

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145202.5 - phase: 0 /pseudo
(1053 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
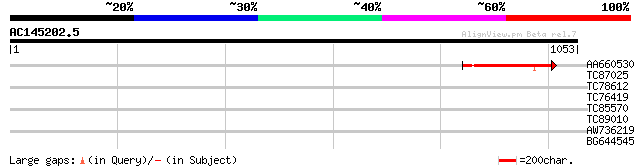
Score E
Sequences producing significant alignments: (bits) Value
AA660530 similar to SP|O24308|TOP DNA topoisomerase II (EC 5.99.... 125 7e-29
TC87025 similar to PIR|S27757|S27757 embryonic abundant protein ... 33 0.75
TC78612 similar to GP|9759092|dbj|BAB09661.1 signal recognition ... 32 1.3
TC76419 similar to SP|Q96558|UGDH_SOYBN UDP-glucose 6-dehydrogen... 32 1.7
TC85570 homologue to SP|Q96558|UGDH_SOYBN UDP-glucose 6-dehydrog... 32 1.7
TC89010 similar to SP|Q96558|UGDH_SOYBN UDP-glucose 6-dehydrogen... 30 3.7
AW736219 similar to GP|23504885|emb hypothetical protein {Plasmo... 30 4.9
BG644545 similar to PIR|H96613|H966 hypothetical protein T15M6.8... 30 6.4
>AA660530 similar to SP|O24308|TOP DNA topoisomerase II (EC 5.99.1.3)
(PsTopII). [Garden pea] {Pisum sativum}, partial (9%)
Length = 566
Score = 125 bits (315), Expect = 7e-29
Identities = 84/184 (45%), Positives = 112/184 (60%), Gaps = 10/184 (5%)
Frame = +1
Query: 842 IVNFKVNVKQEEIATIEKDEELRKKFNLTCTISTSNMYLFDAEGKIKKYNTPEQILEEFC 901
IV+ +V +K E++A I + E L KKF LT TISTSNM+LFDA GKIKK++TP QILEEF
Sbjct: 4 IVDIEVRMKAEKVAAIMQ-EGLFKKFKLTSTISTSNMHLFDAXGKIKKFDTPXQILEEFY 180
Query: 902 TLRLKYYVRRKQYLVKNFTQLLRSLRIKQKFISNIVNEKFNPINR-RAELLIEILKKGKS 960
LRL+YY +RK+Y++ N +LL L K +FI +V+ + NR R ELLIE+ +KG +
Sbjct: 181 PLRLEYYEKRKKYILANLERLLLILDNKVRFILGVVSGEIIVSNRKRTELLIELKQKGFT 360
Query: 961 AEPHVAGAIDYSS------KEQEAGEQESVSQSVS---IEGATLGDYEYLSSLPFENFTL 1011
P +S K +G S +S ++GAT GDYEYL SL TL
Sbjct: 361 PIPEER*ICRTTSCRG***KFXRSGR*LSGKLHLSLSMLKGATWGDYEYLLSLSIGTLTL 540
Query: 1012 ESLK 1015
E +
Sbjct: 541 EXFR 552
>TC87025 similar to PIR|S27757|S27757 embryonic abundant protein 59K -
soybean, partial (56%)
Length = 1353
Score = 32.7 bits (73), Expect = 0.75
Identities = 21/83 (25%), Positives = 38/83 (45%), Gaps = 3/83 (3%)
Frame = +2
Query: 473 YNASDSKKGKELSFDSMQQYEDWKKELGNTAND---WEIKYCQGLSSAQEVREYFQDLGK 529
Y A +K+GK+ + + M +Y+D+ E D W+I E+++ D K
Sbjct: 608 YTAEKAKEGKDATMNKMGEYKDYTAEKAKEGKDNTAWKI---------GEMKDSAVDAAK 760
Query: 530 HRAYFVWDDKDETSIKLAFSKKA 552
++ D K+E K A + +A
Sbjct: 761 RAMGYLGDKKEEAKQKTAETAEA 829
>TC78612 similar to GP|9759092|dbj|BAB09661.1 signal recognition particle
receptor beta subunit-like protein {Arabidopsis
thaliana}, partial (84%)
Length = 963
Score = 32.0 bits (71), Expect = 1.3
Identities = 16/41 (39%), Positives = 22/41 (53%), Gaps = 3/41 (7%)
Frame = +3
Query: 469 KVSMYNASDSKKGKELSFDSMQQYEDWKKE---LGNTANDW 506
K S N + +KGK L F + E WK++ L N AND+
Sbjct: 6 KGSKINQFEERKGKTLEFTMQHELEQWKEQFSHLWNVANDY 128
>TC76419 similar to SP|Q96558|UGDH_SOYBN UDP-glucose 6-dehydrogenase (EC
1.1.1.22) (UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH).
