

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145331.12 - phase: 0
(419 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
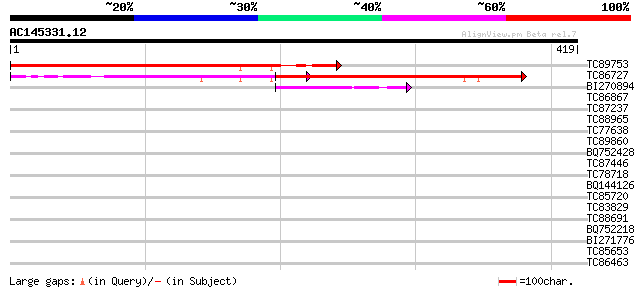
Score E
Sequences producing significant alignments: (bits) Value
TC89753 similar to GP|14334556|gb|AAK59686.1 unknown protein {Ar... 370 e-103
TC86727 similar to GP|4432859|gb|AAD20707.1| unknown protein {Ar... 152 2e-37
BI270894 similar to GP|16209720|gb| At2g35880/F11F19.21 {Arabido... 53 2e-07
TC86867 similar to GP|9294199|dbj|BAB02101.1 gb|AAF04893.1~gene_... 38 0.008
TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-inf... 32 0.46
TC88965 32 0.60
TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protei... 31 0.78
TC89860 similar to PIR|T05209|T05209 hypothetical protein F24J7.... 31 1.0
BQ752428 similar to GP|18376167|emb hypothetical protein {Neuros... 31 1.0
TC87446 similar to GP|19423979|gb|AAL87320.1 unknown protein {Ar... 31 1.0
TC78718 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtM... 30 1.7
BQ144126 29 3.0
TC85720 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pi... 29 3.0
TC83829 similar to GP|18377855|gb|AAL67114.1 AT4g28140/F26K10_20... 29 3.9
TC88691 similar to GP|20259659|gb|AAM14347.1 unknown protein {Ar... 29 3.9
BQ752218 similar to GP|19170914|emb hypothetical protein {Enceph... 28 5.0
BI271776 similar to GP|15724218|gb At2g27920/T1E2.16 {Arabidopsi... 28 5.0
TC85653 weakly similar to PIR|S44261|S44261 SRG1 protein - Arabi... 28 5.0
TC86463 similar to GP|21553582|gb|AAM62675.1 50S ribosomal prote... 28 6.6
>TC89753 similar to GP|14334556|gb|AAK59686.1 unknown protein {Arabidopsis
thaliana}, partial (6%)
Length = 1062
Score = 370 bits (951), Expect = e-103
Identities = 212/277 (76%), Positives = 213/277 (76%), Gaps = 32/277 (11%)
Frame = +1
Query: 1 MDPVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLE 60
MDPVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLE
Sbjct: 202 MDPVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLE 381
Query: 61 STATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSG 120
STATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSG
Sbjct: 382 STATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSG 561
Query: 121 VNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLS----------- 169
VNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLS
Sbjct: 562 VNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKQPSKSEAASS 741
Query: 170 -----KKKPKSLKKGPLDKVQGEGESSL----------------YPFSLRKGESSSLRLR 208
KKKPKSLKKGPLDKVQGEGESSL Y FS R GE
Sbjct: 742 DVAVEKKKPKSLKKGPLDKVQGEGESSLTNREDTKPRRVGTLPNYGFSFRCGE------- 900
Query: 209 KRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQL 245
R +RR V N V P L SL R L K L
Sbjct: 901 ---RAEKRREV----NSSYVLFPPLVSLSRFHLNKLL 990
>TC86727 similar to GP|4432859|gb|AAD20707.1| unknown protein {Arabidopsis
thaliana}, partial (27%)
Length = 1852
Score = 152 bits (384), Expect = 2e-37
Identities = 107/202 (52%), Positives = 128/202 (62%), Gaps = 16/202 (7%)
Frame = +3
