

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145221.2 - phase: 0
(704 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
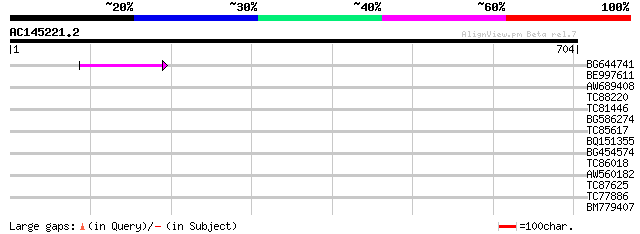
Score E
Sequences producing significant alignments: (bits) Value
BG644741 67 2e-11
BE997611 41 0.001
AW689408 33 0.29
TC88220 similar to GP|15293201|gb|AAK93711.1 unknown protein {Ar... 32 0.83
TC81446 weakly similar to GP|15777865|gb|AAL05893.1 At1g54460/F2... 31 1.4
BG586274 30 2.4
TC85617 similar to GP|2369766|emb|CAA04664.1 hypothetical protei... 30 2.4
BQ151355 homologue to GP|12620658|gb| ID556 {Bradyrhizobium japo... 30 2.4
BG454574 similar to SP|P42762|ERD1 ERD1 protein chloroplast pre... 30 2.4
TC86018 similar to GP|19423939|gb|AAL87312.1 putative RNA helica... 30 2.4
AW560182 similar to PIR|C96543|C96 unknown protein [imported] - ... 30 3.2
TC87625 similar to PIR|T04886|T04886 DAG protein homolog F18F4.1... 30 3.2
TC77886 similar to GP|14532516|gb|AAK63986.1 AT5g05210/K2A11_8 {... 29 7.1
BM779407 similar to GP|6437560|gb|A hypothetical protein {Arabid... 28 9.2
>BG644741
Length = 735
Score = 67.4 bits (163), Expect = 2e-11
Identities = 35/109 (32%), Positives = 54/109 (49%)
Frame = -2
Query: 87 LRLFQHSLIGKAREWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADE 146
LR+F SL+G+A W+ + + + WN L F R++P K ++ + F E
Sbjct: 587 LRVFPLSLMGEADIWFTELPYNSIFTWNQLRDVFLARYYPVSKKLNHNDRVNNFVALPGE 408
Query: 147 SLCDAWERYKSLLRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGS 195
S+ +W+R+ S LR P H DD + F G K++LD A GS
Sbjct: 407 SVSSSWDRFTSFLRSVPNHRIDDDSLKEYFYRGQDDNNKVVLDTIAGGS 261
>BE997611
Length = 547
Score = 41.2 bits (95), Expect = 0.001
Identities = 24/92 (26%), Positives = 47/92 (51%), Gaps = 5/92 (5%)
Frame = -2
Query: 393 KESDVEKDKKKEVGKNVPAHKLPYPKAQSK-----KDLEKHFKRFLDIFKRLEINIPFSE 447
K ++ K K+ +++ A ++ K Q++ K++ F I K ++N+P +E
Sbjct: 546 KR*ELLKSKQNMEDEHINAVEVAEDKPQTEDA*EEKEVPPKFNALHHILKGYKVNMPITE 367
Query: 448 ALEQMPTYAKFMKEILSKKHRYKEDETIQLDA 479
+EQ+P KF++E+L E+E I L +
Sbjct: 366 TVEQIPCCIKFLQELLKTNANLSEEEFISLSS 271
>AW689408
Length = 583
Score = 33.5 bits (75), Expect = 0.29
Identities = 27/94 (28%), Positives = 39/94 (40%), Gaps = 14/94 (14%)
Frame = +2
Query: 306 NQQRQGQFQGQYSGNANQ---------QRFPNNN-----NFQRGNNFRQPWKSNAGPSNQ 351
N Q QF Q+ N NQ +FPN + NF NF Q S++ P+N
Sbjct: 257 NYQHPNQFPNQHLLNPNQFSN*HPQNSNQFPNQHPQNMSNFGFALNFNQ---SSSXPNNF 427
Query: 352 QPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMAN 385
P++ + YP Q + N +M + AN
Sbjct: 428 NPYHRSMMGYPSQTPQ-------FNGYMSMXNAN 508
>TC88220 similar to GP|15293201|gb|AAK93711.1 unknown protein {Arabidopsis
thaliana}, partial (26%)
Length = 1388
Score = 32.0 bits (71), Expect = 0.83
Identities = 35/154 (22%), Positives = 67/154 (42%), Gaps = 16/154 (10%)
Frame = +2
Query: 226 QRKSGILELGSNDANLA-QTKLLTQQMEEMMKQMKEMPKL------VKEQIQKEQ---RH 275
Q + + EL + +L+ Q + ++Q + M +K MP L +K+++Q+ + +
Sbjct: 620 QIRGDVKELSAVRQDLSGQVQAMSQDLSRMTADLKRMPALMVDVEAIKQELQRARAAIEY 799
Query: 276 QQVHFCEMCSG----DHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYS--GNANQQRFPNN 329