[Soybean], complete
Length = 1898
Score = 31.6 bits (70), Expect = 1.7
Identities = 38/168 (22%), Positives = 64/168 (37%), Gaps = 14/168 (8%)
Frame = +3
Query: 421 RYGQLMVMANQDEDGAHFKGLLINFFY-----------SFWPSLLKVSKFMSVFTFPIIK 469
R G + A+ G+ F+ ++N Y ++W ++KV+ + I
Sbjct: 894 RIGPKFLNASVGFGGSCFQKDILNLVYICECNGLPEVANYWKQVIKVNDYQKARFVNRIV 1073
Query: 470 VSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGLSSAQEVREYFQDLGK 529
SM+N KK L F +KK+ G+T I C+GL LG
Sbjct: 1074 SSMFNTVSGKKIAVLGF-------AFKKDTGDTRETPAIDVCKGL------------LGD 1196
Query: 530 HRAYFVWDDK-DETSIKLAFSKKAEEWE--AWIGNSQPGTCNYERKVI 574
++D + E I + K +W+ A + + P T E V+
Sbjct: 1197 KAKLSIYDPQVSEEQILKDLAMKKFDWDHPAHLQPTSPTTSKKEVSVV 1340
>TC85570 homologue to SP|Q96558|UGDH_SOYBN UDP-glucose 6-dehydrogenase (EC
1.1.1.22) (UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH).
[Soybean], complete
Length = 1810
Score = 31.6 bits (70), Expect = 1.7
Identities = 25/105 (23%), Positives = 43/105 (40%), Gaps = 11/105 (10%)
Frame = +3
Query: 421 RYGQLMVMANQDEDGAHFKGLLINFFY-----------SFWPSLLKVSKFMSVFTFPIIK 469
R G + A+ G+ F+ ++N Y +W ++K++ + +
Sbjct: 855 RIGPKFLNASVGFGGSCFQKDILNLVYICECNGLPEVAEYWKQVIKINDYQKSRFVNRVV 1034
Query: 470 VSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL 514
SM+N +KK L F +KK+ G+T I CQGL
Sbjct: 1035ASMFNTVSNKKIAILGF-------AFKKDTGDTRETPAIDVCQGL 1148
>TC89010 similar to SP|Q96558|UGDH_SOYBN UDP-glucose 6-dehydrogenase (EC
1.1.1.22) (UDP-Glc dehydrogenase) (UDP-GlcDH) (UDPGDH).
[Soybean], partial (84%)
Length = 1659
Score = 30.4 bits (67), Expect = 3.7
Identities = 33/152 (21%), Positives = 58/152 (37%), Gaps = 16/152 (10%)
Frame = +3
Query: 379 LSSKLLNATRDSRKLFKKKEIQNLMAILGLVRNKKYSNAKCL-----RYGQLMVMANQDE 433
L S L+ D+ L ++ N M+ L S C+ + G + A+
Sbjct: 894 LWSAELSKLADNAFLAQRISSINAMSALCEATGADVSQVSCVLSKNTKLGSKYLNASVGF 1073
Query: 434 DGAHFKGLLINFFY-----------SFWPSLLKVSKFMSVFTFPIIKVSMYNASDSKKGK 482
G+ F+ ++N Y ++W ++KV+ + + SM+N KK
Sbjct: 1074 GGSCFQKDILNLVYICESNGLVEVANYWKEVIKVNDYQKSRFVKKVVTSMFNTVSGKKIA 1253
Query: 483 ELSFDSMQQYEDWKKELGNTANDWEIKYCQGL 514
L F +KK+ +T I C+GL
Sbjct: 1254 VLGF-------SFKKDTSDTRKTPAIDVCKGL 1328
>AW736219 similar to GP|23504885|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (1%)
Length = 496
Score = 30.0 bits (66), Expect = 4.9
Identities = 9/19 (47%), Positives = 17/19 (89%)
Frame = -1
Query: 566 TCNYERKVINYRNFVNKEL 584
TC+Y RK++ ++NF+N++L
Sbjct: 319 TCDYNRKILIHKNFINRKL 263
>BG644545 similar to PIR|H96613|H966 hypothetical protein T15M6.8 [imported] -
Arabidopsis thaliana, partial (24%)
Length = 749
Score = 29.6 bits (65), Expect = 6.4
Identities = 13/26 (50%), Positives = 20/26 (76%)
Frame = +3
Query: 1007 ENFTLESLKKLEAELDEKENELKTLE 1032
EN +++KKLEAEL++ + ELK L+
Sbjct: 444 ENNIFDTIKKLEAELEKTKTELKMLK 521
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,719,547
Number of Sequences: 36976
Number of extensions: 415328
Number of successful extensions: 2046
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 2032
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2045
length of query: 1053
length of database: 9,014,727
effective HSP length: 106
effective length of query: 947
effective length of database: 5,095,271
effective search space: 4825221637
effective search space used: 4825221637
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC145202.5


