

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
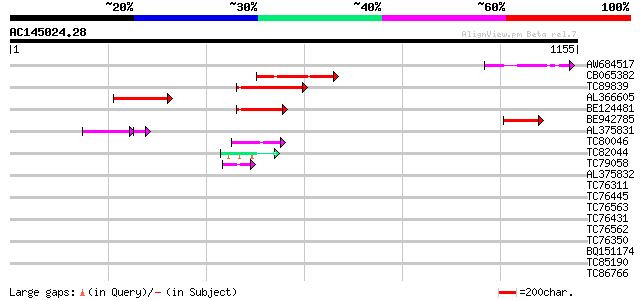
Query= AC145024.28 - phase: 0
(1155 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 151 1e-36
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 148 1e-35
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 138 1e-32
AL366605 132 6e-31
BE124481 111 1e-24
BE942785 84 2e-16
AL375831 66 9e-15
TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor ... 44 4e-04
TC82044 similar to PIR|B86285|B86285 hypothetical protein AAD396... 44 5e-04
TC79058 homologue to GP|20197423|gb|AAM15069.1 unknown protein {... 43 8e-04
AL375832 42 0.001
TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer a... 42 0.001
TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - so... 41 0.003
TC76563 homologue to PIR|C29356|C29356 hydroxyproline-rich glyco... 41 0.003
TC76431 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sat... 40 0.004
TC76562 similar to PIR|T11622|T11622 extensin class 1 precursor ... 39 0.009
TC76350 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sat... 38 0.020
BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xy... 38 0.020
TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protei... 38 0.026
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 36 0.075
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana chromosome
II BAC F26H6; putative retroelement pol polyprotein,
partial (1%)
Length = 488
Score = 151 bits (382), Expect = 1e-36
Identities = 86/186 (46%), Positives = 112/186 (59%), Gaps = 2/186 (1%)
Frame = +1
Query: 967 VTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLKFVSRGKLITVSGESAFLISNLSAF 1026
+TFQVM+I AS+SCLLGRPWIHDAGAVTSTLHQKLKFV G
Sbjct: 1 ITFQVMDINASYSCLLGRPWIHDAGAVTSTLHQKLKFVKNG------------------- 123
Query: 1027 SVIGGSSSDGPSFQGFSAE-ESVGKIETCMASLKDARRVIQEGKTGGWGQLVELPENKRK 1085
S++G +FQG S E K+ MASLKDA++ +QEG+ WG+L++L ENKRK
Sbjct: 124 ------SAEGTAFQGLSMEGAEPKKVGAAMASLKDAQKAVQEGQAADWGKLIQLCENKRK 285
Query: 1086 EGIGFLNSKPGMFDPTRGSFHSAGFIHDSPETNAILDDAPGGV-TPVFVTPGGACCNWIA 1144
EG+ F + + G+FHSAGF+ N + ++ V P+FV PGG +W A
Sbjct: 286 EGLRFSPTS----GVSTGTFHSAGFV------NTLAEEVARFVPRPLFVIPGGIAKDWDA 435
Query: 1145 VDIPSV 1150
VD+PS+
Sbjct: 436 VDVPSI 453
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 148 bits (374), Expect = 1e-35
Identities = 86/175 (49%), Positives = 111/175 (63%), Gaps = 7/175 (4%)
Frame = -2
Query: 503 PYNQQNQKQQFDP----LPMTYGALLPSLLAQN---LVQTIPPPRIPDPLPRWYRPDLHC 555
P +QQ Q QQ P +PM Y LP+LL + + Q PPP DPLP +R DL C
Sbjct: 518 PVHQQRQ*QQARPTFPLIPMLYAE*LPTLLLRGHCTIRQGKPPP---DPLPPRFRSDLKC 348
Query: 556 IYHQGAPGHDVERCFALKKEVQKLINSKELTFTDPDAVAQNNPLPTHGPAVNMIQDDQEE 615
+HQGA GHDVE C+ALK V+KLIN +LTF + +NPLP H AVNMI + EE
Sbjct: 347 DFHQGALGHDVEGCYALKHIVKKLINQGKLTFENNVPHVLDNPLPNHA-AVNMI-EVYEE 174
