

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.14 - phase: 0 /pseudo
(122 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
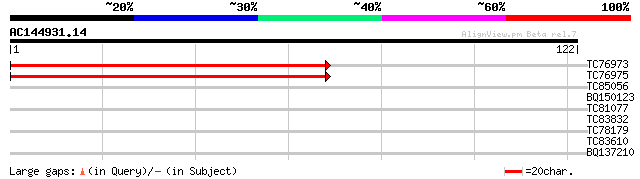
TC76973 homologue to GP|6984222|gb|AAF34799.1| 40S ribosomal pro... 144 8e-36
TC76975 homologue to GP|6984222|gb|AAF34799.1| 40S ribosomal pro... 132 2e-32
TC85056 homologue to GP|9651530|gb|AAF91178.1| ClpB {Phaseolus l... 27 2.5
BQ150123 similar to GP|21109992|gb| TonB-dependent receptor {Xan... 26 3.3
TC81077 similar to GP|19698827|gb|AAL91149.1 unknown protein {Ar... 25 5.6
TC83832 similar to GP|5042435|gb|AAD38274.1| Unknown protein {Ar... 25 5.6
TC78179 weakly similar to GP|7211427|gb|AAF40306.1| RNA helicase... 25 7.3
TC83610 similar to SP|Q9LNQ4|ALA4_ARATH Potential phospholipid-t... 25 7.3
BQ137210 similar to SP|P09789|GRP1_ Glycine-rich cell wall struc... 25 7.3
>TC76973 homologue to GP|6984222|gb|AAF34799.1| 40S ribosomal protein S16
{Euphorbia esula}, partial (96%)
Length = 665
Score = 144 bits (363), Expect = 8e-36
Identities = 69/69 (100%), Positives = 69/69 (100%)
Frame = +3
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV
Sbjct: 66 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 245
Query: 61 DMRIRVKGG 69
DMRIRVKGG
Sbjct: 246 DMRIRVKGG 272
>TC76975 homologue to GP|6984222|gb|AAF34799.1| 40S ribosomal protein S16
{Euphorbia esula}, partial (96%)
Length = 728
Score = 132 bits (333), Expect = 2e-32
Identities = 63/69 (91%), Positives = 67/69 (96%)
Frame = +2
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
+EQVQCFGRKKNAVAVT+CKRGRGLIKING PIELVEPEILRFKA+EPILLLG+HRFAGV
Sbjct: 95 LEQVQCFGRKKNAVAVTYCKRGRGLIKINGAPIELVEPEILRFKAFEPILLLGKHRFAGV 274
Query: 61 DMRIRVKGG 69
DMRIRV GG
Sbjct: 275 DMRIRVNGG 301
>TC85056 homologue to GP|9651530|gb|AAF91178.1| ClpB {Phaseolus lunatus},
partial (25%)
Length = 743
Score = 26.6 bits (57), Expect = 2.5
Identities = 14/21 (66%), Positives = 14/21 (66%)
Frame = +3
Query: 17 THCKRGRGLIKINGCPIELVE 37
THC RGRG NGC ELVE
Sbjct: 45 THCCRGRGYKWCNGC-WELVE 104
>BQ150123 similar to GP|21109992|gb| TonB-dependent receptor {Xanthomonas
axonopodis pv. citri str. 306}, partial (2%)
Length = 1131
Score = 26.2 bits (56), Expect = 3.3
Identities = 10/14 (71%), Positives = 12/14 (85%)
Frame = +1
Query: 107 EGSGGGKRYGRTRG 120
EG+GGGKR G+T G
Sbjct: 151 EGAGGGKRGGKTNG 192
>TC81077 similar to GP|19698827|gb|AAL91149.1 unknown protein {Arabidopsis
thaliana}, partial (25%)
Length = 809
Score = 25.4 bits (54), Expect = 5.6
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = +1
Query: 12 NAVAVTHCKRGRGLIKINGCPIELVEPE 39
NA++V KRG GLI+ C + L E E
Sbjct: 427 NAISVCKYKRGEGLIEGTECSVCLSEFE 510
>TC83832 similar to GP|5042435|gb|AAD38274.1| Unknown protein {Arabidopsis
thaliana}, partial (9%)
Length = 423
Score = 25.4 bits (54), Expect = 5.6
Identities = 8/18 (44%), Positives = 13/18 (71%)
Frame = +3
Query: 4 VQCFGRKKNAVAVTHCKR 21
++CF ++ VA+THC R
Sbjct: 342 IECFNQQGFIVAITHCSR 395
>TC78179 weakly similar to GP|7211427|gb|AAF40306.1| RNA helicase {Vigna
radiata}, partial (12%)
Length = 692
Score = 25.0 bits (53), Expect = 7.3
Identities = 10/14 (71%), Positives = 10/14 (71%)
Frame = +2
Query: 108 GSGGGKRYGRTRGG 121
G GGGK YG RGG
Sbjct: 302 GGGGGKGYGGGRGG 343
>TC83610 similar to SP|Q9LNQ4|ALA4_ARATH Potential phospholipid-transporting
ATPase 4 (EC 3.6.3.1) (Aminophospholipid flippase 4).,
partial (30%)
Length = 1131
Score = 25.0 bits (53), Expect = 7.3
Identities = 17/54 (31%), Positives = 23/54 (42%), Gaps = 1/54 (1%)
Frame = -2
Query: 6 CFGRKKNAVA-VTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFA 58
CF N + +T+CK G P L ++ Y+ LLLGRH A
Sbjct: 275 CFLYHANIICTITYCKSGL--------------PSSLFYQPYDQCLLLGRHTTA 156
>BQ137210 similar to SP|P09789|GRP1_ Glycine-rich cell wall structural
protein 1 precursor. [Petunia] {Petunia hybrida},
partial (13%)
Length = 990
Score = 25.0 bits (53), Expect = 7.3
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = +1
Query: 110 GGGKRYGRTRGG 121
GGGKR GR RGG
Sbjct: 7 GGGKRKGRRRGG 42
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.147 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,605,778
Number of Sequences: 36976
Number of extensions: 25117
Number of successful extensions: 108
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 108
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 108
length of query: 122
length of database: 9,014,727
effective HSP length: 84
effective length of query: 38
effective length of database: 5,908,743
effective search space: 224532234
effective search space used: 224532234
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 52 (24.6 bits)
Medicago: description of AC144931.14


