

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.10 + phase: 0 /pseudo
(416 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
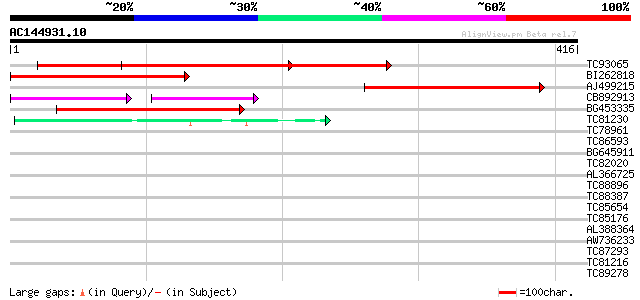
Score E
Sequences producing significant alignments: (bits) Value
TC93065 377 e-105
BI262818 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 124 5e-29
AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 110 1e-24
CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabi... 61 1e-18
BG453335 weakly similar to GP|18071369|g putative gag-pol polypr... 77 9e-15
TC81230 51 9e-07
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 35 0.041
TC86593 similar to GP|20258963|gb|AAM14197.1 putative eukaryotic... 33 0.16
BG645911 32 0.59
TC82020 similar to GP|9279662|dbj|BAB01178.1 gene_id:MIE15.7~unk... 31 0.77
AL366725 31 1.0
TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown prot... 31 1.0
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 30 1.7
TC85654 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 30 2.2
TC85176 homologue to PIR|S56655|S56655 lipoxygenase (EC 1.13.11.... 29 2.9
AL388364 similar to GP|5306274|gb|A expressed protein {Arabidops... 28 5.0
AW736233 GP|21105451|gb small nuclear ribonucleoprotein D1 {Dani... 28 5.0
TC87293 similar to GP|15292795|gb|AAK92766.1 unknown protein {Ar... 28 5.0
TC81216 weakly similar to GP|18252987|gb|AAL62420.1 putative pro... 28 6.5
TC89278 similar to GP|9757719|dbj|BAB08244.1 CRN (crooked neck) ... 28 8.5
>TC93065
Length = 783
Score = 377 bits (969), Expect = e-105
Identities = 184/198 (92%), Positives = 191/198 (95%)
Frame = +2
Query: 83 RTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLP 142
+ K+ + LRREFEALKMKETETVREFSDRISKV+TQIRLLGEDLSDQRVVEKILVCLP
Sbjct: 188 KNKENESAKLRREFEALKMKETETVREFSDRISKVVTQIRLLGEDLSDQRVVEKILVCLP 367
Query: 143 EMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKS 202
EMFEAKISSLEENKNFSEITV ELVNALQASEQRRSLRMEENVEGAFLA+NKGKNQSFKS
Sbjct: 368 EMFEAKISSLEENKNFSEITVAELVNALQASEQRRSLRMEENVEGAFLANNKGKNQSFKS 547
Query: 203 FGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHH 262
FG+KKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQ+ARVVEHH
Sbjct: 548 FGKKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQEARVVEHH 727
Query: 263 QEDEEQLFKASCYLACSS 280
QEDEE LFKASCYLACSS
Sbjct: 728 QEDEEPLFKASCYLACSS 781
Score = 150 bits (378), Expect = 1e-36
Identities = 86/188 (45%), Positives = 116/188 (60%)
Frame = +3
Query: 21 PPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQG 80
PPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNLEIAKEAW+KLMEEYQG
Sbjct: 3 PPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNLEIAKEAWSKLMEEYQG 182
Query: 81 SERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVC 140
SERTKKMKVLNL + K K+ + + F K++ + + + + +K
Sbjct: 183 SERTKKMKVLNLEESLKH*K*KKLKLLESFLTEFLKLLHKSDCWEKISVIKELWKKYWCV 362
