

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144893.5 - phase: 0 /pseudo
(966 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
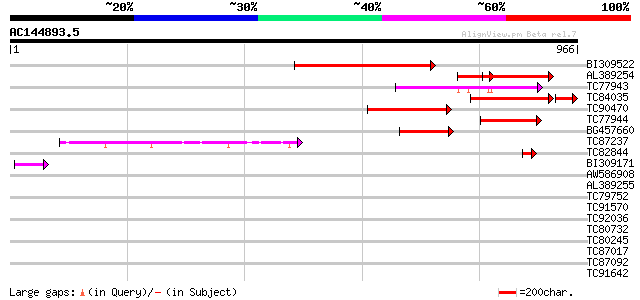
Score E
Sequences producing significant alignments: (bits) Value
BI309522 similar to PIR|A86198|A86 hypothetical protein [importe... 285 6e-77
AL389254 weakly similar to GP|7523514|dbj| Similar to Arabidopsi... 227 1e-59
TC77943 similar to GP|10179602|gb|AAG13810.1 PSTVd RNA-binding p... 213 4e-55
TC84035 similar to PIR|A86198|A86198 hypothetical protein [impor... 175 3e-51
TC90470 weakly similar to GP|9294219|dbj|BAB02121.1 gb|AAF01563.... 127 2e-29
TC77944 similar to GP|10179602|gb|AAG13810.1 PSTVd RNA-binding p... 103 2e-22
BG457660 similar to PIR|T00472|T00 probable RING3 protein [impor... 100 2e-21
TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-inf... 54 2e-07
TC82844 similar to PIR|A86198|A86198 hypothetical protein [impor... 45 2e-04
BI309171 43 5e-04
AW586908 weakly similar to GP|2104814|emb| pleiotropic effects o... 42 0.001
AL389255 similar to GP|15810439|gb| unknown protein {Arabidopsis... 40 0.003
TC79752 similar to GP|11994670|dbj|BAB02898. gene_id:K7P8.17~unk... 40 0.003
TC91570 weakly similar to GP|7208782|emb|CAB76913.1 hypothetical... 40 0.003
TC92036 homologue to EGAD|58716|61404 groucho-related gene 4 pro... 39 0.007
TC80732 similar to GP|21553690|gb|AAM62783.1 unknown {Arabidopsi... 39 0.007
TC80245 similar to PIR|T48581|T48581 hypothetical protein T31B5.... 39 0.010
TC87017 similar to PIR|A96564|A96564 unknown protein 23094-2177... 38 0.016
TC87092 similar to PIR|D84900|D84900 hypothetical protein At2g46... 37 0.028
TC91642 similar to PIR|E96613|E96613 hypothetical protein T15M6.... 36 0.062
>BI309522 similar to PIR|A86198|A86 hypothetical protein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 729
Score = 285 bits (729), Expect = 6e-77
Identities = 149/243 (61%), Positives = 178/243 (72%), Gaps = 2/243 (0%)
Frame = +1
Query: 485 YRHSENVTEEPNLEDRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGY 544
+ H+ ++ LE+ +K LAS SKQE +E+ KLE +L+ VRSLV RIE KQ G +
Sbjct: 1 FEHALRCNDKKELENGVKNGLASRSKQEMREIKRKLEGDLETVRSLVIRIEEKQRMAGRF 180
Query: 545 GNS-NVVLGG-GITNGGGAQRAHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASEGVEKE 602
G NV + + NG GA+RAHSE S GV R+ R QLS + +N SE VEKE
Sbjct: 181 GGGLNVSVDRYRVDNGSGAKRAHSEVASAGVPREANRFTRQLSVMVLENGHGISESVEKE 360
Query: 603 KRMPKANQFYHNSEFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSL 662
KR PKANQFY NSEFLLA DKFPPAESNKKSK + KK G G+MG G + SK+ K+CSSL
Sbjct: 361 KRTPKANQFYSNSEFLLAKDKFPPAESNKKSKSHGKKHGVGEMGHGFGIVSKYLKNCSSL 540
Query: 663 LEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVR 722
LEKLM+H++ WVFNTPVDV+ LGLHDYF II+ PMDLGTVK+RL+KNWYKSPKEFAEDVR
Sbjct: 541 LEKLMKHKHGWVFNTPVDVERLGLHDYFIIISQPMDLGTVKSRLSKNWYKSPKEFAEDVR 720
