

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
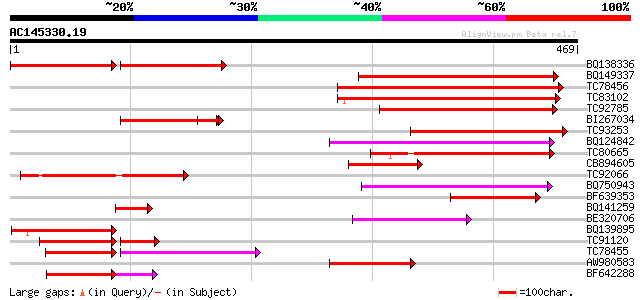
Query= AC145330.19 - phase: 0 /pseudo
(469 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BQ138336 similar to GP|17065360|gb Unknown protein {Arabidopsis ... 187 4e-93
BQ149337 weakly similar to GP|13272459|gb unknown protein {Arabi... 233 1e-61
TC78456 similar to GP|11994404|dbj|BAB02363. gb|AAC28507.1~gene_... 220 1e-57
TC83102 similar to GP|11994404|dbj|BAB02363. gb|AAC28507.1~gene_... 202 2e-52
TC92785 weakly similar to GP|13937234|gb|AAK50109.1 AT5g38030/F1... 172 2e-43
BI267034 weakly similar to PIR|T09556|T095 hypothetical protein ... 151 6e-37
TC93253 weakly similar to GP|14030731|gb|AAK53040.1 AT5g65380/MN... 149 2e-36
BQ124842 weakly similar to GP|19310633|gb unknown protein {Arabi... 146 1e-35
TC80665 weakly similar to GP|19310633|gb|AAL85047.1 unknown prot... 119 2e-27
CB894605 weakly similar to GP|22136880|gb| unknown protein {Arab... 110 9e-25
TC92066 similar to GP|12231296|gb|AAG49032.1 ripening regulated ... 97 1e-20
BQ750943 weakly similar to GP|21627817|emb hypothetical protein ... 79 3e-15
BF639353 similar to GP|17979043|gb| unknown protein {Arabidopsis... 79 5e-15
BQ141259 similar to GP|22136880|gb| unknown protein {Arabidopsis... 74 2e-13
BE320706 weakly similar to GP|23617058|db putative integral memb... 72 5e-13
BQ139895 similar to GP|12231296|gb| ripening regulated protein D... 68 7e-12
TC91120 similar to GP|11994404|dbj|BAB02363. gb|AAC28507.1~gene_... 49 2e-11
TC78455 similar to GP|11994404|dbj|BAB02363. gb|AAC28507.1~gene_... 41 1e-09
AW980583 weakly similar to GP|16797803|db hypothetical membrane ... 54 2e-07
BF642288 similar to GP|1495259|emb orf04 {Arabidopsis thaliana},... 52 3e-07
>BQ138336 similar to GP|17065360|gb Unknown protein {Arabidopsis thaliana},
partial (20%)
Length = 536
Score = 187 bits (475), Expect(2) = 4e-93
Identities = 84/88 (95%), Positives = 85/88 (96%)
Frame = +3
Query: 92 VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDTLRPR*K 151
+GNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDTLRPR*K
Sbjct: 273 LGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDTLRPR*K 452
Query: 152 HIRSCRKHFSLVNSYYVCFYCLIHLPNI 179
HI SCRKHFSLVNSYYV FY LIHLPNI
Sbjct: 453 HIXSCRKHFSLVNSYYVXFYXLIHLPNI 536
Score = 172 bits (437), Expect(2) = 4e-93
Identities = 87/88 (98%), Positives = 88/88 (99%)
Frame = +1
Query: 1 MERDLKQNLLLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAF 60
MERDLKQNLLLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAF
Sbjct: 13 MERDLKQNLLLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAF 192
Query: 61 VGHIGSRELAAFALVFTVLIRFANGILM 88