Query: 616 ARILSVGDIKTPLVPIHVKMCKATLFNHNHEACDICLMDPRGCIQVQNDMQGLLN 670
A L V ++ TPLVP+H+K+C+A+LF+H+H C +P GC VQND+Q L+N
Sbjct: 173 APGLDVRNVTTPLVPLHIKLCQASLFDHDHANCQE*FYNPLGCCVVQNDIQSLMN 9
Score = 32.7 bits (73), Expect = 0.83
Identities = 15/43 (34%), Positives = 27/43 (61%)
Frame = -1
Query: 6 QENAQLREELVNLKGEMERMALMMETMMAEREQAAISNSTPVV 48
+ENAQLR EL +++ E+ + M ++A +EQ++ + T V
Sbjct: 624 EENAQLRTELASMREELAKANDTMTALLAAQEQSSTGSPTTTV 496
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 138 bits (348), Expect = 1e-32
Identities = 71/147 (48%), Positives = 90/147 (60%), Gaps = 4/147 (2%)
Frame = +3
Query: 463 PPLYQPLYNLQQLSQQPYYPQQPYQQRPYN--PPQQQPRPQAPYNQQNQKQQFDPLPMTY 520
PPL Q Q QP PQ+P QQ P P Q P+ P NQ + Q DP+P+ Y
Sbjct: 228 PPLVQ---QRPQSPYQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVKY 398
Query: 521 GALLPSLLAQNLVQTIPPPRIPDPLPRWYRPDLHCIYHQGAPGHDVERCFALKKEVQKLI 580
LLP LL +NL+QT+P PR+P+ LP WYRPDL+C++HQGAPGHD E+C+ LK+EVQKLI
Sbjct: 399 ADLLPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHDTEQCYPLKEEVQKLI 578
Query: 581 NSKELTFTDPD--AVAQNNPLPTHGPA 605
+ +F D D + Q L H A
Sbjct: 579 ENNVWSFDDQDIKVLLQQQHLAPHSGA 659
>AL366605
Length = 422
Score = 132 bits (333), Expect = 6e-31
Identities = 65/124 (52%), Positives = 87/124 (69%), Gaps = 2/124 (1%)
Frame = -3
Query: 211 DLCLVPDVNIPPKFKMPVFEKYQGDTCPQNHLTMYISKMIAYKNNVPLLIHCFQDSLTGP 270
DLCLV V++P KFK+P F++Y G TCPQNH+ Y+ KM YK+N L+IHCFQDSL
Sbjct: 372 DLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMED 193
Query: 271 AHTWFMGL--KGVTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQ 328
A W+ L + TF++LA AF Y +NT L P+R+ L+SL+QK +ESF+EYAQR+
Sbjct: 192 AAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRG 13
Query: 329 KAAQ 332
AA+
Sbjct: 12 AAAR 1
>BE124481
Length = 538
Score = 111 bits (278), Expect = 1e-24
Identities = 54/105 (51%), Positives = 67/105 (63%), Gaps = 2/105 (1%)
Frame = +3
Query: 463 PPLYQPLYNLQQLSQQPYYPQQPYQQRPYN--PPQQQPRPQAPYNQQNQKQQFDPLPMTY 520
PPL Q Q QP PQ+P QQ P P Q P+ P NQ + Q DP+P+ Y
Sbjct: 231 PPLVQ---QRPQSPYQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVKY 401
Query: 521 GALLPSLLAQNLVQTIPPPRIPDPLPRWYRPDLHCIYHQGAPGHD 565
LLP LL +NL+QT+P PR+P+ LP WYRPDL+C++HQGAPGHD
Sbjct: 402 ADLLPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHD 536
>BE942785
Length = 460
Score = 84.3 bits (207), Expect = 2e-16
Identities = 41/81 (50%), Positives = 58/81 (70%), Gaps = 1/81 (1%)
Frame = -2
Query: 1007 GKLITVSGESAFLISNLSAFSVIGGSSSDGPSFQGFSAEESVGKIE-TCMASLKDARRVI 1065
G L+T+ GE A+LIS LS+FS I S++G +FQG + E + K + T MASLKDA+R +
Sbjct: 375 GXLVTIHGEEAYLISQLSSFSCIEAGSAEGTAFQGLTVEGTEPKRDGTAMASLKDAQRAV 196
Query: 1066 QEGKTGGWGQLVELPENKRKE 1086
QE + GWG+L++L ENK K+
Sbjct: 195 QESQAAGWGRLIQLRENKHKD 133
>AL375831
Length = 467
Score = 65.9 bits (159), Expect(2) = 9e-15
Identities = 40/108 (37%), Positives = 54/108 (49%), Gaps = 1/108 (0%)