Query: 723 LTF 725
LTF
Sbjct: 721 LTF 729
>AL389254 weakly similar to GP|7523514|dbj| Similar to Arabidopsis thaliana
chromosome II BAC F19I3 genomic sequence; putative RING3
protein, partial (18%)
Length = 491
Score = 227 bits (579), Expect = 1e-59
Identities = 116/121 (95%), Positives = 117/121 (95%)
Frame = +3
Query: 806 RSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL* 865
RSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSL SLPVEKLDAVVQMMRKKNL
Sbjct: 129 RSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSLQSLPVEKLDAVVQMMRKKNLE 308
Query: 866 LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAEQARERAEALQNSVQSSRL 925
LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRK EQARERAEALQNSVQSS+
Sbjct: 309 LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKGEQARERAEALQNSVQSSQP 488
Query: 926 P 926
P
Sbjct: 489 P 491
Score = 92.4 bits (228), Expect = 7e-19
Identities = 45/62 (72%), Positives = 47/62 (75%)
Frame = +2
Query: 764 MRYGMDYGAPSPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPK 823
MRYGMDYGAPSPLSRRVPAFTPP PLDMRRILDRQEPFARTP+ P K+ K
Sbjct: 2 MRYGMDYGAPSPLSRRVPAFTPPAPLDMRRILDRQEPFARTPKVYE*YSIK*NPGSKEAK 181
Query: 824 AK 825
K
Sbjct: 182 GK 187
>TC77943 similar to GP|10179602|gb|AAG13810.1 PSTVd RNA-binding protein
Virp1a {Lycopersicon esculentum}, partial (43%)
Length = 1783
Score = 213 bits (541), Expect = 4e-55
Identities = 118/277 (42%), Positives = 163/277 (58%), Gaps = 26/277 (9%)
Frame = +3
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K+C +L+KLM+ + W+F++PVD L LHDYF II +PMDLGTVK++L KN Y +P E
Sbjct: 645 KACGQILQKLMKTKIGWIFSSPVDPVALNLHDYFDIIKHPMDLGTVKSKLAKNAYSTPAE 824
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNRE------MRYGMDY 770
FA+DV+LTF NA+TYNPKG DV+ A QL + FE+ + I+ ++ + ++
Sbjct: 825 FADDVKLTFKNALTYNPKGHDVNTAAMQLLEKFEELYRPIQEKFDEKSFDDELQASSWNH 1004
Query: 771 GAPSPLSRRV-----PAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSS----RTPAP-- 819
P RV P PPP + +L + P + N P + RTP+P
Sbjct: 1005VEPERERERVKKKDNPIPIPPPVAKRQELLPEPASTSNQPSTSNPPPLAQSPVRTPSPTR 1184
Query: 820 ---------KKPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQD 870
KPKA+DPNKR+M +EK KL L LP EK++ VVQ++RK+N L Q
Sbjct: 1185ALPVKPLKQPKPKARDPNKREMNVEEKHKLGLGLQILPPEKMEQVVQIIRKRNGHLEQDG 1364
Query: 871 DEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAE 907
DEIE+D +AVD E LWELDR V N+KK ++ K +
Sbjct: 1365DEIELDMEAVDTETLWELDRLVTNWKKMELPDREKVD 1475
>TC84035 similar to PIR|A86198|A86198 hypothetical protein [imported] -
Arabidopsis thaliana, partial (18%)
Length = 768
Score = 175 bits (444), Expect(2) = 3e-51
Identities = 93/143 (65%), Positives = 112/143 (78%), Gaps = 1/143 (0%)
Frame = +2
Query: 785 PPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSL 844
P PPLDMRRI DR + + P M+ T S+RTPAPKKPKA DP+KRDMTY EKQKLST L
