

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145024.2 - phase: 0
(355 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
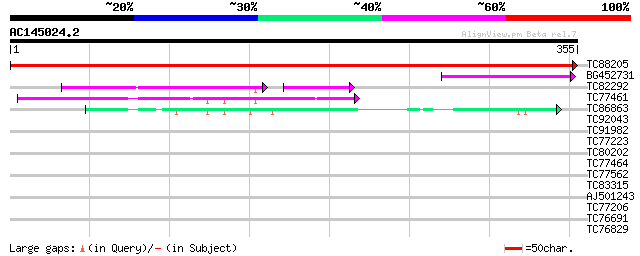
TC88205 similar to GP|15028029|gb|AAK76545.1 unknown protein {Ar... 689 0.0
BG452731 weakly similar to PIR|A96596|A965 hypothetical protein ... 62 4e-10
TC82292 similar to PIR|A96596|A96596 hypothetical protein T18I3.... 47 2e-09
TC77461 similar to GP|10177534|dbj|BAB10929. apospory-associated... 46 2e-05
TC86863 similar to GP|19387263|gb|AAL87174.1 putative apospory-a... 44 9e-05
TC92043 similar to GP|21554299|gb|AAM63374.1 apospory-associated... 35 0.044
TC91982 weakly similar to GP|170525|gb|AAA34202.1|| phenol hydro... 31 0.83
TC77223 weakly similar to GP|19911209|dbj|BAB86931. glucosyltran... 29 2.4
TC80202 similar to GP|4512677|gb|AAD21731.1| unknown protein {Ar... 28 4.1
TC77464 similar to GP|13272433|gb|AAK17155.1 unknown protein {Ar... 28 5.4
TC77562 similar to GP|17104733|gb|AAL34255.1 unknown protein {Ar... 28 7.0
TC83315 weakly similar to GP|19911209|dbj|BAB86931. glucosyltran... 28 7.0
AJ501243 weakly similar to PIR|A40411|A40 translation elongation... 28 7.0
TC77206 similar to GP|18377735|gb|AAL67017.1 putative alpha-gala... 27 9.2
TC76691 similar to SP|O64981|RCA_PHAVU Ribulose bisphosphate car... 27 9.2
TC76829 similar to PIR|T14544|T14544 fructokinase (EC 2.7.1.4) -... 27 9.2
>TC88205 similar to GP|15028029|gb|AAK76545.1 unknown protein {Arabidopsis
thaliana}, partial (45%)
Length = 1371
Score = 689 bits (1777), Expect = 0.0
Identities = 348/356 (97%), Positives = 349/356 (97%), Gaps = 1/356 (0%)
Frame = +1
Query: 1 MASFLSLSLPKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNG 60
MASFLSLSLPKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNG
Sbjct: 115 MASFLSLSLPKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNG 294
Query: 61 SSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGAK 120
SSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGAK
Sbjct: 295 SSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGAK 474
Query: 121 ELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKPV 180
ELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKPV
Sbjct: 475 ELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKPV 654
Query: 181 TLTNAILSHFRFKRRGGA-AIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGA 239
TLTNAILSHFRFK + KGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGA
Sbjct: 655 TLTNAILSHFRFKEKEVVRQFKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGA 834
Query: 240 EPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVI 299
EPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVI
Sbjct: 835 EPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVI 1014
Query: 300 RMGFEDIYVSSPGSMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDNLS 355
RMGFEDIYVSSPGSMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDNLS
Sbjct: 1015RMGFEDIYVSSPGSMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDNLS 1182
>BG452731 weakly similar to PIR|A96596|A965 hypothetical protein T18I3.2