Frame = +2
Query: 148 IPQATMTQAGPTVHVGPQHEE-QIYHSDSIMGDDKAIDWEERFGALEKKMSNMRGKEAVV 206
+PQ T P VH PQ H + EE L ++ RG V
Sbjct: 44 LPQTTAAVTEPLVHTLPQGININTQHRSIPVTKTMEEMMEELAKELRHEIQANRGNADSV 223
Query: 207 QSIYDLCLVPDVNIPPKFKMPVFEKYQGDTCPQNHLTMYISKMIAYKN 254
++ DLCLV V++P KFK+P F++Y G TCPQNH+ Y+ KM YK+
Sbjct: 224 KT-QDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 364
Score = 33.5 bits (75), Expect(2) = 9e-15
Identities = 16/36 (44%), Positives = 21/36 (57%), Gaps = 2/36 (5%)
Frame = +3
Query: 253 KNNVPLLIHCFQDSLTGPAHTWFMGL--KGVTTFEQ 286
K N L+IHCFQDSL A W+ L + TF++
Sbjct: 360 KTNDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDE 467
>TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor (EC
3.2.1.4) (Endo-1 4-beta-glucanase) (Spore germination
protein 270-6), partial (11%)
Length = 643
Score = 43.9 bits (102), Expect = 4e-04
Identities = 36/112 (32%), Positives = 45/112 (40%), Gaps = 2/112 (1%)
Frame = +1
Query: 453 PRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPR--PQAPYNQQNQK 510
P P P PY L + L PQ+P Q+ P PPQ+ P+ PQ P + QK
Sbjct: 199 PLPL*PQPYLLLL*PIPLQTRSLPLPQRPPQRPPQRPPQKPPQRPPQKPPQRPPQRPPQK 378
Query: 511 QQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWYRPDLHCIYHQGAP 562
LP LP Q L Q +P PLP +P L + H P
Sbjct: 379 -----LPQRPPQKLPQRPPQKLPQKLPQKPPQRPLPHPQKPPLRPLPHPQKP 519
Score = 36.2 bits (82), Expect = 0.075
Identities = 35/122 (28%), Positives = 47/122 (37%), Gaps = 3/122 (2%)
Frame = +1
Query: 440 PQMTAPQNPNYQSP-RPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQP 498
P PQ P + P RP P PP P Q+L Q+P P+ Q+P P + P
Sbjct: 343 PPQRPPQRPPQKLPQRPPQKLPQRPPQKLP----QKLPQKPPQRPLPHPQKP--PLRPLP 504
Query: 499 RPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPD--PLPRWYRPDLHCI 556
PQ P + Q PL + +P P+ P PLP +P L +
Sbjct: 505 HPQKPPLRPLPHPQKPPL-----------------RPLPHPQKPPLRPLPHPQKPPLRTL 633
Query: 557 YH 558
H
Sbjct: 634 PH 639
>TC82044 similar to PIR|B86285|B86285 hypothetical protein AAD39642.1
[imported] - Arabidopsis thaliana, partial (16%)
Length = 950
Score = 43.5 bits (101), Expect = 5e-04
Identities = 46/157 (29%), Positives = 53/157 (33%), Gaps = 38/157 (24%)
Frame = +1
Query: 430 SNHPFVANITPQMTA-----PQNPNYQSPRPQGPAPYY---PPLY-------QPLYNLQQ 474
+N+P V N + PQNP YQ P Q P Y PP Y P Y +
Sbjct: 442 NNYPSVGNQNQRSNTQTDPRPQNPYYQ-PSEQPPVSSYAHPPPSYGHSHHQPPPPYQIPP 618
Query: 475 LSQQPYYPQQPYQQRP------------------YN-----PPQQQPRPQAPYNQQNQKQ 511
S PY P Q +QQ+P YN P QQQP P+ PY
Sbjct: 619 TSTAPYPPPQVHQQQPPANHDYGQPAYPGWRAPYYNNAPTHPQQQQPGPRPPY------- 777
Query: 512 QFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRW 548
TIP P P P PRW
Sbjct: 778 -----------------------TIPSP-YPPPSPRW 816
>TC79058 homologue to GP|20197423|gb|AAM15069.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 627
Score = 42.7 bits (99), Expect = 8e-04