Query: 141 LPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSF 200
+ K L++ + F + + L+ + + ++ ++ F + K K ++
Sbjct: 363 FQRCLKQKFHLLKKIRIFLKSQLQSLLMLCKLLNKEDL*GWKKMLKVPFWPTTKAKIKAL 542
Query: 201 KSFGEKKF 208
K G++ F
Sbjct: 543 KVLGKRSF 566
>BI262818 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (24%)
Length = 615
Score = 124 bits (312), Expect = 5e-29
Identities = 56/132 (42%), Positives = 90/132 (67%)
Frame = +1
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
++ YL A LW+ VE+ PL +NPT+AQI+NH + ++ +A A + AA+ ++IF +
Sbjct: 220 IEAYLEANDLWEAVEEDYEVLPLSDNPTMAQIKNHKERKTRKSKARATLFAAVSEEIFTR 399
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQ 120
I+ ++ A E WN L EY+G ER + M+ LNL REFE KMKE+ET++E+++++ + +
Sbjct: 400 IMTIKSAFEIWNFLKTEYEGDERIRGMQALNLIREFEMQKMKESETIKEYANKLISIANK 579
Query: 121 IRLLGEDLSDQR 132
+RLLG +L D R
Sbjct: 580 VRLLGSELXDSR 615
>AJ499215 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (18%)
Length = 567
Score = 110 bits (274), Expect = 1e-24
Identities = 55/132 (41%), Positives = 83/132 (62%)
Frame = +3
Query: 261 HHQEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHV 320
H +EQL S + + WLIDSGCT+HMT++ FKEL++S SKV NG H+
Sbjct: 39 HQ*GSKEQLLAMSTFATKQPSKYWLIDSGCTHHMTHDRDLFKELNKSTISKVRMLNGAHI 218
Query: 321 KVKGKGVVAVETLLGIKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGS 380
+V+G G V V++ G K I +VL+ P++NQSLLSV Q++ K Y + F++ KC I D
Sbjct: 219 EVEGIGTVLVKSHSGYKQISNVLYAPKLNQSLLSVPQLLTKGYKVLFEHEKCVIKDQNNK 398
Query: 381 KLMIVEMRGKSF 392
+++ +M+ + F
Sbjct: 399 EVLRAQMKARKF 434
>CB892913 similar to GP|1769897|emb lectin receptor kinase {Arabidopsis
thaliana}, partial (35%)
Length = 837
Score = 60.8 bits (146), Expect(2) = 1e-18
Identities = 34/89 (38%), Positives = 49/89 (54%)
Frame = +3
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M L A +W+VVEKG ++ T AQ D ++ +AL +I+ AL +D F K
Sbjct: 171 MMALLGAHDVWEVVEKGYKESQDEDSLTKAQRDTLKDSRKRDKKALFLIYQALDEDEFEK 350
Query: 61 ILNLEIAKEAWNKLMEEYQGSERTKKMKV 89
I N AKEAW KL QG ++ KK+++
Sbjct: 351 ISNATSAKEAWEKLKTSCQGEDKVKKVRL 437
Score = 50.1 bits (118), Expect(2) = 1e-18
Identities = 25/78 (32%), Positives = 43/78 (55%)
Frame = +1
Query: 105 ETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVP 164
E VR + R+ + Q++ E L D R++EKIL L FE + +EE K+ +T+
Sbjct: 472 ERVRVYFSRVISISNQLKRNSEKLEDVRIMEKILRSLDPKFEHIVEKIEETKDLETMTIE 651
Query: 165 ELVNALQASEQRRSLRME 182
+L +LQA E++ + +
Sbjct: 652 KLQGSLQAYEEKHKKKKD 705
Score = 31.6 bits (70), Expect = 0.59
Identities = 19/39 (48%), Positives = 23/39 (58%)
Frame = +2
Query: 69 EAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETV 107
EA N L QG E T NLR EFE+L MKE+E++
Sbjct: 386 EA*NLLPRRRQGKEGTSP----NLRGEFESLHMKESESI 490
>BG453335 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (11%)
Length = 660
Score = 77.4 bits (189), Expect = 9e-15
Identities = 38/138 (27%), Positives = 86/138 (61%)