[imported] - Arabidopsis thaliana, partial (14%)
Length = 656
Score = 61.6 bits (148), Expect = 4e-10
Identities = 27/85 (31%), Positives = 49/85 (56%), Gaps = 1/85 (1%)
Frame = +3
Query: 271 KERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMS-DKYGRDYFVCTGPA 329
KER Y P + +ID+GR V R GF++ Y+ SPGS + Y + ++C G A
Sbjct: 207 KERISLVYTNAPRSFTLIDRGRRNSVLVGRNGFDETYLFSPGSSGIESYSKYAYICVGQA 386
Query: 330 SILEPVTVNPGEEFRGAQVIEHDNL 354
++L+P+ ++P + ++G + + N+
Sbjct: 387 AVLQPIVLSPQDVWKGGAYLHNPNI 461
>TC82292 similar to PIR|A96596|A96596 hypothetical protein T18I3.2
[imported] - Arabidopsis thaliana, partial (15%)
Length = 795
Score = 47.0 bits (110), Expect(2) = 2e-09
Identities = 34/132 (25%), Positives = 63/132 (46%), Gaps = 3/132 (2%)
Frame = +2
Query: 33 LNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYT 92
L +F G+ F E + ++ ++NGS ++ LP +TSYK + W +E L
Sbjct: 233 LEQEFNGHGVTFEEVGGSCMAKMELKNGSIATMLLPSGLITSYKAPM-WHGGKVELLNTA 409
Query: 93 IPANETGLYKAKGGIGLVLN-EVLQPGAKELLPSTLEWTVKDVDYDAIDALQVELISTNR 151
+ E G +GG+ L N + E+ S W + ++ ++ +++QV LI+
Sbjct: 410 VSEGEFGDAVIQGGVSLNFNLQTHDDNDDEVSWSPSNWVLHNIKGNSEESIQVVLINKGP 589
Query: 152 F--FDMTYIVSL 161
+ + YIV+L
Sbjct: 590 YDKIGLKYIVTL 625
Score = 32.0 bits (71), Expect(2) = 2e-09
Identities = 14/45 (31%), Positives = 23/45 (51%)
Frame = +3
Query: 172 VKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPPLDS 216
+ N + P+ +T +ILSH GL+ +Y S PP++S
Sbjct: 657 ISNSNSLPLQMTGSILSHLTVSTPEATYAIGLERSNYYSRPPIES 791
>TC77461 similar to GP|10177534|dbj|BAB10929. apospory-associated protein
C-like {Arabidopsis thaliana}, partial (89%)
Length = 1323
Score = 46.2 bits (108), Expect = 2e-05
Identities = 58/227 (25%), Positives = 102/227 (44%), Gaps = 13/227 (5%)
Frame = +3
Query: 6 SLSLPKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESN-SIPIVELTVRNGSSLS 64
S++L +N + + A + ++ N + G+K E ++P + LT GS
Sbjct: 39 SMALCGVNTVTFQNRARYNSGIAFASLGNKEKTTLGVKVTEGEGNLPKLVLTSPAGSEAE 218
Query: 65 LRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKA-KGGIGLVLNEVLQPGAKEL- 122
+ L +TS WK ++LL+ P K GGI + PGA +
Sbjct: 219 IYLFGGCITS------WKVPNGKDLLFVRPDAVFNKKKPISGGIPHCFPQ-FGPGAIQQH 377
Query: 123 -LPSTLEWTVKD---VDYDAIDALQVELISTNRF-----FDMTYIVSLYPVSMATAVIVK 173
L+WTV D V+ + + L+++ + +R F Y V+L S++T + VK
Sbjct: 378 GFARNLDWTVGDSENVEGNPVVTLELKDDTYSRAMWDFSFHALYKVTLNAKSLSTELKVK 557
Query: 174 NKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSYCS-HPPLDSPFQ 219
N K + A+ ++FR GA++KGL+ C + HP ++P +
Sbjct: 558 NTDNKAFSFNTALHTYFR-ASVSGASVKGLKGCKTLNKHPDPNNPIE 695
>TC86863 similar to GP|19387263|gb|AAL87174.1 putative apospory-associated
protein C {Oryza sativa (japonica cultivar-group)},
partial (88%)
Length = 1640
Score = 43.9 bits (102), Expect = 9e-05
Identities = 70/327 (21%), Positives = 123/327 (37%), Gaps = 29/327 (8%)
Frame = +1
Query: 48 NSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKA---- 103
N + V L G+S + L HVTS WK+D EELL+ + ++K
Sbjct: 250 NGLDKVLLRESRGASAEIYLYGGHVTS------WKNDHAEELLFL---SSKAIFKPPKAI 402
Query: 104 KGGIGLVLNEVLQPGAKEL--LPSTLEWTVKD----VDYDAIDALQVELIST-------- 149
+GGI + + G E W ++D + + V+LI
