKFPLSVSCIHFI 382
>CA918168 similar to GP|12056928|gb putative resistance protein {Glycine
max}, partial (4%)
Length = 762
Score = 26.2 bits (56), Expect = 3.6
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +3
Query: 2 SVSFWFCNKFPSIALGVVSA 21
S+SFWF K PSI ++S+
Sbjct: 576 SISFWFRKKIPSINFIIISS 635
>BG644798
Length = 707
Score = 26.2 bits (56), Expect = 3.6
Identities = 10/14 (71%), Positives = 13/14 (92%)
Frame = -2
Query: 102 KEKSNMKDIRFTNP 115
K+KS+M DIRFT+P
Sbjct: 604 KQKSSMNDIRFTDP 563
>CA922316 similar to PIR|F96807|F968 unknown protein T32E8.10 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 746
Score = 26.2 bits (56), Expect = 3.6
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +2
Query: 91 FPFMFSGIHVLKEKSNMKDIRFTNPENDAN 120
FP + GIHV +E+ N+K +PEN ++
Sbjct: 485 FPTVSCGIHVQREQQNLK---MQHPENQSH 565
>AW689680 similar to PIR|G96770|G96 hypothetical protein F1O17.9 [imported] -
Arabidopsis thaliana, partial (43%)
Length = 687
Score = 25.8 bits (55), Expect = 4.7
Identities = 18/52 (34%), Positives = 23/52 (43%), Gaps = 4/52 (7%)
Frame = -1
Query: 16 LGVVSAYTWGYLEHPARVIINDNTFFY----THGRKIDRCSRPDTYHLHLFH 63
L V AY +EHP +IIN T FY H + C+ H L+H
Sbjct: 636 LNPVFAY*AACMEHPFIIIINFCTRFYCNAILHCSRTPGCNCALNLHALLYH 481
>TC78619 similar to GP|6552731|gb|AAF16530.1| T26F17.9 {Arabidopsis
thaliana}, complete
Length = 1428
Score = 25.8 bits (55), Expect = 4.7
Identities = 9/25 (36%), Positives = 15/25 (60%)
Frame = +3
Query: 77 LLENKWNHAEVDFGFPFMFSGIHVL 101
++ NKW ++DF FP S +H +
Sbjct: 201 IIVNKWIFQKLDFKFPLSVSCVHFI 275
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.140 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,303,735
Number of Sequences: 36976
Number of extensions: 82350
Number of successful extensions: 528
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 520
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 520
length of query: 127
length of database: 9,014,727
effective HSP length: 85
effective length of query: 42
effective length of database: 5,871,767
effective search space: 246614214
effective search space used: 246614214
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC145512.11