Frame = +1
Query: 35 HSDEVAKEGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRR 94
H + K+ + L +I +L + F +I +KEA N L + ++G ++ +++K+ +LRR
Sbjct: 109 HKELKKKDCKELFLIQ*SLDEGNFKRISKSTRSKEASNILEKYHEGDDKVEQIKLQSLRR 288
Query: 95 EFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEE 154
+F+ ++M++ + + ++ ++ V+ QI+ GE ++DQ++VEKI+ L F+ + ++ E
Sbjct: 289 KFKLMQMEDEKKIADYISKLINVVNQIKAYGEVVTDQQIVEKIMRTLSPRFDFIVVAIHE 468
Query: 155 NKNFSEITVPELVNALQA 172
+K+ + + EL ++L+A
Sbjct: 469 SKDVKTLKIEELKSSLEA 522
>TC81230
Length = 958
Score = 50.8 bits (120), Expect = 9e-07
Identities = 54/248 (21%), Positives = 95/248 (37%), Gaps = 16/248 (6%)
Frame = +1
Query: 4 YLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEV-AKEGRALAIIHAALHDDIFIKIL 62
+L+ + LW V P + T + +E +K + + I ++
Sbjct: 283 FLKGRRLWRYVTGDKKCPTKGKDDTADAFADKLEEWDSKNHQIITWFRNTSIPSIHMQFG 462
Query: 63 NLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQIR 122
E AKE W+ L + Y S+ + + ++L ++ LK + + V EF ++ + Q+
Sbjct: 463 RFENAKEVWDHLKQRYTISDLSHQYQLL---KDLSNLKQQSGQPVYEFLAQMEVIWNQLT 633
Query: 123 LLGEDLSDQ-------------RVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNA 169
L D R+++ ++ E + SSL +N +P L NA
Sbjct: 634 SCEPSLKDATDMKTYETHRNRVRLIQFLMALTDEYEPVRASSLHQNP------LPTLENA 795
Query: 170 LQA--SEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYR 227
L SE+ R + + AF +N PC HC+K H C
Sbjct: 796 LPCLKSEETRLQLVPPKADLAFAVTNNATK------------PCRHCQKSGHSFSDC--- 930
Query: 228 PGVKCRAC 235
P ++CR C
Sbjct: 931 PTIECRNC 954
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 35.4 bits (80), Expect = 0.041
Identities = 19/53 (35%), Positives = 25/53 (46%), Gaps = 4/53 (7%)
Frame = +2
Query: 211 CPHCKKDTHLDKFC----WYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVV 259
C +C D H + C W KC C + GH+EK CKN + + AR V
Sbjct: 359 CFNCGIDGHWARDCKAGDWKN---KCYRCGERGHIEKNCKNSPKKLSRHARSV 508
>TC86593 similar to GP|20258963|gb|AAM14197.1 putative eukaryotic initiation
factor 4 {Arabidopsis thaliana}, partial (60%)
Length = 2236
Score = 33.5 bits (75), Expect = 0.16
Identities = 27/76 (35%), Positives = 36/76 (46%), Gaps = 13/76 (17%)
Frame = +2
Query: 76 EEYQGSERTKKMKVLN-----------LRREFEALKMKETETVREFSDRISKVIT--QIR 122
EE G E T K +L+ LR E + E ET R DR+ K+ T IR
Sbjct: 299 EESGGKEITFKRLLLDNCQEAFEGAGKLREELAQMTSPEQETERRDKDRLVKIRTLGNIR 478
Query: 123 LLGEDLSDQRVVEKIL 138
L+GE L + V E+I+
Sbjct: 479 LIGELLKQKMVPERIV 526
>BG645911
Length = 617
Score = 31.6 bits (70), Expect = 0.59
Identities = 21/67 (31%), Positives = 30/67 (44%)
Frame = +3
Query: 8 QSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNLEIA 67
+ LWD + +GN HSD +K G+AL I + +I N A
Sbjct: 9 KDLWDHIVEGNLDA------------QHSDWKSKNGQALHAIQLSCGHYTLGQIRNCVTA 152
Query: 68 KEAWNKL 74
+EAWN+L
Sbjct: 153QEAWNRL 173
>TC82020 similar to GP|9279662|dbj|BAB01178.1 gene_id:MIE15.7~unknown
protein {Arabidopsis thaliana}, partial (15%)