Query: 197 LRKGESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQLHYQLSIRSLLL 256
LR+ +S +L LRK F +R+ RR KQ PRR KK RL+ LG+ * LKQL Q S RSLLL
Sbjct: 819 LREEKSFTLSLRKGFMRRKWRRATFKQKPRRAKKLRLRGLGKN*HLKQLQCQASTRSLLL 998
Query: 257 LRWS*RRYQLLEQNLPSSVGRRLQQI-QNLMETVAAVLDKVV*V*MRKCHRATPLQELPL 315
L WS*RRYQL EQNLPS V RR+QQ Q+ M TV AVL+ V*V MRKC R PL+EL +
Sbjct: 999 LGWS*RRYQLQEQNLPSLVVRRVQQCPQS*M*TVTAVLNNAV*VWMRKCLRIIPLKELVM 1178
Query: 316 PIKRSRYGSPFLLDSLLKG---PIQQLH-LHLRQ-----------QRKILAYPKVQGKRK 360
+RS LLD LLKG PIQ LH LHLR+ Q K+ P QGKRK
Sbjct: 1179 FSQRSHKEDLSLLD*LLKGSVHPIQ*LHGLHLRRCMTRKPPYLV*QLKLPPCPLQQGKRK 1358
Query: 361 LRLSQPMKKTVPCQVTQMLLYL 382
L+ Q +++TVP + Q+ L L
Sbjct: 1359 LKQLQLLRRTVPSLMRQVELRL 1424
Score = 126 bits (317), Expect = 1e-29
Identities = 93/258 (36%), Positives = 136/258 (52%), Gaps = 35/258 (13%)
Frame = +2
Query: 1 MDPVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGN-SENINQL 59
MDPVN+ DGL HQ + NS +DAV ++ VT +T + +GN ++N++
Sbjct: 164 MDPVNTSLEDGL---HQQQLF----NSQQDAVDFNV---VTQIKQTVLSNGNFNDNVSME 313
Query: 60 ESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSS 119
E +E EGSN ++G+N+ VSKE E++I TE+SR +K VKN N+K+ S
Sbjct: 314 E---------EEEEGSNGKIEGNNVNVSKEVEIEIVDETEKSRTKKDLVKNNNSKLPSPR 466
Query: 120 GVNASLVKNSKIGKDKQASPA--VSNGTSALDSRPRQPIKNRSSNDRQSQLS-------- 169
G+ + V+ +K GKD++A+ A VSNGTS DS PRQP+ NR+ ND+Q+ LS
Sbjct: 467 GLRMTSVRKNKDGKDEEAAVASSVSNGTSTFDSHPRQPVNNRAVNDKQTHLSKHSGKTDA 646
Query: 170 --------KKKPKSLKKGPLDKVQGEGESSL----------------YPFSLRKGESSSL 205
K +P +KK PLD + G+ ESS Y FS + E +
Sbjct: 647 ASTEAPMEKTRPHLIKKEPLDNLPGKAESSFPTSEDAKPRRVGTMPTYGFSFKCNERA-- 820
Query: 206 RLRKRFRQRRRRRVACKQ 223
RK F + R+ K+
Sbjct: 821 ERRKEFYSKLEERIHAKE 874
>BI270894 similar to GP|16209720|gb| At2g35880/F11F19.21 {Arabidopsis
thaliana}, partial (21%)
Length = 652
Score = 53.1 bits (126), Expect = 2e-07
Identities = 38/101 (37%), Positives = 61/101 (59%)
Frame = +3
Query: 197 LRKGESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQLHYQLSIRSLLL 256
L+ ESSS +++FR+R+ R V CK+N R+ K+ R ++ R * KQL Q+SIR+ LL
Sbjct: 171 LKSEESSSQS*KRKFRKRKLR*VICKRNQRKAKRQRSRN*ERE*HSKQLQCQVSIRN-LL 347
Query: 257 LRWS*RRYQLLEQNLPSSVGRRLQQIQNLMETVAAVLDKVV 297
+ S*+RY QN S L++ ++L+ T ++ V+
Sbjct: 348 QKLS*KRYLRHAQNHQS-----LEETKDLLLTTIQKINPVL 455
>TC86867 similar to GP|9294199|dbj|BAB02101.1
gb|AAF04893.1~gene_id:MXC7.13~similar to unknown protein
{Arabidopsis thaliana}, partial (38%)
Length = 1557
Score = 37.7 bits (86), Expect = 0.008
Identities = 29/76 (38%), Positives = 36/76 (47%)
Frame = +2
Query: 208 RKRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQLHYQLSIRSLLLLRWS*RRYQLL 267
RK +Q + RR+ KQ R+ K+ S G * LKQ Q L LR +*R Y
Sbjct: 818 RKSNKQWKLRRIKMKQGARKRKRKPSNS*GGV*SLKQALCQAFTMRGLHLRLT*RSYHPH 997
Query: 268 EQNLPSSVGRRLQQIQ 283
QNL S G R +Q
Sbjct: 998 VQNLQSWAGERATAVQ 1045
>TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-infected
erythrocyte surface antigen {Plasmodium falciparum},
partial (2%)
Length = 2007
Score = 32.0 bits (71), Expect = 0.46