Sbjct: 8 PAPPLDMRRI-DRSDSITKPPNPMSLTRSARTPAPKKPKANDPDKRDMTYYEKQKLSTHL 184
Query: 845 HSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKR 904
+LP +KLDA+VQ+++K+N L+Q DDEIEVD D+VDAE LWELDR+V NYKKSLSK KR
Sbjct: 185 QNLPPDKLDAIVQIIKKQNSALSQHDDEIEVDIDSVDAETLWELDRYVTNYKKSLSKYKR 364
Query: 905 KAEQA-RERAEALQNSVQSSRLP 926
KAE A + RAEA + + Q S+ P
Sbjct: 365 KAELAIQARAEAERIAQQKSQEP 433
Score = 46.2 bits (108), Expect(2) = 3e-51
Identities = 25/40 (62%), Positives = 29/40 (72%), Gaps = 3/40 (7%)
Frame = +3
Query: 930 QMKEMFPHPCLCKGEVKQIIRVDQVV---QAVILGLLQVV 966
QMKEM PHPCLCKG+ + I+ V QVV QAVIL L V+
Sbjct: 468 QMKEMIPHPCLCKGKFE*IMGVRQVVQAAQAVILALHLVI 587
>TC90470 weakly similar to GP|9294219|dbj|BAB02121.1
gb|AAF01563.1~gene_id:K17E12.8~similar to unknown
protein {Arabidopsis thaliana}, partial (11%)
Length = 772
Score = 127 bits (319), Expect = 2e-29
Identities = 65/147 (44%), Positives = 91/147 (61%), Gaps = 3/147 (2%)
Frame = +3
Query: 610 QFYHNSEFLLANDKFPPAESNKKSKLNWKKQGG---GDMGXGLQMGSKFFKSCSSLLEKL 666
QF L N + P + + N ++ G G ++ K C+++L+ L
Sbjct: 321 QFGEKVSQLDENGRCNPIKKSSLLGSNKREPSGNIEGPKNKRQKLDRKGSHKCATILKCL 500
Query: 667 MEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFH 726
+ H Y+WVF TPVD L + DYFT+I++PMDLGT+K +L+KN Y S +EFA DVRLTF
Sbjct: 501 ISHPYSWVFKTPVDPVALNIPDYFTVISHPMDLGTIKFKLDKNIYYSKEEFAADVRLTFS 680
Query: 727 NAMTYNPKGQDVHAMAEQLSKIFEDRW 753
NAMTYNP DVH MA++L+K+FE +W
Sbjct: 681 NAMTYNPPSNDVHLMAKELNKLFERKW 761
>TC77944 similar to GP|10179602|gb|AAG13810.1 PSTVd RNA-binding protein
Virp1a {Lycopersicon esculentum}, partial (27%)
Length = 1144
Score = 103 bits (258), Expect = 2e-22
Identities = 55/104 (52%), Positives = 68/104 (64%)
Frame = +1
Query: 803 RTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKK 862
R P M P PK PKA+DPNKR+M +EK KL L LP EK++ VVQ++RK+
Sbjct: 13 RLPSPMRALPVKPLKQPK-PKARDPNKREMNVEEKHKLGLGLQILPPEKMEQVVQIIRKR 189
Query: 863 NL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKA 906
N L Q DEIE+D +A D E LWELDR V N+KK +SK KR+A
Sbjct: 190 NGHLEQDGDEIELDMEAGDTETLWELDRLVTNWKKMVSKIKRQA 321
>BG457660 similar to PIR|T00472|T00 probable RING3 protein [imported] -
Arabidopsis thaliana, partial (34%)
Length = 654
Score = 100 bits (250), Expect = 2e-21
Identities = 46/95 (48%), Positives = 65/95 (68%), Gaps = 3/95 (3%)
Frame = +3
Query: 665 KLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRL---NKNWYKSPKEFAEDV 721
++ +H++AW F PVDV+GLGLHDY+ II PMD GT+K ++ + YK+ +E DV
Sbjct: 93 QITQHKWAWPFLEPVDVEGLGLHDYYEIIDKPMDFGTIKNKMEAKDGTGYKNVREIYADV 272
Query: 722 RLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAII 756
RL F NAM YN + DVH MA+ L + FED+W ++
Sbjct: 273 RLIFKNAMKYNNEKHDVHVMAKTLMEKFEDKWLLL 377
>TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-infected
erythrocyte surface antigen {Plasmodium falciparum},