Sbjct: 403 RGGIPICFPQFANHGNLESHGFARNRVWAIEDDPPPFPTNNLSKAFVDLILKPSEEDMKI 582
Query: 150 -NRFFDMTYIVSLYP---VSMATAVIVKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQT 205
N F+ V+L P + + + + N KP + T A ++ ++GL+T
Sbjct: 583 WNHSFEYRLRVALGPGGDLMLTSRIRNTNSDGKPFSFTFAYHTYLSVSDISEVRVEGLET 762
Query: 206 CSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMSRV 265
Y + F T+QG +T E+++ ++
Sbjct: 763 LDYLDNLQNKERF------------------------------TEQGDALTF-ESEVDKI 849
Query: 266 FAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDK----YGRD 321
Y + P+K IID ++ + + + G D V +P K +G D
Sbjct: 850 ------------YLSTPTKIAIIDHEKKRTFVLRKDGLPDAVVWNPWDKKAKAMADFGDD 993
Query: 322 ---YFVCTGPASILEPVTVNPGEEFRG 345
+ +C A+I + +T+ PGEE++G
Sbjct: 994 EYKHMLCVEAANIEKAITLKPGEEWKG 1074
>TC92043 similar to GP|21554299|gb|AAM63374.1 apospory-associated protein C
{Arabidopsis thaliana}, partial (37%)
Length = 1163
Score = 35.0 bits (79), Expect = 0.044
Identities = 26/105 (24%), Positives = 48/105 (44%), Gaps = 7/105 (6%)
Frame = +2
Query: 248 WTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIY 307
+T+QG +T E+++ RV Y S ++D R+ + + + G D+
Sbjct: 551 FTEQGDALTF-ESEVDRV------------YLDSSSMVAVLDHERKRTFVIRKEGLPDVV 691
Query: 308 VSSPGSMSDKYGRDY-------FVCTGPASILEPVTVNPGEEFRG 345
V +P K D+ +C A++ +P+T+ PGEE+ G
Sbjct: 692 VWNPWEKKSKSIVDFGDEEYKQMLCVDGAAVGKPITLKPGEEWTG 826
>TC91982 weakly similar to GP|170525|gb|AAA34202.1|| phenol hydroxylase
{Trichosporon cutaneum}, partial (15%)
Length = 1328
Score = 30.8 bits (68), Expect = 0.83
Identities = 18/46 (39%), Positives = 27/46 (58%)
Frame = -2
Query: 141 ALQVELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKPVTLTNAI 186
AL+ ELIS+ R+F+ T S+YP ++A V + +TLT I
Sbjct: 1303 ALREELISSRRYFESTQAFSMYPS*TSSARGVAYLLARKLTLTTNI 1166
>TC77223 weakly similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13
{Vigna angularis}, partial (50%)
Length = 1628
Score = 29.3 bits (64), Expect = 2.4
Identities = 19/55 (34%), Positives = 26/55 (46%)
Frame = +3
Query: 47 SNSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLY 101
S+ +PIVE T R L P+ HVT P + D + L TIP N ++
Sbjct: 57 SHLVPIVEFTKR----LVTNHPNFHVTCIIPSLGSPPDSSKSYLETIPPNINSIF 209
>TC80202 similar to GP|4512677|gb|AAD21731.1| unknown protein {Arabidopsis
thaliana}, partial (11%)
Length = 765
Score = 28.5 bits (62), Expect = 4.1
Identities = 15/36 (41%), Positives = 24/36 (66%)
Frame = -1
Query: 3 SFLSLSLPKLNLIKASSAATTTTPLSTPEALNDKFG 38
SF+S+SLP++ ++ S+ TT TP+A +DK G
Sbjct: 699 SFISISLPRVGTLQLSNGLTTFFCFITPQA-SDK*G 595
>TC77464 similar to GP|13272433|gb|AAK17155.1 unknown protein {Arabidopsis
thaliana}, partial (24%)
Length = 1109
Score = 28.1 bits (61), Expect = 5.4
Identities = 14/25 (56%), Positives = 16/25 (64%)
Frame = +2
Query: 10 PKLNLIKASSAATTTTPLSTPEALN 34
P LNLIK SA TTTT T +L+
Sbjct: 287 PFLNLIKTDSATTTTTTTPTVSSLS 361
>TC77562 similar to GP|17104733|gb|AAL34255.1 unknown protein {Arabidopsis
thaliana}, partial (75%)
Length = 1720
Score = 27.7 bits (60), Expect = 7.0
Identities = 14/40 (35%), Positives = 25/40 (62%)
Frame = +2