Identities = 30/74 (40%), Positives = 33/74 (44%), Gaps = 6/74 (8%)
Frame = +1
Query: 434 FVANITPQMTAPQNPNYQSPRPQG--PAPYYPPLYQPLYNLQQLSQQPY----YPQQPYQ 487
F+ NI Q P P PQG P YPP P Y Q QQ Y YPQQ Y
Sbjct: 73 FINNIMSYYDHQQQPPVGVPPPQGYPPKDAYPP---PGYPQQGYPQQGYPPQGYPQQGYP 243
Query: 488 QRPYNPPQQQPRPQ 501
Q+ Y P Q +PQ
Sbjct: 244 QQGYPPQQYAQQPQ 285
>AL375832
Length = 535
Score = 42.4 bits (98), Expect = 0.001
Identities = 19/34 (55%), Positives = 25/34 (72%), Gaps = 1/34 (2%)
Frame = +3
Query: 1118 NAILDDAPG-GVTPVFVTPGGACCNWIAVDIPSV 1150
NAI ++A G G+ P FVTPGG +W A+DIPS+
Sbjct: 9 NAISEEATGSGLRPAFVTPGGIASDWDAIDIPSI 110
>TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer arietinum},
partial (77%)
Length = 973
Score = 42.0 bits (97), Expect = 0.001
Identities = 33/120 (27%), Positives = 43/120 (35%), Gaps = 11/120 (9%)
Frame = +2
Query: 440 PQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY-----PQQPYQQRPY--- 491
P + P Y SP P P+P P Y+ PYY P P PY
Sbjct: 5 PSPSPPPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYQ 184
Query: 492 NPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQN---LVQTIPPPRIPDPLPRW 548
+PP P P PY+ ++ P Y + P + T PPP P P P +
Sbjct: 185 SPPPPSPTPHTPYHYKSPPPPTASPPPPYHYVSPPPPTSSPPPYHYTSPPPPSPAPAPTY 364
Score = 30.0 bits (66), Expect = 5.4
Identities = 24/91 (26%), Positives = 29/91 (31%), Gaps = 10/91 (10%)
Frame = +2
Query: 482 PQQPYQQRPY---NPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPP 538
P P PY +PP P P PY ++ P Y PP
Sbjct: 2 PPSPSPPPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPPPY------------YYKSPP 145
Query: 539 PRIPDPLPRWY-------RPDLHCIYHQGAP 562
P P P P +Y P H YH +P
Sbjct: 146 PPSPSPPPPYYYQSPPPPSPTPHTPYHYKSP 238
>TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - soybean
(fragment), partial (76%)
Length = 821
Score = 40.8 bits (94), Expect = 0.003
Identities = 35/126 (27%), Positives = 48/126 (37%), Gaps = 1/126 (0%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQ 483
P P Y P P +P +P Y +SP P P+P P Y+ S P P
Sbjct: 11 PPPYYYKSP-----PPPSPSPPSPYYYKSPPPPSPSPPPPYYYK--------SPPPPSPS 151
Query: 484 QPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPD 543
P +PP P P PY+ Q+ P P+++ N ++ PPP
Sbjct: 152 PPPPYYYKSPPPPSPSPPPPYHYQSPP---PPSPISH--------PPNYYKSPPPPSPSP 298
Query: 544 PLPRWY 549
P P Y
Sbjct: 299 PPPYHY 316
Score = 33.5 bits (75), Expect = 0.49
Identities = 28/89 (31%), Positives = 34/89 (37%), Gaps = 11/89 (12%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY-- 481
P P Y P P +P P Y +SP P P+P P YQ +S P Y
Sbjct: 107 PPPYYYKSP-----PPPSPSPPPPYYYKSPPPPSPSPPPPYHYQSPPPPSPISHPPNYYK 271
Query: 482 ---PQQPYQQRPYN-----PPQQQPRPQA 502
P P PY+ PP + P P A
Sbjct: 272 SPPPPSPSPPPPYHYVSPPPPVKSPPPPA 358
>TC76563 homologue to PIR|C29356|C29356 hydroxyproline-rich glycoprotein
(clone Hyp2.13) - kidney bean (fragment), partial (34%)
Length = 507