VGHIGSRELAAFALVFTVLIRFANGIL+
Sbjct: 193 VGHIGSRELAAFALVFTVLIRFANGILL 276
>BQ149337 weakly similar to GP|13272459|gb unknown protein {Arabidopsis
thaliana}, partial (21%)
Length = 622
Score = 233 bits (594), Expect = 1e-61
Identities = 111/166 (66%), Positives = 143/166 (85%)
Frame = +2
Query: 289 FMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEV 348
F AAASVRV+NELG+GS++ AKFSIV+ LTS AIG FL FLF +++L+YIFTS+ +V
Sbjct: 2 FFAAASVRVANELGRGSSRDAKFSIVINALTSFAIGFIFFLIFLFLKKKLSYIFTSDSDV 181
Query: 349 AAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNII 408
A AVG+LS L++S+LLNSVQPVLSGV++GAGWQS VAYVNIGCYY+IGIPVG+V+G +
Sbjct: 182 ANAVGDLSFWLALSMLLNSVQPVLSGVSVGAGWQSIVAYVNIGCYYLIGIPVGVVIGVVY 361
Query: 409 HWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVNRWS 454
+ +KGIW+GMLFGT +QTI+L+IIT KT+WD+QV +A+ RVNRW+
Sbjct: 362 NLGIKGIWIGMLFGTFVQTIMLIIITTKTDWDKQVEIAQWRVNRWA 499
>TC78456 similar to GP|11994404|dbj|BAB02363.
gb|AAC28507.1~gene_id:MIL23.25~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (40%)
Length = 920
Score = 220 bits (560), Expect = 1e-57
Identities = 104/188 (55%), Positives = 144/188 (76%), Gaps = 1/188 (0%)
Frame = +1
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC ++GW MIS+GF AAASVRVSNELG G++K+A FS+VV + S I + + L
Sbjct: 64 LSICTTVSGWTFMISVGFQAAASVRVSNELGAGNSKSASFSVVVVTVISFIICAIIALVV 243
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L R+ ++Y+FT +EVAAAV LSPLL+++I+LN VQPVLSGVA+G GWQ+ VAYVN+G
Sbjct: 244 LALRDVISYVFTEGEEVAAAVSNLSPLLALAIVLNGVQPVLSGVAVGCGWQTFVAYVNVG 423
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY +GIP+G VLG + KGIW+GML GT++QTI+L+ +T++T+W+ +V + KR+N
Sbjct: 424 CYYGLGIPLGAVLGFYFKFGAKGIWLGMLGGTVLQTIILMWVTFRTDWNNEVVESNKRLN 603
Query: 452 RW-SKVES 458
+W K ES
Sbjct: 604 KWEGKTES 627
>TC83102 similar to GP|11994404|dbj|BAB02363.
gb|AAC28507.1~gene_id:MIL23.25~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (36%)
Length = 1048
Score = 202 bits (515), Expect = 2e-52
Identities = 97/187 (51%), Positives = 136/187 (71%), Gaps = 3/187 (1%)
Frame = +3
Query: 272 VSIC---LNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLF 328
++IC ++GW MIS+GF AAASVRVSNELG + K+A FS+ V + S I
Sbjct: 192 ITICDCSTTVSGWVFMISVGFNAAASVRVSNELGARNPKSASFSVKVVTVISFIISVIAA 371
Query: 329 LFFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYV 388
L L R+ ++Y+FT + VAAAV +L PLLS+S++LN +QPVLSGVA+G GWQ+ VAYV
Sbjct: 372 LIVLALRDVISYVFTEGEVVAAAVSDLCPLLSLSLVLNGIQPVLSGVAVGCGWQAFVAYV 551
Query: 389 NIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARK 448
N+GCYYI GIP+G LG ++ KGIW+GML GT +QTI+L+ +T++T+W+++V A K
Sbjct: 552 NVGCYYIAGIPLGAGLGFYFNFGAKGIWLGMLGGTTMQTIILMWVTFRTDWNKEVKEAAK 731