Query: 148 STNRFFDMTYIVSLYPVSMATAVIVKNKSPKPVTLTNAIL 187
ST+ ++ ++SL S+ T +V+NK+PKP +L + L
Sbjct: 332 STHLLRFVSRVLSLRHPSIETEQVVRNKNPKPYSLHHVTL 451
>TC83315 weakly similar to GP|19911209|dbj|BAB86931. glucosyltransferase-13
{Vigna angularis}, partial (42%)
Length = 1048
Score = 27.7 bits (60), Expect = 7.0
Identities = 18/55 (32%), Positives = 26/55 (46%)
Frame = +2
Query: 47 SNSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLY 101
S+ +PIVE + R L P+ HVT P + D + L TIP N ++
Sbjct: 17 SHLVPIVEFSKR----LVTNHPNFHVTCIIPSLGSPPDSSKSYLETIPPNINSIF 169
>AJ501243 weakly similar to PIR|A40411|A40 translation elongation factor
eEF-2 - Caenorhabditis elegans, partial (18%)
Length = 497
Score = 27.7 bits (60), Expect = 7.0
Identities = 21/69 (30%), Positives = 27/69 (38%)
Frame = -3
Query: 279 NTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRDYFVCTGPASILEPVTVN 338
NTP + +D G+E R+G E + SP S D G C G V +
Sbjct: 276 NTPNGRLRALDIGQEDRLHQSRLGRELGGIESPSSRWDDLGASSVHCVGMERYFHDVVPD 97
Query: 339 PGEEFRGAQ 347
F GAQ
Sbjct: 96 STHAF-GAQ 73
>TC77206 similar to GP|18377735|gb|AAL67017.1 putative alpha-galactosidase
{Arabidopsis thaliana}, partial (80%)
Length = 1786
Score = 27.3 bits (59), Expect = 9.2
Identities = 13/37 (35%), Positives = 20/37 (53%)
Frame = +3
Query: 269 PPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFED 305
PPK+R PP + + + GR+IFY + G +D
Sbjct: 726 PPKKRY------PPMRDALNETGRKIFYSICEWGVDD 818
>TC76691 similar to SP|O64981|RCA_PHAVU Ribulose bisphosphate
carboxylase/oxygenase activase chloroplast precursor
(RuBisCO activase) (RA)., partial (93%)
Length = 1931
Score = 27.3 bits (59), Expect = 9.2
Identities = 32/138 (23%), Positives = 57/138 (41%), Gaps = 12/138 (8%)
Frame = +1
Query: 4 FLSLSLPKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNGSSL 63
FL ++P ++ + +++ +T + TP +LN P V + G+SL
Sbjct: 250 FLFCAIP-ISAVSMAASVSTVGAVKTPMSLNGSVVAG----------PTVSNSAFFGTSL 396
Query: 64 ---SLRLPDAHVTSYKPKVFWKD---------DGLEELLYTIPANETGLYKAKGGIGLVL 111
+ RLP+ V+S KVF K+ D L Y I ++ + + KG + V
Sbjct: 397 KKLTARLPNTKVSSGSFKVFAKEIDEGKQTDGDRWRGLAYDISDDQQDITRGKGMVDSVF 576
Query: 112 NEVLQPGAKELLPSTLEW 129
G + S+ E+
Sbjct: 577 QAPQDTGTHYAVMSSYEY 630
>TC76829 similar to PIR|T14544|T14544 fructokinase (EC 2.7.1.4) - beet,
partial (95%)
Length = 1408
Score = 27.3 bits (59), Expect = 9.2
Identities = 17/34 (50%), Positives = 21/34 (61%)
Frame = -1
Query: 34 NDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRL 67
N KFG +K + S+ I+ L VRN SSLSL L
Sbjct: 769 NFKFGCHNVKAVASS---ILSLPVRNSSSLSLTL 677
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,297,024
Number of Sequences: 36976
Number of extensions: 136394
Number of successful extensions: 709
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 704
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 706
length of query: 355
length of database: 9,014,727
effective HSP length: 97
effective length of query: 258
effective length of database: 5,428,055
effective search space: 1400438190
effective search space used: 1400438190
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC145024.2