partial (2%)
Length = 2007
Score = 54.3 bits (129), Expect = 2e-07
Identities = 78/451 (17%), Positives = 182/451 (40%), Gaps = 36/451 (7%)
Frame = +1
Query: 85 SSRLEDGSSIQQQLGDQNLVEEHASSRTGDRNSPQQQLEELNSAQPHASFQLADGNSPQQ 144
+ ++DG IQQ+ ++ +E S + + N +++ ++ N + + + G ++
Sbjct: 169 NEEIKDGEKIQQE--NEENKDEEKSQQENEENKDEEKSQQENELKKNEGGEKETGEITEE 342
Query: 145 --KVENQDSAHPQASLRIGD-----GNSPQQQLVEPNLAQPQASLRAGDGNSPQRQLEEE 197
K EN++++ + + + G+ ++Q+ E + + +G + E +
Sbjct: 343 KSKQENEETSETNSKDKENEESNQNGSDAKEQVGENHEQDSKQGTEETNGTEGGEKEEHD 522
Query: 198 NSAQPQAS-SRTGDGNLPHQQLEDQYSAQPPASLRIGDGNSLR----------------- 239
+ +S ++ DG ++ E+ YS AS + D S
Sbjct: 523 KIKEDTSSDNQVQDGEKNNEAREENYSGD-NASSAVVDNKSQESSNKTEEQFDKKEKNEF 699
Query: 240 --QKVEGHNLVQPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNLAQ 297
+ + N S I + + +DQ+ + + Q+Q ++QN
Sbjct: 700 ELESQKNSNETTESTDSTITQNSQGNESEKDQAQTENDTPKGSASESDEQKQEQEQN-NT 876
Query: 298 THVSSRTEDVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEVNSRLEDRSFPQPELNS 357
T +T D +S ++T+K ++ + NS+ ED+S N+ + +S
Sbjct: 877 TKDDVQTTDTSSQNGNDTTEKQNETSED-ANSKKEDSSALNTTPNNEDSKSGVAGDQADS 1053
Query: 358 KLEDMTSQQQDNSI----LEDGNSSQPQLNLRLEGSSLQPVANSTSEDLNLAQPLSQSVS 413
+S+ QD + +D + P+ N EG+ +++T ++ + A S
Sbjct: 1054TTTTSSSETQDGNTNHGEYKDTTNENPEKNSGQEGTQESGSSSNTFDNKDAA-------S 1212
Query: 414 DDLHINQQAEPSNLNVRQEDDRPSSPIHRQGAISDNLHSHQQVEPSNPNVLLEDDGPSSP 473
+ + + ++ S + +++D+ S+ + + +DN +S Q S+ + D+ SS
Sbjct: 1213NKVQLTTTSDTS--SEQKKDESSSAESKSESSQNDNANSGQSNTTSDESA--NDNKDSSQ 1380
Query: 474 I-----HRHKAVPSTEYRHSENVTEEPNLED 499
+ + + +TE EN + N E+
Sbjct: 1381VTTSSENSAEGNSNTENNSDENQNDSKNNEN 1473
Score = 38.5 bits (88), Expect = 0.013
Identities = 39/168 (23%), Positives = 67/168 (39%), Gaps = 9/168 (5%)
Frame = +1
Query: 41 KDENIPTTIVTSDTDNNNG-------TTLENIDEAPEFAVLGDGNLAKAEGSSRLEDGSS 93
KDE+ + + N+N T+ E+ ++ + + + + AEG+S E+ S
Sbjct: 1261 KDESSSAESKSESSQNDNANSGQSNTTSDESANDNKDSSQVTTSSENSAEGNSNTENNSD 1440
Query: 94 IQQQLGDQNLVEEHASSRTGDRNSPQQQLEELNSAQPHASFQLAD--GNSPQQKVENQDS 151
Q N + + + D N + Q E N+AQ S D S + K EN +S
Sbjct: 1441 ENQNDSKNNENTNDSGNTSNDANVNENQNE--NAAQTKTSENEGDAQNESVESKKENNES 1614
Query: 152 AHPQASLRIGDGNSPQQQLVEPNLAQPQASLRAGDGNSPQRQLEEENS 199
AH + NS Q + ++ Q R G P+ E ++
Sbjct: 1615 AHKDVD---NNSNSNDQGSSDTSVTQDDKESRVDLGTLPESNGESHHN 1749
>TC82844 similar to PIR|A86198|A86198 hypothetical protein [imported] -
Arabidopsis thaliana, partial (9%)
Length = 834
Score = 44.7 bits (104), Expect = 2e-04
Identities = 19/24 (79%), Positives = 21/24 (87%)
Frame = +2
Query: 874 EVDFDAVDAEILWELDRFVLNYKK 897
EVD D+VDAE LWELDR+V NYKK
Sbjct: 38 EVDIDSVDAETLWELDRYVTNYKK 109
Score = 39.7 bits (91), Expect = 0.006
Identities = 22/41 (53%), Positives = 26/41 (62%), Gaps = 4/41 (9%)
Frame = +1
Query: 930 QMKEMFPHPCLCKGEVKQII----RVDQVVQAVILGLLQVV 966