Score = 40.8 bits (94), Expect = 0.003
Identities = 30/108 (27%), Positives = 39/108 (35%), Gaps = 8/108 (7%)
Frame = +2
Query: 450 YQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY-----PQQPYQQRPY---NPPQQQPRPQ 501
Y+SP P P+P P Y+ + + PYY P P PY +PP P P
Sbjct: 149 YKSPPPPSPSPPPPYYYKSPPPPSPIPKTPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPP 328
Query: 502 APYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWY 549
PY ++ P Y PPP P P P +Y
Sbjct: 329 PPYYYKSPPPPSPSPPPPY------------YYKSPPPPSPSPPPPYY 436
Score = 38.1 bits (87), Expect = 0.020
Identities = 31/104 (29%), Positives = 39/104 (36%), Gaps = 9/104 (8%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY-- 481
P P Y P P P+ P Y +SP P P+P P Y+ PYY
Sbjct: 182 PPPYYYKSP-----PPPSPIPKTPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYK 346
Query: 482 ---PQQPYQQRPY---NPPQQQPRPQAPYNQQNQKQQFDPLPMT 519
P P PY +PP P P PY ++ P+P T
Sbjct: 347 SPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPP-SPIPHT 475
Score = 37.0 bits (84), Expect = 0.044
Identities = 26/89 (29%), Positives = 34/89 (37%), Gaps = 9/89 (10%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY-- 481
P P+ + + P +P P Y +SP P P+P P Y+ PYY
Sbjct: 215 PSPIPKTPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYK 394
Query: 482 ---PQQPYQQRPY---NPPQQQPRPQAPY 504
P P PY +PP P P PY
Sbjct: 395 SPPPPSPSPPPPYYYKSPPPPSPIPHTPY 481
Score = 36.6 bits (83), Expect = 0.057
Identities = 35/133 (26%), Positives = 41/133 (30%), Gaps = 11/133 (8%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY-- 481
P P Y P P +P P Y +SP P P P P Y+ PYY
Sbjct: 134 PPPYYYKSP-----PPPSPSPPPPYYYKSPPPPSPIPKTPYYYKSPPPPSPSPPPPYYYK 298
Query: 482 --------PQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLV 533
P PY + PP P P Y P P Y +
Sbjct: 299 SPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKS----------- 445
Query: 534 QTIPPPRIPDPLP 546
PPP P P+P
Sbjct: 446 ---PPP--PSPIP 469
Score = 29.3 bits (64), Expect = 9.2
Identities = 19/62 (30%), Positives = 25/62 (39%), Gaps = 1/62 (1%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQ 483
P P Y P P +P P Y +SP P P+P P Y+ + PYY +
Sbjct: 326 PPPYYYKSP-----PPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHTPYYYK 490
Query: 484 QP 485
P
Sbjct: 491 SP 496
>TC76431 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sativum},
partial (30%)
Length = 890
Score = 40.4 bits (93), Expect = 0.004
Identities = 33/139 (23%), Positives = 55/139 (38%), Gaps = 21/139 (15%)
Frame = +1
Query: 422 HGGPQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYN---------- 471
H P P HP+ + +P +P+ P Y+ P P P ++ P P+Y+
Sbjct: 49 HPHPHPHPHPHPYPVHHSPP-PSPKKP-YKYPSPPPPTHHHYPHPHPVYHSPPPPKYKYS 222
Query: 472 -----LQQLSQQPYYPQQPYQQRPYN----PPQQQP--RPQAPYNQQNQKQQFDPLPMTY 520
+ +S +PYY P ++PY PP P P+ Y+ + P P+ +
Sbjct: 223 SPPPPVHHVSPKPYYHSPPPPKKPYKYSSPPPPVHPPYTPKPVYHSPPVHPPYTPHPVYH 402