Identities = 40/204 (19%), Positives = 75/204 (36%), Gaps = 25/204 (12%)
Frame = +1
Query: 26 NSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEI-----EGSNDNVD 80
N G + + ++ + + NS++ ES G+ A +++ + S +
Sbjct: 301 NEGGEKETGEITEEKSKQENEETSETNSKDKENEESNQNGSDAKEQVGENHEQDSKQGTE 480
Query: 81 GSNLTVSKEKEV--KIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV------------ 126
+N T EKE KIK T + KN A+ + SG NAS
Sbjct: 481 ETNGTEGGEKEEHDKIKEDTSSDNQVQDGEKNNEAREENYSGDNASSAVVDNKSQESSNK 660
Query: 127 ------KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGP 180
K K + ++ + T + DS Q + S Q+Q PK
Sbjct: 661 TEEQFDKKEKNEFELESQKNSNETTESTDSTITQNSQGNESEKDQAQTENDTPKGSASES 840
Query: 181 LDKVQGEGESSLYPFSLRKGESSS 204
++ Q + +++ ++ ++SS
Sbjct: 841 DEQKQEQEQNNTTKDDVQTTDTSS 912
Score = 28.1 bits (61), Expect = 6.6
Identities = 28/145 (19%), Positives = 59/145 (40%), Gaps = 8/145 (5%)
Frame = +1
Query: 10 DGLDDVHQNGVHDEPSNSGEDA----VSNDLDPHVTVNTETFVPDGNSENINQLESTATG 65
D + N DE +N +D+ S++ NTE + +++ N + +G
Sbjct: 1309 DNANSGQSNTTSDESANDNKDSSQVTTSSENSAEGNSNTENNSDENQNDSKNNENTNDSG 1488
Query: 66 NSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSS----GV 121
N++ N N + + T + E E + + +S+ + +K+ S+S
Sbjct: 1489 NTSNDANVNENQNENAAQ-TKTSENEGDAQNESVESKKENNESAHKDVDNNSNSNDQGSS 1665
Query: 122 NASLVKNSKIGKDKQASPAVSNGTS 146
+ S+ ++ K + + SNG S
Sbjct: 1666 DTSVTQDDKESRVDLGTLPESNGES 1740
>TC88965
Length = 1595
Score = 31.6 bits (70), Expect = 0.60
Identities = 29/128 (22%), Positives = 56/128 (43%), Gaps = 3/128 (2%)
Frame = +2
Query: 99 EQSRAQKGPVKNKNAKVG---SSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQP 155
++S A P +N+ +V S++ V S N + +ASP V + P +
Sbjct: 644 QKSLASSPPTENEAPEVRVTKSTADVKISDNPNDHV----RASPLVKPAKHVRPNDPAKA 811
Query: 156 IKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRR 215
+ R +DRQ +++ K+ K +K + + + ++ S + E + +K F RR
Sbjct: 812 GRKRGPSDRQEKIAAKRLKKIK--DIKALAADKTAANQKTSEARREDKAASSQKTFEARR 985
Query: 216 RRRVACKQ 223
+ A Q
Sbjct: 986 EDKAASSQ 1009
>TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protein {Oryza
sativa (japonica cultivar-group)}, partial (62%)
Length = 2345
Score = 31.2 bits (69), Expect = 0.78
Identities = 35/155 (22%), Positives = 61/155 (38%), Gaps = 4/155 (2%)
Frame = +1
Query: 13 DDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEI 72
D+ + D P + EDA D + V++E + ++E + E T T E
Sbjct: 514 DNSNARQFEDNPGDLPEDATKGDSN----VSSEEKSEENSTEKSS--EDTKT------ED 657
Query: 73 EGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKN---- 128
EG +GSN +K+ E +E ++K +N K S S N
Sbjct: 658 EGKKTEDEGSNTENNKDGEEASTKESESDESEKKDESEENNKSDSDESEKKSSDSNETTD 837
Query: 129 SKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSND 163
S + + + S + +A + K++SSN+
Sbjct: 838 SNVEEKVEQSQNKESDENASEKNTDDNAKDQSSNE 942
>TC89860 similar to PIR|T05209|T05209 hypothetical protein F24J7.50 -
Arabidopsis thaliana, partial (16%)
Length = 1139
Score = 30.8 bits (68), Expect = 1.0
Identities = 27/96 (28%), Positives = 43/96 (44%), Gaps = 2/96 (2%)
Frame = +2
Query: 4 VNSL-PADGLDDVHQNGVHDEPSNSGED-AVSNDLDPHVTVNTETFVPDGNSENINQLES 61