Query: 449 RVNRWSK 455
R+N+W +
Sbjct: 732 RLNKWEE 752
>TC92785 weakly similar to GP|13937234|gb|AAK50109.1 AT5g38030/F16F17_30
{Arabidopsis thaliana}, partial (30%)
Length = 533
Score = 172 bits (436), Expect = 2e-43
Identities = 78/147 (53%), Positives = 112/147 (76%)
Frame = +1
Query: 307 KAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLN 366
+ A+FS+VV V+TS+ IG L L + R++ FT++KEV V +L+PLL++ +++N
Sbjct: 4 RTARFSLVVAVITSILIGLLLALVLIISRDKYPAYFTTDKEVQDLVKDLTPLLALCVVIN 183
Query: 367 SVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQ 426
+VQPVLSGVAIGAGWQ+ VAYVNI CYY+ GIPVG++LG ++ VKGIW GM+ GT++Q
Sbjct: 184 NVQPVLSGVAIGAGWQAAVAYVNIACYYLFGIPVGLILGYKVNLGVKGIWCGMMSGTILQ 363
Query: 427 TIVLLIITYKTNWDEQVTVARKRVNRW 453
T VLLI+ YKTNW+++ ++A R+ W
Sbjct: 364 TCVLLIMVYKTNWNKEASLAEDRIRNW 444
>BI267034 weakly similar to PIR|T09556|T095 hypothetical protein L73G19.20 -
Arabidopsis thaliana, partial (13%)
Length = 428
Score = 151 bits (381), Expect = 6e-37
Identities = 69/84 (82%), Positives = 73/84 (86%)
Frame = +1
Query: 92 VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDTLRPR*K 151
VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDTLRPR*K
Sbjct: 166 VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDTLRPR*K 345
Query: 152 HIRSCRKHFSLVNSYYVCFYCLIH 175
HIRS ++ F L + +C L H
Sbjct: 346 HIRSLQEAF-LFGQFLLCXLLLXH 414
Score = 46.6 bits (109), Expect = 2e-05
Identities = 20/22 (90%), Positives = 20/22 (90%)
Frame = +2
Query: 156 CRKHFSLVNSYYVCFYCLIHLP 177
CRKHFSLVNSYYV FY LIHLP
Sbjct: 359 CRKHFSLVNSYYVXFYXLIHLP 424
>TC93253 weakly similar to GP|14030731|gb|AAK53040.1 AT5g65380/MNA5_11
{Arabidopsis thaliana}, partial (25%)
Length = 633
Score = 149 bits (376), Expect = 2e-36
Identities = 68/131 (51%), Positives = 100/131 (75%), Gaps = 1/131 (0%)
Frame = +2
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
+ F + AYIFT++ V AV ++S LL+++ILLNSVQP+LSGVA+G+GWQ VAYVNIG
Sbjct: 2 MIFHRQFAYIFTTSPPVLEAVNDMSILLAVTILLNSVQPILSGVAVGSGWQVFVAYVNIG 181
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLF-GTLIQTIVLLIITYKTNWDEQVTVARKRV 450
CYY+IG+P+GI++G + + V+GIW GM+F GT IQT++L+I+T + +W+ + AR V
Sbjct: 182 CYYLIGLPLGILMGWVFNTGVEGIWGGMIFGGTAIQTLILIIVTARCDWENEAKKARSSV 361
Query: 451 NRWSKVESTDQ 461
++WS + DQ
Sbjct: 362 SKWSVTKPDDQ 394
>BQ124842 weakly similar to GP|19310633|gb unknown protein {Arabidopsis
thaliana}, partial (31%)
Length = 706
Score = 146 bits (369), Expect = 1e-35
Identities = 68/186 (36%), Positives = 110/186 (58%)
Frame = +2
Query: 265 PFSRRNVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIG 324
P +V+SICLN G MI G A S+RVSNELG G+ A ++ V + S
Sbjct: 149 PVIETSVLSICLNTFGLAWMIPFGCSCAVSIRVSNELGGGNPNGASLAVRVALSISFIAA 328