++ F E + ++E+ + N +++ + GN Q PN
Sbjct: 800 EKKGFTENYEHGQVMEKKLVAMAREMEKLRAEIANAEKRAHATAAATAAGNPGQGYNPNY 979
Query: 330 NNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPP 363
N + G P+ + G NQ+P NFQQY P
Sbjct: 980 GNAETGYG-GNPYPAYYG-MNQRPGGENFQQYAP 1075
>TC81446 weakly similar to GP|15777865|gb|AAL05893.1 At1g54460/F20D21_28
{Arabidopsis thaliana}, partial (36%)
Length = 1420
Score = 31.2 bits (69), Expect = 1.4
Identities = 52/269 (19%), Positives = 104/269 (38%), Gaps = 17/269 (6%)
Frame = +2
Query: 180 LQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAAKHNRSGT------QRKSGILE 233
+ +P + + G + +SP+ I + D K + G Q + +L
Sbjct: 101 MDKKPNGLASNGSFGDRIRVSPK----IAKMVQAMDHEIKESAEGNSFVEKHQERKDVLN 268
Query: 234 LGSNDANLAQTKLLTQQMEE-MMKQMKEM--PKLVKEQIQKEQRHQQVHFCEMCSGDHPT 290
S N ++ ++ EE M+ KE+ P + + KE H +
Sbjct: 269 TKSTKLNAGLSEEKNEKSEEPKMEGDKELTSPTAITIPVDKE---------------HTS 403
Query: 291 GYCPPQ----LEEEVNYVGNQQRQGQFQGQ-YSGNANQQRFPNNNNFQRGNNFRQPWKSN 345
P Q E+ V + + GQ S NAN PN NN N+ + +SN
Sbjct: 404 PSAPQQSDQATEKHVTHTQTVDTEAVANGQDLSPNANNTHSPNTNNTHSPNSSKNS-QSN 580
Query: 346 AGPSNQQPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKD---KK 402
+ P + + + +++ +D ++ ++T+ RI++ ++ K+ K
Sbjct: 581 S-PFSSRKLAQHDKKHHDDEDNWSVASSSVASTRKVTVGTAPTFRIYERAEKRKEFYMKL 757
Query: 403 KEVGKNVPAHKLPYPKAQSKKDLEKHFKR 431
+E + +L Y +A+ K++ E K+
Sbjct: 758 EEKNRAREEERLQY-EARRKEEEEAALKQ 841
>BG586274
Length = 546
Score = 30.4 bits (67), Expect = 2.4
Identities = 36/147 (24%), Positives = 60/147 (40%), Gaps = 15/147 (10%)
Frame = -3
Query: 298 EEEVNYVGN-QQRQGQFQGQYSGNANQQR--FPNNN--------NFQRGNNFRQPWKSNA 346
+E++++VGN Q + G Q+ F NNN NFQ NN++Q N
Sbjct: 472 QEQLHFVGNPSQDTPTVVHEVEGLEGQEELCFINNNGSWYKKEPNFQY-NNYQQKSYPNN 296
Query: 347 GPSNQQPFNNNFQQYPPQQD----RTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKK 402
S P NN Y PQQ+ + E + + ++ + +Q + H +++
Sbjct: 295 HQSGYPPRNNQQGSYQPQQNPSSGSSAPQESSTDTLLKQILESQTRSEKHVGYELKNLHS 116
Query: 403 KEVGKNVPAHKLPYPKAQSKKDLEKHF 429
K G + A + K+LE F
Sbjct: 115 KIDGSYNELNNKFSHLASTVKNLENQF 35
>TC85617 similar to GP|2369766|emb|CAA04664.1 hypothetical protein {Citrus x
paradisi}, partial (85%)
Length = 1489
Score = 30.4 bits (67), Expect = 2.4
Identities = 14/40 (35%), Positives = 19/40 (47%)
Frame = +1
Query: 298 EEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNN 337
+ VN+ GN +YSGN + P+NNN NN
Sbjct: 388 KSNVNFAGNMN-----SNKYSGNVQLNKDPHNNNSNNNNN 492
>BQ151355 homologue to GP|12620658|gb| ID556 {Bradyrhizobium japonicum},
partial (4%)
Length = 1420
Score = 30.4 bits (67), Expect = 2.4
Identities = 25/102 (24%), Positives = 40/102 (38%)
Frame = +2
Query: 305 GNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQ 364
G QR+G +G+ G Q+R N +G + N GP P ++ ++ P
Sbjct: 170 GPSQREGTPRGESRGKKKQKRREKRQN--KGEKGTKQKPQNRGPG---PGDHPQRKQPKN 334
Query: 365 QDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVG 406
+ + T Q + R KE K+KKK+ G
Sbjct: 335 RGGEGRHSPTTKAENQEPPRTKDEKRKEKEDKERKEKKKQEG 460
>BG454574 similar to SP|P42762|ERD1 ERD1 protein chloroplast precursor.