QMKEM P P LCKG+ + II +V Q QAVI L QV+
Sbjct: 235 QMKEMLPRPWLCKGKFESIIMGIRQVAQAAQAVIPALQQVI 357
>BI309171
Length = 547
Score = 43.1 bits (100), Expect = 5e-04
Identities = 27/61 (44%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Frame = +2
Query: 9 VREIQRYSEGKVYRRKVFKGIKINPNIEETVLKDENIPTT---IVTSDTDNNNGTTLENI 65
VREIQRYSEGKVY RK FK + N E V + TT V DN N +++ +
Sbjct: 365 VREIQRYSEGKVYTRKNFKVSEDKVNFLEVVPHTHSGATTATVTVDGTKDNGNNVSVQLL 544
Query: 66 D 66
D
Sbjct: 545 D 547
>AW586908 weakly similar to GP|2104814|emb| pleiotropic effects on cellular
differentiation and slug behaviour {Dictyostelium
discoideum}, partial (8%)
Length = 668
Score = 42.0 bits (97), Expect = 0.001
Identities = 44/180 (24%), Positives = 74/180 (40%), Gaps = 1/180 (0%)
Frame = +2
Query: 87 RLEDGSSIQQQL-GDQNLVEEHASSRTGDRNSPQQQLEELNSAQPHASFQLADGNSPQQK 145
+L+ QQQL G QNL + + SPQQ ++ QP + + +G +
Sbjct: 80 QLQQQQQRQQQLAGSQNLPQPQQQQPQLTQQSPQQPQQQQQHQQPCQNTTMNNGTIGSNQ 259
Query: 146 VENQDSAHPQASLRIGDGNSPQQQLVEPNLAQPQASLRAGDGNSPQRQLEEENSAQPQAS 205
+ NQ P ++ QQQL+ ++ Q AG L +++ Q Q
Sbjct: 260 IPNQCVQQPVTYSQL------QQQLLSGSMQSQQNLQSAGKNGLMMTSLPQDSQFQQQID 421
Query: 206 SRTGDGNLPHQQLEDQYSAQPPASLRIGDGNSLRQKVEGHNLVQPQASSIIEDGNSPRQK 265
+ G L QQ ++Q +SL++ L+Q + QPQ S + S +Q+
Sbjct: 422 QQQA-GLLQRQQQQNQLQ---QSSLQL-----LQQSMLQRGPQQPQMSQTVPQNISDQQQ 574
Score = 38.9 bits (89), Expect = 0.010
Identities = 56/249 (22%), Positives = 90/249 (35%), Gaps = 5/249 (2%)
Frame = +2
Query: 129 QPHASFQLADGNSPQQKVENQDSAHPQASLRIGDGNSPQQQLV-EPNLAQPQASLRAGDG 187
QP AS+ + QQ+ + S P ++ QQQL NL QPQ
Sbjct: 5 QPPASW-----SQQQQQQQKMQSLLPMPLDQLQQQQQRQQQLAGSQNLPQPQQQQPQLTQ 169
Query: 188 NSPQRQLEEENSAQPQASSRTGDGNLPHQQLEDQYSAQPPASLRIGDGNSLRQKVEGHNL 247
SPQ+ +++ QP ++ +G + Q+ +Q QP ++ +Q + G
Sbjct: 170 QSPQQPQQQQQHQQPCQNTTMNNGTIGSNQIPNQCVQQPVTYSQL-----QQQLLSGSMQ 334
Query: 248 VQPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNLAQTHVSSRTEDV 307
Q S ++G +D QQQ++ Q
Sbjct: 335 SQQNLQSAGKNGLMMTSLPQDSQF---------------QQQIDQQQAGLLQ-------- 445
Query: 308 NSPRPQNSTQKDGDSPQILENSR----PEDASLTQPEVNSRLEDRSFPQPELNSKLEDMT 363
R Q Q S Q+L+ S P+ ++Q V + D+ Q +L KL+
Sbjct: 446 ---RQQQQNQLQQSSLQLLQQSMLQRGPQQPQMSQ-TVPQNISDQQ-QQLQLLQKLQQQQ 610
Query: 364 SQQQDNSIL 372
QQQ L
Sbjct: 611 QQQQQQQPL 637
>AL389255 similar to GP|15810439|gb| unknown protein {Arabidopsis thaliana},
partial (6%)
Length = 522
Score = 40.4 bits (93), Expect = 0.003
Identities = 20/21 (95%), Positives = 21/21 (99%)
Frame = +2
Query: 946 KQIIRVDQVVQAVILGLLQVV 966
KQIIRVDQVVQAVILGLLQV+
Sbjct: 2 KQIIRVDQVVQAVILGLLQVI 64
>TC79752 similar to GP|11994670|dbj|BAB02898. gene_id:K7P8.17~unknown
protein {Arabidopsis thaliana}, partial (2%)
Length = 2137
Score = 40.4 bits (93), Expect = 0.003
Identities = 52/198 (26%), Positives = 76/198 (38%), Gaps = 21/198 (10%)
Frame = +1