Query: 325 SFLFLFFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQST 384
F+ L + R+ ++++ +K+V V + P+L+IS L+++Q LSGV G GWQ
Sbjct: 329 LFMVLSMILARKVWGHLYSDDKQVIRYVSAMMPILAISSFLDAIQSTLSGVLAGCGWQKI 508
Query: 385 VAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVT 444
AYVN+G +Y++G+P +VL +H G+W+G++ ++QT + +I T ++NW+E+
Sbjct: 509 GAYVNLGSFYVVGVPCAVVLAFFVHMHAMGLWLGIISAFIVQTSLYIIFTIRSNWEEEAK 688
Query: 445 VARKRV 450
A+ RV
Sbjct: 689 KAQSRV 706
>TC80665 weakly similar to GP|19310633|gb|AAL85047.1 unknown protein
{Arabidopsis thaliana}, partial (31%)
Length = 894
Score = 119 bits (299), Expect = 2e-27
Identities = 56/156 (35%), Positives = 96/156 (60%), Gaps = 4/156 (2%)
Frame = +3
Query: 299 NELGKGSAKAAKFSI----VVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVGE 354
NELG G+ +AA+ ++ V+ ++ S+ +G+ + L R Y +++ +EV V
Sbjct: 3 NELGAGNPRAARLAVYVVVVIAIIESIVVGAVIILI----RNIWGYAYSNEEEVVKYVAI 170
Query: 355 LSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKG 414
+ P++++S L+ +Q VLSG A G GWQ AYVN+G YY++GIP +VL ++H KG
Sbjct: 171 MLPIIAVSNFLDGIQSVLSGTARGVGWQKIGAYVNLGSYYLVGIPAAVVLAFVLHVGGKG 350
Query: 415 IWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRV 450
+W+G++ +Q + L IIT +T+W+++ A RV
Sbjct: 351 LWLGIICALFVQVVSLTIITIRTDWEKEAKKATDRV 458
>CB894605 weakly similar to GP|22136880|gb| unknown protein {Arabidopsis
thaliana}, partial (12%)
Length = 183
Score = 110 bits (276), Expect = 9e-25
Identities = 57/61 (93%), Positives = 58/61 (94%)
Frame = +1
Query: 281 WEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAY 340
WEMMISLGFMAAASVRVSNELGKGS KAAKFSIVVT LTSL IGSFLFLFFLFFRER+AY
Sbjct: 1 WEMMISLGFMAAASVRVSNELGKGSEKAAKFSIVVTGLTSLGIGSFLFLFFLFFRERIAY 180
Query: 341 I 341
I
Sbjct: 181 I 183
>TC92066 similar to GP|12231296|gb|AAG49032.1 ripening regulated protein
DDTFR18 {Lycopersicon esculentum}, partial (34%)
Length = 668
Score = 97.1 bits (240), Expect = 1e-20
Identities = 52/139 (37%), Positives = 84/139 (60%)
Frame = +3
Query: 10 LLKKPEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSREL 69
LL + EEE E+++L+ ++VW E+K +W + PAIF+R +++ + +I+Q+F GH+G EL
Sbjct: 153 LLSRNEEEEEEQDLT--RKVWIESKKLWHIVGPAIFSRIASYMMLVITQSFAGHLGDLEL 326
Query: 70 AAFALVFTVLIRFANGILMKIVVGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNC 129
AA ++ V++ F G+L+ G C+ N +WTSIW K I + SI + IMDS I
Sbjct: 327 AAISIANNVVVGFDFGLLL----GMAKCITNFMWTSIWSKTILHVRSIHATIMDSTIHMS 494
Query: 130 TCSSSCVCLHNPDFDTLRP 148
S + + N T+ P
Sbjct: 495 DFSLTNLPFCNTSVKTIGP 551
>BQ750943 weakly similar to GP|21627817|emb hypothetical protein {Aspergillus
fumigatus}, partial (7%)
Length = 711
Score = 79.3 bits (194), Expect = 3e-15