Length = 873
Score = 31.2 bits (69), Expect = 0.77
Identities = 25/90 (27%), Positives = 44/90 (48%)
Frame = +1
Query: 118 ITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRR 177
++Q L + + Q+ +K+ LP S ++EN +E PE+ N+ +++RR
Sbjct: 502 LSQSNPLDKSSTKQKGKKKLGTDLP-------SEMKENLGDAEKPDPEIPNSSSKNKERR 660
Query: 178 SLRMEENVEGAFLASNKGKNQSFKSFGEKK 207
E+ VEG + KGK +S G +K
Sbjct: 661 RRGKEKIVEGE-ESDQKGKKKSSHRHGRRK 747
>AL366725
Length = 485
Score = 30.8 bits (68), Expect = 1.0
Identities = 16/54 (29%), Positives = 23/54 (41%)
Frame = +2
Query: 193 NKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCK 246
N G+ + K+ C C K H+ C R + C CN+ GH+ CK
Sbjct: 314 NYGEKGHKSNVCPKEIKKCVRCDKKGHIVADC-KRNDIVCFNCNEEGHIGSQCK 472
>TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown protein
{Arabidopsis thaliana}, partial (27%)
Length = 1097
Score = 30.8 bits (68), Expect = 1.0
Identities = 36/139 (25%), Positives = 58/139 (40%), Gaps = 14/139 (10%)
Frame = +2
Query: 62 LNLEIAKEAWNKLMEEYQGSERT---KKMKVLNLRREFEALKMKETETVREFSDRISKVI 118
L L+ EA +E+ +G + K+ + L E EA KM E+ R D K +
Sbjct: 2 LELQTEIEALKHELEKAKGYDEKLAEKETLIEQLNVESEAAKMAESYA-RSVLDECRKKV 178
Query: 119 TQIRLLGEDLSD-QRVVEKILVCLPEMFEAK----------ISSLEENKNFSEITVPELV 167
++ + E+ + +R L + E K ISSL+E E+TV
Sbjct: 179 EELEMKVEEANQLERSASLSLETATKQLEGKNELLHDAESEISSLKEKLGMLEMTVGRQR 358
Query: 168 NALQASEQRRSLRMEENVE 186
L+ +E+ EEN+E
Sbjct: 359 GDLEDAERCLLAAKEENIE 415
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 30.0 bits (66), Expect = 1.7
Identities = 14/35 (40%), Positives = 18/35 (51%)
Frame = +1
Query: 211 CPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVC 245
C +C+K HL + C P C CN GHV + C
Sbjct: 730 CNNCRKTGHLARDCPNDP--ICNLCNISGHVARQC 828
>TC85654 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (32%)
Length = 1307
Score = 29.6 bits (65), Expect = 2.2
Identities = 26/86 (30%), Positives = 40/86 (46%), Gaps = 1/86 (1%)
Frame = +3
Query: 209 PPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQ 268
P P +D HL+ +C +K A +G +EK K KTN+ + H +++
Sbjct: 531 PSIPKPFRD-HLETYCLE---LKQLAFTIIGRMEKTLKIKTNELVEFFEDAIHQGNEDKL 698
Query: 269 LFKASCYLACS-SKETWLIDSGCTNH 293
L S AC+ SK ++ GC NH
Sbjct: 699 LSSMSTTRACNWSKSSF*Y--GCLNH 770
>TC85176 homologue to PIR|S56655|S56655 lipoxygenase (EC 1.13.11.12) loxG -
garden pea, partial (19%)
Length = 663
Score = 29.3 bits (64), Expect = 2.9
Identities = 15/53 (28%), Positives = 29/53 (54%)
Frame = +1
Query: 154 ENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEK 206
+N +++T+ E+++ + EQ + E +EG S+ +SFK FG+K
Sbjct: 205 KNDTLTDLTIIEVLSRHASDEQY----LGERIEGDLWTSDSQPKESFKRFGKK 351
>AL388364 similar to GP|5306274|gb|A expressed protein {Arabidopsis
thaliana}, partial (4%)
Length = 524
Score = 28.5 bits (62), Expect = 5.0
Identities = 17/56 (30%), Positives = 25/56 (44%)
Frame = +1