[Mouse-ear cress] {Arabidopsis thaliana}, partial (19%)
Length = 672
Score = 30.4 bits (67), Expect = 2.4
Identities = 24/85 (28%), Positives = 40/85 (46%), Gaps = 5/85 (5%)
Frame = +1
Query: 502 PVAIGKVNVGKALIDLGSSI-----NLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMG 556
P+ +G+ VGK I G +I + P+ + KRV LD+ L + A+ G
Sbjct: 265 PILLGEAGVGKTAIAEGLAILISRAEVAPFLLTKRVMSLDVGLL--------MAGAKERG 420
Query: 557 IAEDVLVKVDKFVFPVDFVVMDIEE 581
ED + K+ K + V++ I+E
Sbjct: 421 ELEDRVTKLIKDIIESGDVILFIDE 495
>TC86018 similar to GP|19423939|gb|AAL87312.1 putative RNA helicase
{Arabidopsis thaliana}, partial (12%)
Length = 885
Score = 30.4 bits (67), Expect = 2.4
Identities = 26/94 (27%), Positives = 43/94 (45%), Gaps = 5/94 (5%)
Frame = +1
Query: 320 NANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNN--FQQYP--PQQDRTQK-IEDT 374
N N+ R+P RG S G N P N N FQQ P QQ Q+ +
Sbjct: 457 NNNRGRYPPGIGLGRG--------SGGGGLNSNPNNANAGFQQRPHYQQQQYVQRHLMQN 612
Query: 375 LNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKN 408
NQ Q +QQN + +++ + ++ +++ + +N
Sbjct: 613 QNQHQQHYQHHQQNQQQYQQQNQQQQQQQWLRRN 714
>AW560182 similar to PIR|C96543|C96 unknown protein [imported] - Arabidopsis
thaliana, partial (32%)
Length = 706
Score = 30.0 bits (66), Expect = 3.2
Identities = 16/34 (47%), Positives = 24/34 (70%), Gaps = 2/34 (5%)
Frame = +2
Query: 671 EEEKEIEECLKEL--ETLKEIPPKKAKVEELKKE 702
E+E++ E ++E+ E KEI KA+VEELK+E
Sbjct: 212 EKERKARELIEEVCDELAKEIGEDKAEVEELKRE 313
>TC87625 similar to PIR|T04886|T04886 DAG protein homolog F18F4.120 -
Arabidopsis thaliana, partial (27%)
Length = 1019
Score = 30.0 bits (66), Expect = 3.2
Identities = 21/81 (25%), Positives = 31/81 (37%)
Frame = +2
Query: 289 PTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGP 348
P + P Q + +++ QQ GQ Q ++ QRFP
Sbjct: 749 PQNFPPQQNHGQASHIPPQQNIGQQQPNIRIDSTSQRFPP-------------------- 868
Query: 349 SNQQPFNNNFQQYPPQQDRTQ 369
QQ ++ Q+YPPQQ Q
Sbjct: 869 --QQSYDQASQRYPPQQSYDQ 925
>TC77886 similar to GP|14532516|gb|AAK63986.1 AT5g05210/K2A11_8 {Arabidopsis
thaliana}, partial (46%)
Length = 1439
Score = 28.9 bits (63), Expect = 7.1
Identities = 20/69 (28%), Positives = 29/69 (41%)
Frame = +3
Query: 31 GKEKEKQVELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLF 90
G K+++ + K TIQ + LD E P L + A+E EEE ++ F
Sbjct: 318 GLNKKQKEKAKKKTIQNIKKSKVQRLDPEKPVTTLELLKKSLGKEKANEGEEEEDVVKPF 497
Query: 91 QHSLIGKAR 99
L G R
Sbjct: 498 VSELDGDDR 524
>BM779407 similar to GP|6437560|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (16%)
Length = 532
Score = 28.5 bits (62), Expect = 9.2
Identities = 18/74 (24%), Positives = 30/74 (40%)
Frame = +3
Query: 303 YVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYP 362
Y G Q G G S N N+Q FP + N + + +P + PF +++ +
Sbjct: 165 YTGPYQLSGSSVGPSSRNQNEQSFPASPNQPEYHYYSKP--------EELPFAPSYRYHK 320
Query: 363 PQQDRTQKIEDTLN 376
P K+ L+
Sbjct: 321 PPDMSAPKVNAVLS 362
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,426,429
Number of Sequences: 36976
Number of extensions: 255228
Number of successful extensions: 1420
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1394
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1419
length of query: 704
length of database: 9,014,727
effective HSP length: 103
effective length of query: 601
effective length of database: 5,206,199
effective search space: 3128925599
effective search space used: 3128925599
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC145221.2