Identities = 43/158 (27%), Positives = 77/158 (48%)
Frame = -1
Query: 292 AASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAA 351
AAS R++N +G G AAK + V +L +G F + F R L +F++++EV A
Sbjct: 684 AASTRIANLIGAGLVDAAKTTGKVAFGAALTVGVFNVILFGGLRFHLPRLFSNDEEVIAI 505
Query: 352 VGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQ 411
V + PL ++ + + + G+ G G S Y N+ YY + +P+ ++ W+
Sbjct: 504 VANVLPLCAVMQVFDGLAAGAHGLLRGIGRPSIGGYANLAVYYGVALPLSFSTAFLLDWK 325
Query: 412 VKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+KG+W G+ G + + + T+W + A R
Sbjct: 324 LKGLWFGVTVGLALVSAIEYAFVALTDWHKAAKEAETR 211
>BF639353 similar to GP|17979043|gb| unknown protein {Arabidopsis thaliana},
partial (16%)
Length = 551
Score = 78.6 bits (192), Expect = 5e-15
Identities = 35/75 (46%), Positives = 50/75 (66%)
Frame = +1
Query: 365 LNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTL 424
L+SVQ VLSGVA G GWQ YVN+ +Y+IG+P+ +LG + Q KG+W+G++ G +
Sbjct: 1 LDSVQGVLSGVARGCGWQHLTVYVNLATFYLIGLPISCLLGFKTNLQYKGLWIGLICGLV 180
Query: 425 IQTIVLLIITYKTNW 439
QT LL++T W
Sbjct: 181 CQTGALLLLTRHVKW 225
>BQ141259 similar to GP|22136880|gb| unknown protein {Arabidopsis thaliana},
partial (5%)
Length = 467
Score = 73.6 bits (179), Expect = 2e-13
Identities = 31/31 (100%), Positives = 31/31 (100%)
Frame = +2
Query: 88 MKIVVGNGDCVVNPLWTSIWCKRIWNDGSIS 118
MKIVVGNGDCVVNPLWTSIWCKRIWNDGSIS
Sbjct: 374 MKIVVGNGDCVVNPLWTSIWCKRIWNDGSIS 466
>BE320706 weakly similar to GP|23617058|db putative integral membrane protein
{Oryza sativa (japonica cultivar-group)}, partial (18%)
Length = 330
Score = 72.0 bits (175), Expect = 5e-13
Identities = 41/99 (41%), Positives = 58/99 (58%)
Frame = +2
Query: 284 MISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFT 343
+I G AAAS RVSNELG G + AK ++ V++ S+ +G L +F +F+
Sbjct: 32 LIIYGLSAAASTRVSNELGAGQPERAKHAMRVSLKLSILLGFCFALMIVFGHGIWIRLFS 211
Query: 344 SNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQ 382
S+ + ++P L+ISILL+SVQ VLSGV GWQ
Sbjct: 212 SSPTIKHEFASIAPFLAISILLDSVQGVLSGVVKACGWQ 328
>BQ139895 similar to GP|12231296|gb| ripening regulated protein DDTFR18
{Lycopersicon esculentum}, partial (16%)
Length = 392
Score = 68.2 bits (165), Expect = 7e-12
Identities = 30/92 (32%), Positives = 58/92 (62%), Gaps = 5/92 (5%)
Frame = +2
Query: 2 ERDLKQNLLLKK-----PEEENEQEELSLGKRVWNETKLMWVVAAPAIFTRFSTFGIQII 56
+ +L Q+LL K+ + + +E GK++W ET +W + P+IF+R ++F + ++
Sbjct: 68 DHNLIQSLLSKEISTINKHQHEDDDEQEFGKKLWFETXKLWHIVGPSIFSRVASFTMNVV 247
Query: 57 SQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
+QAF GH+G +LA+ ++ TV++ F G+L+
Sbjct: 248 TQAFAGHLGDVQLASISIANTVIVGFNFGLLL 343
>TC91120 similar to GP|11994404|dbj|BAB02363.