Query: 95 QQQLGDQNLVEEHASSRTGDRNSPQQQLEELNSAQPHASFQLADG-NSPQQKVENQDSAH 153
QQQ + E + R QQQL N+ PH Q +SPQQ N A
Sbjct: 721 QQQAYIRLAKERQLQQQQQQRYLQQQQLSATNALIPHVQAQAQPPISSPQQ---NSSQAQ 891
Query: 154 P-----QASLRIGDGNSP------QQQLVEPNLAQPQASLRAGDGNSPQRQLEEENSAQP 202
P Q S+ +SP Q Q + +L QP S G + Q + + AQ
Sbjct: 892 PQNSSQQVSVSPATPSSPLTPMSSQHQQQKHDLPQPGFSRNPGSSVTSQAVKQRQRQAQQ 1071
Query: 203 QASSRTGDGNLPHQQLEDQYSAQPPASLRIGDGNSLRQKVEGHNLVQPQ---------AS 253
+ ++ + P+Q Q Q IG GN+L + +N V P +
Sbjct: 1072RQYQQSARQH-PNQPQHAQAQQQAKILKGIGRGNTL---IHQNNSVDPSHINGLSVAPGT 1239
Query: 254 SIIEDGNSPRQKTEDQSL 271
+E G+ Q T+ Q+L
Sbjct: 1240QPVEKGDQITQMTQGQTL 1293
Score = 36.2 bits (82), Expect = 0.062
Identities = 50/195 (25%), Positives = 75/195 (37%), Gaps = 15/195 (7%)
Frame = +1
Query: 214 PHQQLEDQYSAQPPASLRIGDGNSLRQKVEGHNLVQPQASSIIEDGNSPRQKTEDQSLAQ 273
PH Q + + Q A +R+ L+Q+ + L Q Q S+ + P Q+ AQ
Sbjct: 688 PHLQGPNHPNNQQQAYIRLAKERQLQQQQQQRYLQQQQLSAT--NALIPHV----QAQAQ 849
Query: 274 PPVSSRTDDGDSAQQQVEDQNLAQTHVSSRT-EDVNSPRPQNSTQKDGDSPQILENSRPE 332
PP+SS + AQ Q Q Q VS T +P Q+ D PQ SR
Sbjct: 850 PPISSPQQNSSQAQPQNSSQ---QVSVSPATPSSPLTPMSSQHQQQKHDLPQ-PGFSRNP 1017
Query: 333 DASLTQPEVNSR---LEDRSFPQPELNSKLEDMTSQQQD-----------NSILEDGNSS 378
+S+T V R + R + Q + +Q Q N+++ NS
Sbjct: 1018GSSVTSQAVKQRQRQAQQRQYQQSARQHPNQPQHAQAQQQAKILKGIGRGNTLIHQNNSV 1197
Query: 379 QPQLNLRLEGSSLQP 393
P + G S+ P
Sbjct: 1198DPS---HINGLSVAP 1233
>TC91570 weakly similar to GP|7208782|emb|CAB76913.1 hypothetical protein
{Cicer arietinum}, partial (17%)
Length = 1446
Score = 40.4 bits (93), Expect = 0.003
Identities = 60/272 (22%), Positives = 115/272 (42%), Gaps = 20/272 (7%)
Frame = +2
Query: 191 QRQLEEENSAQPQASSRTG---DGNLPHQQLEDQYSAQPPASLR--IGDGNSLRQKVEGH 245
QR+ EEN+ QP+ +S G + L + LE++ + +P SL D SL QK
Sbjct: 92 QRKYLEENNDQPKKNSNGGAKEEKVLGKKTLEEK-NDKPKKSLNGSAKDEKSLEQKTLEE 268
Query: 246 NLVQPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNLAQTHVSSRTE 305
N +P+ S +G++ +K+ Q + ++ D ++ + + + +T+
Sbjct: 269 NNDKPKKSL---NGSAKEEKSLGQKTPE-------ENNDKPKKSLNGGVKEEKDLGQKTQ 418
Query: 306 DVNSPRPQNS--------------TQKDGDSP-QILENSRPEDASLTQPEVNSRLEDRSF 350
+ N+ +P+ S TQ++ D P + L E+ L Q ++ E+
Sbjct: 419 EENNDKPKKSLNGGVKEEKDLGQKTQENNDKPKKSLSGGAKEEKGLGQ---KAQEENNGK 589
Query: 351 PQPELNSKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPVANSTSEDLNLAQPLSQ 410
P+ LN ++ +Q LE + P L+ + SEDLN
Sbjct: 590 PKKSLNGSAKEEKGLRQ--KALEQEGHNDPNAVLQTSMKKVGDEEIGVSEDLN----GGD 751
Query: 411 SVSDDLHINQQAEPSNLNVRQEDDRPSSPIHR 442
+ + + + + P+N N QE++ ++P R
Sbjct: 752 KIISNNYQRRASLPANFN-DQENEIHNTPSPR 844
>TC92036 homologue to EGAD|58716|61404 groucho-related gene 4 protein (Grg4)
{Mus musculus}, partial (3%)
Length = 1046
Score = 39.3 bits (90), Expect = 0.007
Identities = 46/209 (22%), Positives = 92/209 (44%), Gaps = 2/209 (0%)
Frame = -2