Query: 197 NQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQ 252
++S K G F C +D YR G K + C Q+GH+ C + T Q+
Sbjct: 97 SRSCKKLGAYNFEGCNKSS**EDIDML--YR*G*KQQKCTQIGHLVATCSDTTIQE 258
>AW736233 GP|21105451|gb small nuclear ribonucleoprotein D1 {Danio rerio},
partial (14%)
Length = 510
Score = 28.5 bits (62), Expect = 5.0
Identities = 17/68 (25%), Positives = 30/68 (44%)
Frame = +1
Query: 230 VKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSG 289
++C+ C + H C + +Q HH+ + +S Y S+ W DSG
Sbjct: 292 LQCQICARNNHDAARCCFRYDQASSSQA---HHRAPPSEYAASSSY----SEAPWYPDSG 450
Query: 290 CTNHMTNN 297
++H+T N
Sbjct: 451 ASHHLTFN 474
>TC87293 similar to GP|15292795|gb|AAK92766.1 unknown protein {Arabidopsis
thaliana}, partial (29%)
Length = 1101
Score = 28.5 bits (62), Expect = 5.0
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = -2
Query: 366 HFKNMKCTIFDPYGSKLMIVEMRGKSFPRGVEEDILACFSK 406
HF+ + + F P S L+I + G+SFP ACFSK
Sbjct: 947 HFQKLPVSFFIP--SILLITKPGGESFPSS*ARTSQACFSK 831
>TC81216 weakly similar to GP|18252987|gb|AAL62420.1 putative protein
{Arabidopsis thaliana}, partial (44%)
Length = 1159
Score = 28.1 bits (61), Expect = 6.5
Identities = 30/139 (21%), Positives = 58/139 (41%), Gaps = 22/139 (15%)
Frame = +2
Query: 13 VVEKGNNPPPLPNNPTLAQIRNHSDEVAKEG--------------RALAIIHAALHD-DI 57
VV + L N+ +A + NH D VAKE +A+A HD D
Sbjct: 83 VVSSSDGTDNLQNDAAVADVANHIDTVAKEEVKSDASRDEVVRILQAIASTGKFWHDWDN 262
Query: 58 FIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSD----- 112
+L+ ++ +++ EY ++ T + + +L + L K E + F D
Sbjct: 263 LKSMLSFQL-----KQVLSEYPEAKLTSEQQCASLGESYSDLVNKLDEALTCFIDGPPFT 427
Query: 113 --RISKVITQIRLLGEDLS 129
R+ +++ + + + +LS
Sbjct: 428 LQRVCEILLEAKNIYPNLS 484
>TC89278 similar to GP|9757719|dbj|BAB08244.1 CRN (crooked neck) protein
{Arabidopsis thaliana}, partial (64%)
Length = 1612
Score = 27.7 bits (60), Expect = 8.5
Identities = 33/120 (27%), Positives = 49/120 (40%), Gaps = 12/120 (10%)
Frame = +1
Query: 82 ERTKKMKVLNLRREFEA------LKMKETETVREFSDRISKV----ITQIRLLGEDLSDQ 131
+RTK +KV EFEA L++ E E + R KV + R DL ++
Sbjct: 1009 DRTKHLKVWQSYAEFEATAIDESLELSEQEQKEQCLQRARKVFEDALNHFRSSAPDLKEE 1188
Query: 132 R--VVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAF 189
R ++EK L EA L + +L Q S + S R+EE ++ F
Sbjct: 1189 RAMLLEKWL-----NLEASSGELGDVSLLQSKLPKKLKKRRQVSTEDGSSRIEEFIDYLF 1353
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,355,093
Number of Sequences: 36976
Number of extensions: 191121
Number of successful extensions: 1085
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 1069
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1084
length of query: 416
length of database: 9,014,727
effective HSP length: 99
effective length of query: 317
effective length of database: 5,354,103
effective search space: 1697250651
effective search space used: 1697250651
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144931.10