gb|AAC28507.1~gene_id:MIL23.25~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (35%)
Length = 665
Score = 49.3 bits (116), Expect(2) = 2e-11
Identities = 23/64 (35%), Positives = 41/64 (63%)
Frame = +3
Query: 25 LGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFAN 84
+G W E +L++++AAPA+F + + + +Q F GH+G+ ELAA +L T + FA
Sbjct: 159 IGSATWIELRLLFLLAAPAVFVYLINYVMSMSTQIFSGHLGNLELAAASLGNTGIQIFAY 338
Query: 85 GILM 88
G+++
Sbjct: 339 GLML 350
Score = 37.0 bits (84), Expect(2) = 2e-11
Identities = 15/33 (45%), Positives = 23/33 (69%)
Frame = +2
Query: 92 VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDS 124
VG G C + +WTSIW ++IW+ +I +KI +S
Sbjct: 347 VGYGKCS*DTMWTSIWSRKIWHARNIFTKINNS 445
>TC78455 similar to GP|11994404|dbj|BAB02363.
gb|AAC28507.1~gene_id:MIL23.25~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (48%)
Length = 1060
Score = 41.2 bits (95), Expect(2) = 1e-09
Identities = 20/59 (33%), Positives = 36/59 (60%)
Frame = +1
Query: 30 WNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
W E KL++ +AAP++ + + + +Q F GH+G+ ELAA +L + FA G+++
Sbjct: 202 WVEFKLLFYLAAPSVIVYLINYVMSMSTQIFSGHLGNLELAAASLGNNGIQIFAYGLML 378
Score = 39.3 bits (90), Expect(2) = 1e-09
Identities = 31/117 (26%), Positives = 51/117 (43%), Gaps = 1/117 (0%)
Frame = +3
Query: 92 VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDTLRPR*K 151
VG+G+C + +WTSIW K+I N I +KI + + ++ + + +F+ R K
Sbjct: 375 VGHGECC*DTMWTSIWSKKI*NVRHIFAKINSASHNSRFNFNNNIHILRTNFNISRRITK 554
Query: 152 HIRSCRKHFSLVNSYYVCFYCLIHLPNIPSITKQEHHHCI-LGSFFHNHSCIPLLAF 207
+ S N +C + P I S TK CI + S C+ L +
Sbjct: 555 NCISSIPFCLWFNPTNICLCYKLSNPKISSSTKHSSTKCIHISSNIGYSPCVKLCCY 725
>AW980583 weakly similar to GP|16797803|db hypothetical membrane protein-1
{Marchantia polymorpha}, partial (21%)
Length = 631
Score = 53.5 bits (127), Expect = 2e-07
Identities = 27/71 (38%), Positives = 43/71 (60%)
Frame = +3
Query: 265 PFSRRNVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIG 324
P + +V++IC+NI M+S GF AAS+RVSNELG G+ +AA ++ V V+ ++
Sbjct: 405 PKLQTSVLAICMNIASVVWMLSSGFTGAASIRVSNELGAGNPRAAHLAVCVVVVLNITEA 584
Query: 325 SFLFLFFLFFR 335
F+ + R
Sbjct: 585 FFVGTVMILLR 617
>BF642288 similar to GP|1495259|emb orf04 {Arabidopsis thaliana}, partial
(29%)
Length = 654
Score = 51.6 bits (122), Expect(2) = 3e-07
Identities = 23/58 (39%), Positives = 37/58 (63%)
Frame = +3
Query: 31 NETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
NE+K +W +A PAIFT S + + ++Q F G +G+ +LAA ++ +V+ F GI M
Sbjct: 264 NESKKLWYLAGPAIFTSISQYSLGAVTQVFAGQVGTLQLAAVSVENSVIAGFCLGITM 437
Score = 20.4 bits (41), Expect(2) = 3e-07
Identities = 10/37 (27%), Positives = 21/37 (56%)
Frame = +2
Query: 86 ILMKIVVGNGDCVVNPLWTSIWCKRIWNDGSISSKIM 122
+L++ G+G + + +WTS + + +I +KIM
Sbjct: 416 LLLRHHDGDGKRIGDTMWTSFRSRETRHVRNIHAKIM 526
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.329 0.141 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,895,734
Number of Sequences: 36976
Number of extensions: 411153
Number of successful extensions: 3645
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 3571
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3636
length of query: 469
length of database: 9,014,727
effective HSP length: 100
effective length of query: 369
effective length of database: 5,317,127
effective search space: 1962019863
effective search space used: 1962019863
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC145330.19