Query: 242 VEGHNLVQPQASSIIEDGNSPRQKTE-DQSLAQPPVSSRTDDGDSAQQQVEDQN-LAQTH 299
V NL QP + ++ D R TE D + P+SS T++G+ + + + ++
Sbjct: 979 VSEENLDQPTETQMVSD-EMQRDHTEIDNQQSTLPLSSETEEGEMLPEAGDPEGGFDGSN 803
Query: 300 VSSRTEDVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEVNSRLEDRSFPQPELNSKL 359
+ ++ +P P + D D+ + E + PE ++ + + ED + +L + +
Sbjct: 802 MENQESREATPEPSPARVDDDDALEAGEINSPEISTDDKNDEGDLAEDAADGSDKL-ADV 626
Query: 360 EDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPVANSTSEDLNLAQPLSQSVSDDLHIN 419
TS + D + +P PVA+ + NL +++S S L ++
Sbjct: 625 NKATSVESDQVV-------EPA-----------PVASES----NLQSSVAESSSSKLPVS 512
Query: 420 QQAEPSNLNVRQEDDRPSSPIHRQGAISD 448
+Q + + + ED +P+SP+ ISD
Sbjct: 511 KQGATRSPS-KTEDVKPTSPVKPTSPISD 428
>TC80732 similar to GP|21553690|gb|AAM62783.1 unknown {Arabidopsis
thaliana}, partial (16%)
Length = 760
Score = 39.3 bits (90), Expect = 0.007
Identities = 34/132 (25%), Positives = 54/132 (40%), Gaps = 15/132 (11%)
Frame = +2
Query: 304 TEDVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPE-------VNSRLEDRSFPQPELN 356
+++ + P P+ +Q P+ N P+ T+PE V E S P PE+N
Sbjct: 137 SQNESQPSPEPQSQNGAPEPESNSNHEPDPQPETEPEQPESQEQVQLEPESGSRPDPEVN 316
Query: 357 SKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSS-------LQPVAN-STSEDLNLAQPL 408
S + + N ++ +S PQL + EGS L + N S E +++ P
Sbjct: 317 SDADLKETAINSNDVVGTNSSPHPQLK-KDEGSRTFTMRELLHGLKNDSEPEREDVSSPY 493
Query: 409 SQSVSDDLHINQ 420
SQ H Q
Sbjct: 494 SQDSQQQQHTEQ 529
>TC80245 similar to PIR|T48581|T48581 hypothetical protein T31B5.160 -
Arabidopsis thaliana, partial (60%)
Length = 895
Score = 38.9 bits (89), Expect = 0.010
Identities = 42/160 (26%), Positives = 75/160 (46%), Gaps = 7/160 (4%)
Frame = +2
Query: 772 APSPLSRRVPAFTPPPPLDMRRILD--RQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNK 829
+PS R + +P P RR R+ +R+ S++ P SRTP K K P
Sbjct: 83 SPSYRRRHSRSRSPSPSPSHRRSRHHRRRRHRSRSSSSLSPPPRSRTPTLKHNKKDHPPT 262
Query: 830 RDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWEL- 888
+ ++Q+ L L E + + +R KN+ +E++V+ + AE + +L
Sbjct: 263 LNNKSSKRQQ-DEELKLLEEETARRIEEAIR-KNVEEKLNSEEVKVEIERRVAEGVKKLF 436
Query: 889 DRFVLNYKK----SLSKNKRKAEQARERAEALQNSVQSSR 924
D + +K +L++ +RK EQAR+ E L ++ +R
Sbjct: 437 DDVEVQLEKEKQDALTEARRKEEQARKEREELDKMLEENR 556
>TC87017 similar to PIR|A96564|A96564 unknown protein 23094-21772
[imported] - Arabidopsis thaliana, partial (30%)
Length = 1701
Score = 38.1 bits (87), Expect = 0.016
Identities = 32/154 (20%), Positives = 65/154 (41%), Gaps = 12/154 (7%)
Frame = +3
Query: 307 VNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEV-----------NSRLEDRSFPQPEL 355
+ P P ST K G++ + + +D QPE+ N+ + DR+ +L
Sbjct: 321 IRLPAPTESTAKSGEATSETQPEKAKDEETKQPEIKAPEVEDKSTSNNDVADRNNAGKDL 500
Query: 356 NSKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPVANSTSE-DLNLAQPLSQSVSD 414