+N+L P D L NGV ++ SNS D A +ND P G S +E
Sbjct: 251 INTLFPLDALSSGDPNGVEEDNSNSYHDLATNNDARP--------MADTGQSNAEQNVEQ 406
Query: 62 TATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVS 97
T + + + K G + +++ ++S EK+ K S
Sbjct: 407 TDSTDESKKPNRGHSKSLE----SISTEKDQKKSAS 502
>BQ752428 similar to GP|18376167|emb hypothetical protein {Neurospora
crassa}, partial (3%)
Length = 690
Score = 30.8 bits (68), Expect = 1.0
Identities = 39/159 (24%), Positives = 67/159 (41%), Gaps = 12/159 (7%)
Frame = +2
Query: 92 VKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVK-----NSKIGK----DKQASPAVS 142
+++++ E RAQ VG S + ++++ NS + D P +S
Sbjct: 110 LRVQLFLELRRAQSRTTAGHLCVVGYSPQSSVAMLRWPLHDNSPLSDPHLYDHSFFPNMS 289
Query: 143 NGTSALDSRPR---QPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRK 199
+ + A SR R +P N S+N +SQ + ++L K P ++ P RK
Sbjct: 290 DPSLARCSRGRYT*RPASN-SNNRPKSQRCPRTAQNLPKSPSPRMPSPNPRRSPPRRSRK 466
Query: 200 GESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKSLGR 238
E + RK R R +R RR+ +PR ++ R
Sbjct: 467 RERRPMARRKARRATRTKR-------RRICRPRRLTVTR 562
>TC87446 similar to GP|19423979|gb|AAL87320.1 unknown protein {Arabidopsis
thaliana}, partial (18%)
Length = 1812
Score = 30.8 bits (68), Expect = 1.0
Identities = 19/51 (37%), Positives = 25/51 (48%), Gaps = 1/51 (1%)
Frame = -3
Query: 81 GSNLTVSKEKEVKIKVS-TEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSK 130
GSNL V + V S +KGP K+ A VG + G++ L NSK
Sbjct: 949 GSNLLVPNLELVDASESMVSNGPLEKGPTKSPTAFVGEAKGLSGRLCSNSK 797
>TC78718 similar to SP|O80961|ML12_ARATH MLO-like protein 12 (AtMlo12).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (42%)
Length = 1625
Score = 30.0 bits (66), Expect = 1.7
Identities = 17/71 (23%), Positives = 35/71 (48%)
Frame = +2
Query: 24 PSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGSN 83
P +S + ++ + P ++ +TF GNS+++ T+ + ++EG +N
Sbjct: 1019 PYSSRQSTPTHGMSPVHLLHRQTF---GNSDSLQTSPRTSNYENEQWDVEGGGSTSPRNN 1189
Query: 84 LTVSKEKEVKI 94
TV+ E E+ I
Sbjct: 1190 QTVASEIEIPI 1222
>BQ144126
Length = 698
Score = 29.3 bits (64), Expect = 3.0
Identities = 31/94 (32%), Positives = 46/94 (47%), Gaps = 3/94 (3%)
Frame = +2
Query: 1 MDPVNSLPADGLDDVHQNGVHDEPSNSGEDAVS-NDLDPHVTVNTETFVPDGNSENINQL 59
MDP+N+ DGL + + NS +DA ND VT +T + +GN +
Sbjct: 188 MDPLNTSLEDGLHQL-------QLFNSQQDADDFND----VTQIKQTGLTNGNLND---- 322
Query: 60 ESTATGNSAMKEIEG--SNDNVDGSNLTVSKEKE 91
N++M+E E SN + G+N+ VSKE E
Sbjct: 323 ------NASMEEEEK**SNGTI*GNNVNVSKELE 406
>TC85720 similar to SP|P08283|H1_PEA Histone H1. [Garden pea] {Pisum
sativum}, partial (73%)
Length = 968
Score = 29.3 bits (64), Expect = 3.0
Identities = 25/72 (34%), Positives = 35/72 (47%), Gaps = 11/72 (15%)
Frame = +1
Query: 196 SLRKGESSSLRLRKRFRQRRRRRVACKQNPRRVKK-----------PRLKSLGRA*LLKQ 244
SL + + S LR+R QRR+ Q R++ + PRLK L R LL++
Sbjct: 292 SLLRLKVHSSFLRRRNHQRRKSLQPLSQRQRQLLR*RQ*RPSPALSPRLKLLSRLRLLQK 471
Query: 245 LHYQLSIRSLLL 256
L+ L SLLL
Sbjct: 472 LNLLLQSPSLLL 507
>TC83829 similar to GP|18377855|gb|AAL67114.1 AT4g28140/F26K10_20