Query: 521 GALLPSLLAQNLVQTIPPP 539
P + + PPP
Sbjct: 403 S---PPPPVHSYIYASPPP 450
>TC76562 similar to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 1391
Score = 39.3 bits (90), Expect = 0.009
Identities = 31/108 (28%), Positives = 40/108 (36%), Gaps = 8/108 (7%)
Frame = +2
Query: 450 YQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY-----PQQPYQQRPY---NPPQQQPRPQ 501
Y+SP P P+P P Y+ + PYY P P PY +PP P P
Sbjct: 20 YKSPPPPSPSPPPPYYYKSPPPPSPIPHTPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPH 199
Query: 502 APYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWY 549
PY ++ P P P +L PPP + P P Y
Sbjct: 200 TPYYYKS------PPP-------PKVLPPPYYYNSPPPPVAYPHPHPY 304
Score = 35.0 bits (79), Expect = 0.17
Identities = 30/116 (25%), Positives = 42/116 (35%), Gaps = 15/116 (12%)
Frame = +2
Query: 451 QSPRPQGPAPYY---PPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPP---------QQQP 498
+ P+P P PYY PP P+Y +Y PY + PP + P
Sbjct: 695 EKPKPS-PTPYYYKSPPPPSPVYKYNSPPPPVHYYSPPYTYKSPPPPVKAAPTPYYYKSP 871
Query: 499 RPQAP---YNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWYRP 551
P AP YN + P TY + P + A P P P+ ++ P
Sbjct: 872 PPPAPVYKYNSPPPPVHYYSPPYTYKSPPPPVKAAPTPYYYKSPPPPAPVYKYNSP 1039
Score = 34.3 bits (77), Expect = 0.28
Identities = 23/80 (28%), Positives = 29/80 (35%), Gaps = 1/80 (1%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQ 483
P P Y P P P P Y +SP P P+P P Y+ + PYY +
Sbjct: 53 PPPYYYKSP-----PPPSPIPHTPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHTPYYYK 217
Query: 484 QPYQQRPYNPPQQQPRPQAP 503
P + PP P P
Sbjct: 218 SPPPPKVLPPPYYYNSPPPP 277
Score = 29.6 bits (65), Expect = 7.0
Identities = 24/82 (29%), Positives = 33/82 (39%), Gaps = 1/82 (1%)
Frame = +2
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNY-QSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQ 483
P P+ + + P +P P Y +SP P P P+ P Y+ + L PYY
Sbjct: 86 PSPIPHTPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHTPYYYKSPPPPKVLPP-PYYYN 262
Query: 484 QPYQQRPYNPPQQQPRPQAPYN 505
P PP P P PY+
Sbjct: 263 SP------PPPVAYPHPH-PYH 307
>TC76350 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sativum},
partial (63%)
Length = 1008
Score = 38.1 bits (87), Expect = 0.020
Identities = 45/167 (26%), Positives = 59/167 (34%), Gaps = 29/167 (17%)
Frame = +2
Query: 425 PQPVYSNHPFVANITP---QMTAPQNPNYQSPRP-----QGPAPYY---PPLYQ------ 467
P PVYS P V + P + +P P Y P P P P Y PP+Y+
Sbjct: 212 PPPVYSPPPPVYSPPPPVYKYKSPPPPVYSPPPPVYKYKSPPPPVYSPPPPVYKYKSPPP 391
Query: 468 -------PLYNLQQLSQQPYYPQQP-YQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMT 519
P+Y + Y P P Y+ + PP P P + + P P
Sbjct: 392 PVYSPPPPVYKYKSPPPPVYSPPPPVYKYKSPPPPVYSPPPPVYKYKSPPPPVYSPPPPV 571
Query: 520 YGALLPSLLAQNLVQTIPPP----RIPDPLPRWYRPDLHCIYHQGAP 562
Y P V + PPP + P P P + P H IY P
Sbjct: 572 YKYKSP----PPPVYSPPPPVYKYKSPPP-PAYSPPPPHYIYSSPPP 697
>BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xylostella
granulovirus}, partial (42%)
Length = 909
Score = 38.1 bits (87), Expect = 0.020
Identities = 31/110 (28%), Positives = 42/110 (38%), Gaps = 4/110 (3%)
Frame = +1
Query: 410 PKRNAQEVGMVAHGGPQPVYSNHPFVANITPQMTAPQN----PNYQSPRPQGPAPYYPPL 465
PKR +E A PQP + Q AP+ P Q P P P P PP
Sbjct: 28 PKRPPKEKDTQARN-PQPEEATKK-------QKRAPEKKKNPPRGQPPPPPPPPPQAPPT 183
Query: 466 YQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDP 515
P + + P P P P PP Q P+P P++++ + P
Sbjct: 184 KPPTRQTPKNNTPPPPPHTPPDPTPPQPPPQPPKP--PHHEKRPRTPEPP 327
>TC85190 similar to GP|8894548|emb|CAB95829.1 hypothetical protein {Cicer
arietinum}, partial (50%)
Length = 1496
Score = 37.7 bits (86), Expect = 0.026
Identities = 19/61 (31%), Positives = 33/61 (53%), Gaps = 2/61 (3%)
Frame = -3
Query: 454 RPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYN--QQNQKQ 511
+P+ P P+ +P ++ QQ PY PQ P +P+ Q++ + Q PY Q+ Q+Q
Sbjct: 420 KPR*FLP*KKPIEKPCFHQQQFHPPPYQPQLPPASQPHQQQQEEDQQQLPYQWWQR*QRQ 241
Query: 512 Q 512
+
Sbjct: 240 R 238
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 36.2 bits (82), Expect = 0.075
Identities = 39/142 (27%), Positives = 50/142 (34%), Gaps = 11/142 (7%)
Frame = +2
Query: 421 AHGGPQPVYS----NHPFVANITPQMTAPQNPNYQSPRP---QGPAPYYPPLYQPLYNLQ 473
AH P P S H + P ++ P P SP P P P PP +P
Sbjct: 137 AHSPPPPPXSPPPPPHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPPP 316
Query: 474 QLSQQPYYPQQPY-QQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNL 532
+P P P Q PY+PP P +P P P Y + PS +
Sbjct: 317 PPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYP-----SPPPPVYTSPPPSPV---Y 472
Query: 533 VQTIPPPRI---PDPLPRWYRP 551
PPP + P P P + P
Sbjct: 473 AYPSPPPPVYSSPPPPPVYEGP 538
Score = 32.3 bits (72), Expect = 1.1
Identities = 28/107 (26%), Positives = 32/107 (29%)
Frame = +2
Query: 445 PQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPY 504
P P P P PAP + P PYY P ++PP P P
Sbjct: 32 PPPPPNSPPPPPPPAPVFSP----------PPPVPYYYSSPPPPPAHSPPPPPXSPPPP- 178
Query: 505 NQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWYRP 551
P P Y L P PPP + P P Y P
Sbjct: 179 -------PHSPTPPVYPYLSPP----------PPPPVHSPPPPVYSP 268
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.133 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,276,201
Number of Sequences: 36976
Number of extensions: 539858
Number of successful extensions: 5313
Number of sequences better than 10.0: 226
Number of HSP's better than 10.0 without gapping: 3800
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4810
length of query: 1155
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1048
effective length of database: 5,058,295
effective search space: 5301093160
effective search space used: 5301093160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC145024.28