K D + + DN +DG+ ++E + VA+ S ++N P + ++
Sbjct: 501 AEKETDSDNSKVDNEQSKDGS--------KIENEDKKEVADKESAGEINKEPPTEEKNTE 656
Query: 415 DLHINQQAEPSNLNVRQEDDRPSSPIHRQGAISD 448
+ N +E + ++D+ S ++ +SD
Sbjct: 657 NSDKNGNSES-----KDKEDKVSESKDKEDKVSD 743
Score = 34.3 bits (77), Expect = 0.24
Identities = 25/124 (20%), Positives = 50/124 (40%), Gaps = 7/124 (5%)
Frame = +3
Query: 264 QKTEDQSLAQPPVSS-RTDDGDSAQQQVEDQNLAQTHVSSRTEDVNSPRPQNSTQKDG-- 320
+K +D+ QP + + +D ++ V D+N A ++ + D ++ + N KDG
Sbjct: 387 EKAKDEETKQPEIKAPEVEDKSTSNNDVADRNNAGKDLAEKETDSDNSKVDNEQSKDGSK 566
Query: 321 ----DSPQILENSRPEDASLTQPEVNSRLEDRSFPQPELNSKLEDMTSQQQDNSILEDGN 376
D ++ + + + P E+ + ED S+ +D ED
Sbjct: 567 IENEDKKEVADKESAGEINKEPPTEEKNTENSDKNGNSESKDKEDKVSESKDK---EDKV 737
Query: 377 SSQP 380
S +P
Sbjct: 738 SDEP 749
>TC87092 similar to PIR|D84900|D84900 hypothetical protein At2g46240
[imported] - Arabidopsis thaliana, partial (5%)
Length = 1878
Score = 37.4 bits (85), Expect = 0.028
Identities = 39/212 (18%), Positives = 83/212 (38%), Gaps = 16/212 (7%)
Frame = +3
Query: 261 SPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNLAQTHVSSRTEDVNSPRPQNSTQKDG 320
+P Q+ + + V+S + Q+ +++ +A SS SP+ Q + D
Sbjct: 282 NPCQQPHEDAKDPVEVTSLNVQNEKLNQEQQEEKVASEKDSSEGTSDGSPKEQFCMKDDD 461
Query: 321 DSPQILENSRPEDASLTQPEVNSRLEDRS------------FPQPELNSKLEDMTSQQQD 368
+ + + T+P V+S L +R+ P + + D TS D
Sbjct: 462 GRSESRSHVDSASSERTKPHVDSALSERTKTTMLPNGLINEDSSPVMAADASDSTSDLVD 641
Query: 369 NSILEDGNSSQ----PQLNLRLEGSSLQPVANSTSEDLNLAQPLSQSVSDDLHINQQAEP 424
+ LE + S+ P + +L+ ++L+ ++D S+ + D+H ++
Sbjct: 642 KTDLECKSKSEVIDIPIVVDKLDTTALKDSPVGANDDNISDNSASEGLDSDMHALKELP- 818
Query: 425 SNLNVRQEDDRPSSPIHRQGAISDNLHSHQQV 456
+ V ED + G + +H+ +V
Sbjct: 819 --VGVLDEDTATFEGTNTSGNVQSEVHAENEV 908
>TC91642 similar to PIR|E96613|E96613 hypothetical protein T15M6.22
[imported] - Arabidopsis thaliana, partial (11%)
Length = 949
Score = 36.2 bits (82), Expect = 0.062
Identities = 16/36 (44%), Positives = 23/36 (63%)
Frame = +3
Query: 662 LLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPM 697
++ K+M+ A FN PV+ + LG+ DYF II PM
Sbjct: 840 IIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPM 947
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.128 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,166,285
Number of Sequences: 36976
Number of extensions: 349801
Number of successful extensions: 2086
Number of sequences better than 10.0: 92
Number of HSP's better than 10.0 without gapping: 1950
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2057
length of query: 966
length of database: 9,014,727
effective HSP length: 105
effective length of query: 861
effective length of database: 5,132,247
effective search space: 4418864667
effective search space used: 4418864667
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144893.5