{Arabidopsis thaliana}, partial (9%)
Length = 1054
Score = 28.9 bits (63), Expect = 3.9
Identities = 14/44 (31%), Positives = 20/44 (44%)
Frame = +1
Query: 163 DRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLR 206
D S +S +P K P + Q + ++ PFS SSS R
Sbjct: 610 DEASNMSHNRPYKKSKSPQRENQNQNQNQFQPFSFPSSASSSSR 741
>TC88691 similar to GP|20259659|gb|AAM14347.1 unknown protein {Arabidopsis
thaliana}, partial (14%)
Length = 746
Score = 28.9 bits (63), Expect = 3.9
Identities = 32/123 (26%), Positives = 48/123 (39%)
Frame = +2
Query: 88 KEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSA 147
KEK + S S NK+ +SS N L + + + +P VSN + A
Sbjct: 230 KEKRFGHRFSVANSDQLSTDASNKDIGSTASSSANDQLTSTNP-AQAESPTPEVSNVSVA 406
Query: 148 LDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRL 207
S QP+ RSS + ++ + + K P G S L F ES +
Sbjct: 407 --SPKSQPVTTRSSQKVRERIRAARVLNQSKEPKTAKSEMGSSVLAAFK----ESDRGKK 568
Query: 208 RKR 210
R+R
Sbjct: 569 RRR 577
>BQ752218 similar to GP|19170914|emb hypothetical protein {Encephalitozoon
cuniculi}, partial (5%)
Length = 637
Score = 28.5 bits (62), Expect = 5.0
Identities = 13/28 (46%), Positives = 21/28 (74%)
Frame = -3
Query: 206 RLRKRFRQRRRRRVACKQNPRRVKKPRL 233
RL++R R+RRRR K+ PRR+++ R+
Sbjct: 461 RLKRRSRRRRRRP---KRRPRRIRRSRI 387
>BI271776 similar to GP|15724218|gb At2g27920/T1E2.16 {Arabidopsis thaliana},
partial (40%)
Length = 610
Score = 28.5 bits (62), Expect = 5.0
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = -3
Query: 163 DRQSQLSKKKPKSLKKGPLDKVQGEGESSLYP 194
D S LS SL++GP D + GE L+P
Sbjct: 545 DFSSPLSSSLETSLRRGPHDNTKSSGEIQLFP 450
>TC85653 weakly similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis
thaliana, partial (20%)
Length = 660
Score = 28.5 bits (62), Expect = 5.0
Identities = 19/53 (35%), Positives = 27/53 (50%), Gaps = 4/53 (7%)
Frame = +2
Query: 84 LTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGS----SSGVNASLVKNSKIG 132
+T K K + + +Q +A KG KN K S + GVN SL++ KIG
Sbjct: 71 VTGQKAKMARERNLEKQKQAGKGSQLEKNKKAMSIQIINHGVNTSLIEKVKIG 229
>TC86463 similar to GP|21553582|gb|AAM62675.1 50S ribosomal protein L27
{Arabidopsis thaliana}, partial (58%)
Length = 950
Score = 28.1 bits (61), Expect = 6.6
Identities = 19/45 (42%), Positives = 25/45 (55%), Gaps = 2/45 (4%)
Frame = +3
Query: 191 SLYPFSLRKGESSSLRLRK--RFRQRRRRRVACKQNPRRVKKPRL 233
S+YP R S RLRK F+ RR R+ A K+ RR +P+L
Sbjct: 477 SVYPQEERVPNPDSYRLRKIKYFKMRRERKKAAKE--RRAFQPQL 605
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.137 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,143,443
Number of Sequences: 36976
Number of extensions: 144543
Number of successful extensions: 1229
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 1196
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1218
length of query: 419
length of database: 9,014,727
effective HSP length: 99
effective length of query: 320
effective length of database: 5,354,103
effective search space: 1713312960
effective search space used: 1713312960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC145331.12


