

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145331.1 - phase: 0
(520 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
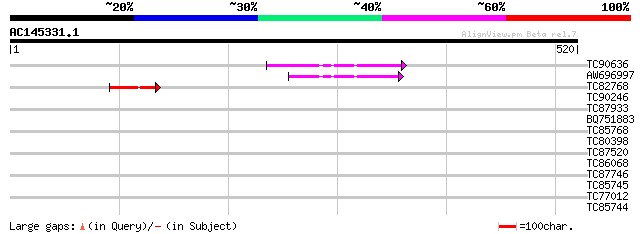
Score E
Sequences producing significant alignments: (bits) Value
TC90636 similar to GP|21593548|gb|AAM65515.1 unknown {Arabidopsi... 75 8e-14
AW696997 similar to GP|21593548|gb unknown {Arabidopsis thaliana... 55 7e-08
TC82768 similar to GP|21593548|gb|AAM65515.1 unknown {Arabidopsi... 44 2e-04
TC90246 similar to PIR|T00872|T00872 probable protein kinase At2... 31 1.3
TC87933 weakly similar to GP|5832707|dbj|BAA84071.1 cytochrome P... 30 1.7
BQ751883 homologue to GP|22762042|db Ste12-like transcription fa... 30 2.3
TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle pr... 29 5.0
TC80398 similar to GP|20521392|dbj|BAB91903. putative FtsH prote... 28 6.6
TC87520 similar to GP|9757998|dbj|BAB08420.1 cell division prote... 28 6.6
TC86068 similar to GP|12407658|gb|AAG53613.1 eukaryotic initiati... 28 6.6
TC87746 similar to GP|15021761|gb|AAK77908.1 AAA-metalloprotease... 28 8.6
TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome re... 28 8.6
TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulator... 28 8.6
TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15... 28 8.6
>TC90636 similar to GP|21593548|gb|AAM65515.1 unknown {Arabidopsis
thaliana}, partial (30%)
Length = 842
Score = 74.7 bits (182), Expect = 8e-14
Identities = 46/129 (35%), Positives = 67/129 (51%)
Frame = +1
Query: 236 IIVATSNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYL 295
I+V+TSNRAP +L E + + F ++ L+E C +GS DYR+ S + YL
Sbjct: 175 ILVSTSNRAPDNLYEGGLQRDLFLPFIATLKERCIAHEIGSSTDYRKMT---SGGQGFYL 345
Query: 296 WPIERETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRP 355
+ ++ +KK+Q G G + V+ GR L VP G A F F+ LC +P
Sbjct: 346 --VGSDSSGFLKKKFQQLIGE--GTPTPQEVEVVMGRKLHVPLGANGCAYFPFEELCDKP 513
Query: 356 LGAADYIAV 364
LGAADY +
Sbjct: 514 LGAADYFGL 540
>AW696997 similar to GP|21593548|gb unknown {Arabidopsis thaliana}, partial
(21%)
Length = 742
Score = 55.1 bits (131), Expect = 7e-08
Identities = 36/106 (33%), Positives = 51/106 (47%)
Frame = +2
Query: 256 EFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATG 315
+ F ++ L+E C +GS DYR+ S + YL + ++ +KK+Q G
Sbjct: 86 DLFLPFIATLKERCIAHEIGSSTDYRKMT---SGGQGFYL--VGSDSSGFLKKKFQQLIG 250
Query: 316 RFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADY 361
G + V+ GR L VP G A F F LC +PLGAADY
Sbjct: 251 E--GTPTPQEVEVVMGRELHVPLGANGCAYFPFKELCDKPLGAADY 382
>TC82768 similar to GP|21593548|gb|AAM65515.1 unknown {Arabidopsis
thaliana}, partial (23%)
Length = 780
Score = 43.9 bits (102), Expect = 2e-04
Identities = 22/48 (45%), Positives = 32/48 (65%), Gaps = 1/48 (2%)
Frame = +2
Query: 92 LYIY-GNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHK 138
+YI+ G VG+GKTMLMD+FY K ++R HFH+ ML ++ + K
Sbjct: 614 VYIFMGGVGTGKTMLMDLFYDQLPSNWK-KKRIHFHDFMLNVHSLLQK 754
>TC90246 similar to PIR|T00872|T00872 probable protein kinase At2g45590
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1451
Score = 30.8 bits (68), Expect = 1.3
Identities = 37/134 (27%), Positives = 57/134 (41%), Gaps = 17/134 (12%)
Frame = +2
Query: 12 EKKRENERRRIL-----MDEV--------EKQQNDKDWWK--RLNNKITERWTNSRKRP- 55
EKK+E +R++ L MDE EK+++ ++WWK E+ N +K+
Sbjct: 272 EKKKEKKRKQKLEWWESMDEEIVGGMMRKEKRKSVREWWKEEHCEEVAKEKKKNKKKKKG 451
Query: 56 -ENVDPGVGKWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATE 114
EN D G+ ++ + D+L G R KG GN SG +D +
Sbjct: 452 VENGDDGISNGDNWWMSD---DALYGDR------KKGKRRSGNNRSGN---IDWWLDGFS 595
Query: 115 GIVKHRRRYHFHEA 128
G + RR F A
Sbjct: 596 GDLWRARRNSFDSA 637
>TC87933 weakly similar to GP|5832707|dbj|BAA84071.1 cytochrome P450
{Antirrhinum majus}, partial (68%)
Length = 1614
Score = 30.4 bits (67), Expect = 1.7
Identities = 17/44 (38%), Positives = 25/44 (56%)
Frame = +1
Query: 66 VSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMF 109
+SYLKR KKL + R P PP+P + I G++ K ++ F
Sbjct: 94 LSYLKRNKKLHA---RLPPHPPSPPAIPIIGHLHLLKPLIHQAF 216
>BQ751883 homologue to GP|22762042|db Ste12-like transcription factor
{Colletotrichum lagenarium}, partial (23%)
Length = 652
Score = 30.0 bits (66), Expect = 2.3
Identities = 28/101 (27%), Positives = 45/101 (43%), Gaps = 8/101 (7%)
Frame = -3
Query: 190 ADKFFLDREGEEKGANILCFD-----EIQTVDVFAIVALSGILSRLLSSGTIIVATSNRA 244
+D FL +E EE +L EI V +A+ L G+L R + + +
Sbjct: 584 SDAVFLVKEVEEGALGLLERRISPGLEISQVGEYALFELLGVLHRAAEG----LESE*QT 417
Query: 245 PKDLNEANM---VPEFFQNLLSNLEEHCEKVLVGSEIDYRR 282
P D+ ++ VP+ N+ + +E VL+G ID RR
Sbjct: 416 PDDVRAGDVEEIVPQHAGNVFACWQEEPANVLIGLPIDRRR 294
>TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle protein 48
homolog (Valosin containing protein homolog) (VCP).
[Soybean], partial (95%)
Length = 2785
Score = 28.9 bits (63), Expect = 5.0
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = +2
Query: 71 REKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
R +L +G +P PKG+ +YG GSGKT++
Sbjct: 779 RHPQLFKSIGVKP-----PKGILLYGPPGSGKTLI 868
>TC80398 similar to GP|20521392|dbj|BAB91903. putative FtsH protease {Oryza
sativa (japonica cultivar-group)}, partial (26%)
Length = 981
Score = 28.5 bits (62), Expect = 6.6
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = +2
Query: 66 VSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
V YL+ K+ L G+ PKG+ + G G+GKTML
Sbjct: 878 VHYLRDPKRFTRLGGK------LPKGVLLVGPPGTGKTML 979
>TC87520 similar to GP|9757998|dbj|BAB08420.1 cell division protein FtsH
protease-like {Arabidopsis thaliana}, partial (53%)
Length = 1667
Score = 28.5 bits (62), Expect = 6.6
Identities = 16/40 (40%), Positives = 21/40 (52%)
Frame = +2
Query: 66 VSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
V YLK K L G+ PKG+ + G G+GKT+L
Sbjct: 26 VEYLKNPAKFTRLGGK------LPKGILLTGAPGTGKTLL 127
>TC86068 similar to GP|12407658|gb|AAG53613.1 eukaryotic initiation factor
3E subunit {Arabidopsis thaliana}, partial (98%)
Length = 1602
Score = 28.5 bits (62), Expect = 6.6
Identities = 24/101 (23%), Positives = 43/101 (41%), Gaps = 18/101 (17%)
Frame = +1
Query: 92 LYIYGNVGSGKTMLMDMF------------------YSATEGIVKHRRRYHFHEAMLRIN 133
L+I+ N +G+T ++D+F Y AT IV RRR F + +++
Sbjct: 730 LFIFFNHDNGRTQIIDLFNQDKYLNAIQTSAPHLLRYLATAFIVNKRRRPQFKD-FIKVI 906
Query: 134 EHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEE 174
+ ++K P+ ++ +N FD K+ EE
Sbjct: 907 QQEQNSYK-----DPITEFLACVYVNYDFDGAQKKMRECEE 1014
>TC87746 similar to GP|15021761|gb|AAK77908.1 AAA-metalloprotease FtsH {Pisum
sativum}, partial (49%)
Length = 1274
Score = 28.1 bits (61), Expect = 8.6
Identities = 16/50 (32%), Positives = 25/50 (50%)
Frame = +2
Query: 56 ENVDPGVGKWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
E + ++V +LK KK + L + PKG + G G+GKT+L
Sbjct: 1055 EEAKQEIMEFVHFLKNPKKYEELGAK------IPKGALLVGPPGTGKTLL 1186
>TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome regulatory
particle triple-A ATPase subunit2b {Oryza sativa
(japonica cultivar-group), partial (98%)
Length = 1696
Score = 28.1 bits (61), Expect = 8.6
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +3
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSAT 113
PKG+ +YG G+GKT+L ++T
Sbjct: 753 PKGVILYGEPGTGKTLLAKAVANST 827
>TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulatory subunit
6A homolog (TAT-binding protein homolog 1) (TBP-1)
(Mg(2+), complete
Length = 1793
Score = 28.1 bits (61), Expect = 8.6
Identities = 20/89 (22%), Positives = 43/89 (47%)
Frame = +1
Query: 17 NERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGKWVSYLKREKKLD 76
N+ +++D + + + + ++ K TE + + + + V V + +++
Sbjct: 544 NKDSYLILDTLPSEYDSRVKAMEVDEKPTEDYNDIGGLEKQIQELVEAIVLPMTHKERFQ 723
Query: 77 SLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
L G RP PKG+ +YG G+GKT++
Sbjct: 724 KL-GIRP-----PKGVLLYGPPGTGKTLM 792
>TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15.70 -
Arabidopsis thaliana, complete
Length = 1629
Score = 28.1 bits (61), Expect = 8.6
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +2
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSAT 113
PKG+ +YG G+GKT+L ++T
Sbjct: 755 PKGVILYGEPGTGKTLLAKAVANST 829
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,197,678
Number of Sequences: 36976
Number of extensions: 220335
Number of successful extensions: 1413
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1395
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1411
length of query: 520
length of database: 9,014,727
effective HSP length: 100
effective length of query: 420
effective length of database: 5,317,127
effective search space: 2233193340
effective search space used: 2233193340
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC145331.1


